- TECHNOLOGY FEATURE
Find a home for every imaging data set

Credit: The Project Twins
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 579, 162-163 (2020)
doi: https://doi.org/10.1038/d41586-020-00594-4
References
Zivanov, J., Nakane, T. & Scheres, S. H. W. IUCrJ https://doi.org/10.1107/S2052252520000081 (2020).
Dorkenwald, S. et al. Preprint at bioRxiv https://doi.org/10.1101/2019.12.29.890319 (2019).
McDole, K. et al. Cell 175, 859–876 (2018).
Williams, E. et al. Nature Meth. 14, 775–781 (2017).
Colavizza, G., Hrynaszkiewicz, I., Staden, I., Whitaker, K. & McGillivray, B. Preprint at https://arxiv.org/abs/1907.02565 (2019).

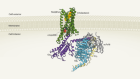
 Cryo-electron microscopy shapes up
Cryo-electron microscopy shapes up
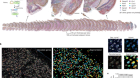
 The microscope makers putting ever-larger biological samples under the spotlight
The microscope makers putting ever-larger biological samples under the spotlight
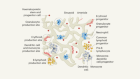
 Web service makes big data available to neuroscientists
Web service makes big data available to neuroscientists
 NatureTech hub
NatureTech hub