- NEWS AND VIEWS
The clock that controls spine development modelled in a dish
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 580, 32-34 (2020)
doi: https://doi.org/10.1038/d41586-020-00322-y
References
Herrgen, L. et al. Curr. Biol. 20, 1244–1253 (2010).
Hubaud, A. & Pourquié, O. Nature Rev. Mol. Cell Biol. 15, 709–721 (2014).
Oates, A. C., Morelli, L. G. & Ares, S. Development 139, 625–639 (2012).
Diaz-Cuadros, M. et al. Nature 580, 113–118 (2020).
Matsuda, M. et al. Nature 580, 124–129 (2020).
Yoshioka-Kobayashi, K. et al. Nature 580, 119–123 (2020).
Chal, J. et al. Nature Biotechnol. 33, 962–969 (2015).
Loh, K. M. et al. Cell 166, 451–467 (2016).
Xi, H. et al. Cell Rep. 18, 1573–1585 (2017).
Bessho, Y. et al. Genes Dev. 15, 2642–2647 (2001).
Hofmann, M. et al. Genes Dev. 18, 2712–2717 (2004).
Wahl, M. B., Deng, C., Lewandoski, M. & Pourquié, O. Development 134, 4033–4041 (2007).
Sparrow, D. B. et al. Am. J. Hum. Genet. 78, 28–37 (2006).
Whittock, N. V. et al. Am. J. Hum. Genet. 74, 1249–1254 (2004).
Sparrow, D. B., Guillén-Navarro, E., Fatkin, D. & Dunwoodie, S. L. Hum. Mol. Genet. 17, 3761–3766 (2008).
Jiang, Y.-J. et al. Nature 408, 475–479 (2000).
Jouve, C. et al. Development 127, 1421–1429 (2000).
Isomura, A., Ogushi, F., Kori, H. & Kageyama, R. Genes Dev. 31, 524–535 (2017).

 Raed the paper: In vitro characterization of the human segmentation clock
Raed the paper: In vitro characterization of the human segmentation clock
 Read the paper: Coupling delay controls synchronized oscillation in the segmentation clock
Read the paper: Coupling delay controls synchronized oscillation in the segmentation clock
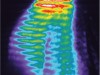 Read the paper: Recapitulating the human segmentation clock with pluripotent stem cells
Read the paper: Recapitulating the human segmentation clock with pluripotent stem cells
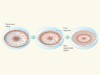 Segmentation within scale
Segmentation within scale
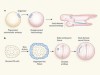 Human embryonic stem cells get organized
Human embryonic stem cells get organized