This is a transcript of the 19th January, 2017 edition of the weekly Nature Podcast. Audio files for the current show and archive episodes can be accessed from the Nature Podcast index page (http://www.nature.com/nature/podcast), which also contains details on how to subscribe to the Nature Podcast for FREE, and has troubleshooting top-tips. Send us your feedback to podcast@nature.com.
[Jingle]
Noah Baker: Coming up – explaining the mysterious 'fairy circles' in the Namibian desert.
Corina Tarnita: I find that, in some sense, a whole lot more satisfying than fairies [laughs].
Noah Baker: And the news that viruses can talk to each other.
Nonia Pariente: I had heard this story, and it didn't disappoint.
Noah Baker: Plus, a new analysis aims to reproduce cancer studies. Do the results hold up? This is The Nature Podcast for January 19th, 2017. I'm Noah Baker.
Adam Levy: And I'm Adam Levy.
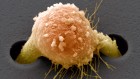
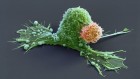
Noah Baker: Call it déjà vu, but this week, a handful of papers are being published that might ring a bell. That's because they're replications, do-overs of high profile papers published in the last few years. Just how easy is it to do the same experiment and get the same result? Kerri takes a look.
Kerri Smith: At the end of October 2013, cancer biologist Erkki Ruoslahti got an email he wasn't expecting.
Erkki Ruoslahti: Well, I wasn't too happy about it.
Kerri Smith: The email told Erkki that a paper of his, a high profile article from 2010 about a way of delivering drugs to tumors, had been chosen for an ambitious replication effort. His wasn't the only one. An independent team of scientists were planning to dissect 50 cancer papers, assemble the kits and reagents they needed to do them over again, and try to reproduce the results.
Erkki Ruoslahti: The people from the program contacted me. I can't say that it's a pleasure.
Kerri Smith: The effort is part of a program called the Reproducibility Project: Cancer Biology. It's run by Elizabeth Iorns.
Elizabeth Iorns: Elizabeth Iorns, CEO of Science Exchange.
Kerri Smith: And Tim Errington.
Timothy Errington: Timothy Errington. I'm meta-science manager at the Center for Open Science.
Kerri Smith: And it aims to examine key findings in cancer biology to see what proportion of them stand up.
Jingle
It came about because some pharmaceutical companies reported having trouble with replicating work in the area.
Elizabeth Iorns: So, Amgen and Bayer had published studies showing that their rates of being able to reproduce published results were around 20 to 30 percent. And obviously there's been a lot of media attention around this issue, but there hasn't been any open projects to actually examine the rates of reproducibility by doing replication studies. And so this project was the first to attempt that.
Kerri Smith: The project leaders knew it wasn't going to be easy to do carbon copies of the experiments.
Elizabeth Iorns: We anticipated that there would be challenges around the models behaving as expected, and being able to access enough information about the protocols.
Timothy Errington: The materials that weren't commercially available.
Elizabeth Iorns: Being able to access original reagents, or even identify unique reagents.
Timothy Errington: Trying to just understand what to do.
Kerri Smith: They got going in 2013, with over $1 million of funding from a foundation and 50 papers to reproduce. A couple of years later, they scaled back their efforts to about 30 papers, citing the expense. And now they published the results of the first five replications. Erkki Ruoslahti's paper is one of them. Spoiler alert. Scientists have not been able to get his drug to work.
Erkki Ruoslahti: Well, I was disappointed, but not entirely surprised. No.
Kerri Smith: Erkki thinks the replication failed for a few reasons. He doesn't think they used enough mice. He says they didn't spend time troubleshooting once they got one negative result. The first experiments the team tried to replicate were to see if Erkki's peptide actually built up in tumors in the first place.
Erkki Ruoslahti: Once they found that they didn't do that, then there was no point in going further. But they just went on and did a treatment study, which based on the first result was doomed.
Kerri Smith: The reason this might have happened is because the reproducibility lab was obliged to stick to the recipes and procedures they'd set out in meticulous reports they published in advance of their experiments. These reports are peer reviewed, and Erkki and others fed back on the one concerning his work. The other four studies being published this week fared marginally better. Two mostly substantiated, but too inconclusive.
Timothy Errington: These first five show that it's mixed, that there's quite a range of variability between one study and the other, and I think it really points to what we're potentially going to start seeing with these other ones.
Kerri Smith: Nature spoke to all five corresponding authors from the original papers for a news article, which you can read on our website. Some praise the effort, and some worry that negative results might hold repercussions for them. Elizabeth Iorns is quick to add that these results just add a few new data points, rather than overturning the work that came before.
Elizabeth Iorns: We want people to read the papers and interpret the results themselves, and have an open discussion about what reproducibility really means.
Kerri Smith: For some scientists, replication is just something done as standard. Erkki says upwards of a dozen other labs have already replicated the results from his original paper.
Erkki Ruoslahti: And then there's more in the pipeline.
Kerri Smith: He thinks that should be enough. But the project is about more than just this handful of papers. It also illustrates some of the fundamental and really quite ordinary challenges to redoing science. In a lot of cases, it's boring old logistics that could let future reproducibility down.
Elizabeth Iorns: So for example it was difficult for us to obtain information about the original published studies, just because it wasn't standard practice to publish raw data. It wasn't standard practice to publish full protocols, and it wasn't standard practice to publish unique identifiers for the reagents using those studies.
Kerri Smith: So much for doing the replication, communicating the result is also going to be important. Especially for the scientists whose work was chosen. Here's Erkki.
Elizabeth Iorns: I'm concerned that the negative methods and the outsized publicity that this will receive will bias people against maybe our grants and some other things we do.
Kerri Smith: In fact, he has a more pressing concern.
Erkki Ruoslahti: We're about halfway through doing the preclinical work that is needed to get this into the clinic. I'm really worried that this will spook potential investors.
Kerri Smith: Erkki says that could damage his attempts to produce a cancer drug. Elizabeth Iorns and Tim Errington urge others not to read these studies as simple yes or no answers.
Timothy Errington: These are single replications. Just like what we're replicating is a single study. And right now the current system is that we publish, we find some result, and then we move on, and what we're trying to say is well how does that look, how does any piece of evidence look? This is really getting not so much at trying to say anything about the entire field. There's lots of pieces of evidence. This is trying to get more at looking at one individual piece of evidence, and really getting at how are we conducting our research, and how are we communicating our research?
Kerri Smith: So what would be their one tip for a study that will successfully replicate? I'll give you a clue. Organization, organization, organization.
Timothy Errington: Plan ahead from the beginning. The very first time when you're conducting that study think about your future self, and think about all the other scientists who are going to want to read this, understand it, eventually reuse it and build upon it.
Kerri Smith: The first five replication studies, together with their registered reports, those planning documents published before the experiments began, are being published tomorrow, Thursday, January 19, by the journal E-Life. You can find all the details at ELifeSciences.org. With luck, says Tim, the rest of the 29 studies they're working on will be out by the end of 2017. Tim Errington is from the Open Science Framework. He co-runs the project with Elizabeth Iorns of Science Exchange. Erkki Ruoslahti is at the Sanford-Burnham Medical Research Institute in La Jolla, California.
Noah Baker: Still to come in the research highlights – an ambitious diving somersault, and a tiny little knot. But, before that, Adam's been investigating a mysterious pattern.
Adam Levy: If you fly over the Namib Desert in Southern Africa, you might spot something strange, something almost magical in the grasses below. In places, the vegetation is dotted with strange scars. Round patches measuring a few meters across, where the grass doesn't grow.
Corina Tarnita: Hundreds of thousands, there must be. I mean, they stretch on hundreds of kilometers.
Adam Levy: This is ecologist Corina Tarnita. What's more, these mysterious barren patches aren't just scattered about randomly. They're organized.
Corina Tarnita: If you look at any one of these circles, and you count their neighbors, it's always roughly the same number. It's always six circles. They're very regularly spaced. So it looks kind of like a polka dot dress.
Adam Levy: These Namibian polka dots have a name: fairy circles. So where did these fairy circles come from? There are many suggestions for their origin, as mystical as their name suggests. Depending on the story, their creators could be anything from dragons to ancient ancestors. There are plenty of more empirical proposals too. At the moment, the two most prominent hypotheses involve either underground termite colonies competing with neighbors for territory, or the plants themselves competing for a scarce supply of water. Both hypotheses hope to explain the size, shape, and organization of these circles. Physicist Ehud Meron has been investigating how the second of these hypotheses, plant competition, could lead to these fairy circles.
Ehud Meron: We developed, long ago, a mathematical model to explain vegetation patterns in general. This works beautifully for vegetation patterns. This model, we don't have termites at all.
Adam Levy: Ehud's model can produce many of the observed qualities of fairy circles with no help from termites. So that's one-nil for the plant hypothesis. But termite mounds are found in the fairy circles of Namibia. Corina and her team wondered whether a model of competition between termites could also explain the patches.
Corina Tarnita: We started to wonder, might it be possible that you could get superficially the same patterns from very different processes? So the first thing that we wanted to do is see, can we make a model that takes into account insect behavior, and see if it's possible for termites to produce these kinds of patterns.
Adam Levy: Lo and behold, when they used their termite model to try and explain the fairy circles, it also seemed to capture the key elements of the rings. So that's one all. We're back where we started, with seemingly little way to choose between hypotheses.
Corina Tarnita: For us, the immediate insight was what if both of them are right?
Adam Levy: So, in a paper out this week, Corina and collaborators set about combining these two hypotheses into one by using a model that simulates competition between termite mound and competition between plants for scarce water. They found that like the individual hypotheses, it captured the key fairy ring features.
Corina Tarnita: But it also made this additional prediction that if both of these mechanisms are occurring in this system simultaneously, then the vegetation, when it self organizes, it forms this tiny clump of grass, and every clump of grass should have about six neighbors.
Adam Levy: So the model not only predicted that each fairy circle should have six fairy circle neighbors, but that the vegetation in between the circles should be clumped, and these clumps should also have six neighbors. This prediction seemed to offer a clear cut way to finally settle what causes the fairy circles. If true, it would mean that everyone had been so distracted by the circles themselves that they'd missed this far smaller pattern in the vegetation. Corina's team quickly arranged to get more detailed photos of the rings.
Corina Tarnita: We were waiting on the edge of our seats to see what will happen when they send us the photos. And they did. And it was really very satisfying. Because it became immediately clear we were finding this exact same regularity that the model was predicting.
Adam Levy: Exactly the same regularity. So surely that finally closes the case on the fairy circle mystery.
Ehud Meron: It's a very nice paper. Has interesting results.
Adam Levy: This is Ehud Meron, who we heard from earlier. But Ehud is not convinced we should abandon the plant competition hypothesis just yet. He reckons that if the plant competition model is set up to take account of water transport by plant roots, they could also get the same result.
Ehud Meron: We can lift the surface due to the roots. And get wherever there is vegetation, get the small scale pictures. I'm pretty sure we can do it. We haven't done it yet.
Adam Levy: Although Ehud and his collaborators haven't tried that particular experiment, he explains they have another reason to stick with their termite free hypothesis. The Namib Desert isn't the only place with fairy circles. Similar patterns have been spotted in Australia, which seem to lack the regular termite mounds found in their Namibian counterparts. Ultimately though, Corina thinks that this debate won't be settled until the hypotheses are truly tested. And to truly test a hypothesis, researchers need to get out to the field.
Corina Tarnita: Models are always very helpful to bring all these ideas together and to make new predictions. But I don't think the case can be closed until you have some manipulative experimental field experiments done in the system, where termites are killed or displaced to see what happens to the circles, or plants are killed on certain areas, or water is added on certain areas.
Adam Levy: So hopefully one day researchers will carry out experiments like these, and the fairy circle mystery will finally be put to bed. But wouldn't it be kind of sad for this incredible phenomenon to lose its mystique? Corina doesn't think so.
Corina Tarnita: To me, knowing what's behind them, and knowing that tiny things like termites or plants can create patterns on the scales of hundreds of kilometers, that, to me, is mind boggling. I find that in some sense a whole lot more satisfying than fairies [laughs].
Adam Levy: That was Corina Tarnita, Princeton University in the US. Her paper is out now at http://Nature.com/nature. You also heard from Ehud Meron, who is at Ben-Gurion University in Israel.
Noah Baker: Stay tuned for the weather. The space weather, that is. That's in the News chat. Now, though, it's time for the Research Highlights, read this week by Shamini Bundell.
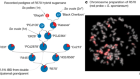
Shamini Bundell: You are not going to believe this. Scientists have plaited three tiny synthetic strands into the most complex knot yet made in the lab. The strands cross in eight places, and the whole thing is only 20 nanometers long. Flexible polymers can knot themselves, but so far, scientists have only managed to engineer very simple loops. This team used little specks of iron to control the positions of the strands as they assembled into the knot. Structures like this could be useful for making new materials that are tough and flexible. Find that paper in Science.Olympic divers are always keen to try out the most ambitious tricks. The more complicated the dive, the more points they can get. Now Australian mathematicians are helping out by modeling a new dive, the calculations for which are published in a study on the pre-print server archive. Involving one and a half somersaults, and five twists, the dive is currently completely theoretical but they claim it should be physical possible. It may be mathematically sound, but it requires such precise and rapid movements that a representative of British diving, quoted in the Guardian, described the dive as a bit of a non-starter.
Noah Baker: Imagine you're a virus particle, floating outside your next bacterial host. You're preparing to infect the cell, injecting your DNA into the bacterium, and tricking its cellular machinery into replicating your own DNA. But you have a choice. You see, you're a temperate bacteria phage, phi3T, to be exact. And viruses like you have two phage cycles to choose from. Here's nature and microbiology editor, Nonia Pariente, to explain.
Nonia Pariente: When a temperate virus infect, it can choose between two different types of cycle. In one it will prime its own reproduction, so it will go into the cell, it will produce many copies of itself, and, in leaving the bacterial cell, these copies will lyse the bacteria. And so a lot of virus will be produced, but the bacteria will die.
Noah Baker: This is called the lytic cycle. But you have a second choice.
Nonia Pariente: The lysogenic cycle is where a virus will integrate its genomic material into the genome of the bacterium – so if it's host – and it will stay there dormant. The bacteria will not die there, for it will just have an extra piece of DNA. But this virus could potentially reactivate in the future, so it's saving itself from death, if you will. Although talking of death without life is weird. But –
Noah Baker: Viruses are not considered to be alive, so can't really die, as such. But we'll talk more about that later. Back to your choice. As you float outside your bacterial host cell, ready to infect and kill it by replicating, or let it stay alive, and lie dormant. If you're the first of your kind on the scene, surely you'd like to kill and replicate, infecting all the bacteria you can. But if you're later to the party, most of the bacteria will already have been killed, and your progeny may not be able to find any more bacteria to infect. In that case, you should lie in wait until bacterial numbers recover. So how do you choose? Especially since you don't have a brain.With trepidation, you engage your bacterial host and set about injecting your DNA into its cytoplasm. And then, in the distance, you notice a signal. This signal is a peptide molecule called arbitrium, which means decision, and it acts like a simple messenger. The arbitrium binds to receptors that you've had the bacteria make for you. Arbitrium is produced by other viruses like you. Indeed, you go ahead and instruct the bacteria to produce some arbitrium for you, too. The more viruses that have been in the vicinity, the more arbitrium will have been produced. And if there's enough, it will overrun your receptors. The chemical din in your bacterium is deafening.Now, here's the clever bit. That receptor, which arbitrium is binding to, has another job. It inhibits lysogeny, the cycle in which viruses lie dormant in the bacterial genome, and don't kill it. All this arbitrium has bound to your receptors which means they can no longer do their other job of stopping lysogeny from happening. It's like the brakes on lysogeny have been lifted, and so you lysogenize, and lie dormant in the bacteria's genome. In this way, you've detected how many viruses are in the vicinity, and decided which cycle to progress down, all without a brain.This simple but ingenious process is called quorum sensing. And, up until now, it's only been seen in bacteria. For Nonia, finding it in viruses was an exciting moment.
Nonia Pariente: You read a lot of papers, and some good and some bad, but you can remember less than five per year I would say that are truly groundbreaking, that you feel excited about. And it didn't disappoint. And actually my notes say this. You know I had heard this story and it didn't disappoint.
Noah Baker: It was a surprise, too, to the authors of the paper. They thought the communication was between the bacteria. Here's Rotem Sorek, from the Weizmann Institute of Science in Israel.
Rotem Sorek: What we were looking for, we chased the hypothesis that maybe bacteria communicate between them in order to alert that phage of infected culture. It was a little bit different to the hypothesis that we were following, and we were completely surprised to find basically the communication appearing from the phage.
Noah Baker: Rotem didn't just find these viruses could talk to each other. He also found out that in some rudimentary way they have different languages.
Rotem Sorek: Different phages have different kinds of molecules, and so one phage's tissue, it only understands its own language. So basically many different phages stick with different languages, and each phage understands its own language.
Noah Baker: This in particular excited Nonia.
Nonia Pariente: The viruses that a virus spawns, if you will, will be able to detect what its progenitors produce, and not what progenitors of other kinds produce. So they talk to each other in very specific languages, and of course it also opens the possibility that if they talk about this one particular decision, they may talk about other things as well.
Noah Baker: Rotem says that not all temperate viruses communicate like this. But it could be more common than scientists might have thought.
Rotem Sorek: We found it in hundreds of cases in nature, and we don't know yet if this kind of communication system is more broad or not. I hope that our paper causes future research that we look whether other temperate phages use such communication.
Noah Baker: Rotem's work brings up an even more fundamental question. With the help of their host, viruses can replicate, they can evolve, and speciate, and now they can communicate: all things which are associated with life. And yet viruses are not considered to be alive. Which raises the question, should they be?
Nonia Pariente: It's a very hard question, philosophical question at the root. So I am not sure. I really think that this takes us a step closer to them having the behavior of living. But well I am a very biased person, as a virologist, and I've always thought [laughs] viruses should be alive.
Noah Baker: That was Nonia Pariente, a senior editor for nature microbiology, who handled this paper for Nature. And, before her, you heard Rotem Sorek from the Weizmann Institute of Science in Israel. You can read the paper, along with the news and views all about the discovery, over at http://Nature.com/nature/.
Adam Levy: Finally, this week, it's the News Chat, and Davide Castelvecchi joins in the studio. Hi, Davide.
Davide Castelvecchi: Hi, Adam.
Adam Levy: First to India. Now, India seemed on the verge of approving GM – genetically modified mustard. But there's a holdup. So what's going on here?
Davide Castelvecchi: Yeah. So the Indian or an agency of the Indian Environmental Ministry had one a safety study, and concluded there were no safety problems with this crop, which would be the first genetically modified transgenic food crop to be approved in the country. But there is now a lawsuit, which is pending with the Supreme Court, and it's been there for a while and it's not clear when the Supreme Court will actually make the decision.
Adam Levy: What are the benefits of this genetically modified crop meant to be over conventional mustard?
Davide Castelvecchi: The main benefit that has been mentioned is an increase in yields. This is a crop that is used mostly for making oil, the seeds are pressed to make oil, and it was traditionally used in Northern India and Pakistan and Bangladesh. But it has been taken over by a lot of other cheaper oils, some of which may be imported, so that the argument was we increase the yield of mustard seeds and of mustard oil, and we will be able to produce more of it for cheaper.
Adam Levy: So what about accusations here, what's the cause of the holdup?
Davide Castelvecchi: Part of it is the familiar concerns about transgenic crops. About safety in particular , the fact that you take a gene from one organism, you put it into another, and there could be unforeseen consequences. And then there's also the worry that if this particular type of crop takes over, then the company that produces it will gain too much power over the market.And also the plaintiffs are saying that the benefits, the potential benefits of this crop, have been exaggerated, and particularly the safety tests were not really designed to maximize the yield of the crops that were being compared. And so it's not clear exactly how much more yield one could get from this particular crop.
Adam Levy: How does this fit into the wider GM debate in India?
Davide Castelvecchi: So the wider GM debate… There was one non-food crop which is currently approved, which is cotton, a type of cotton, which has been enormously controversial. There have been people who have blamed it for causing farmers to commit suicide, and a lot of these claims were not very well substantiated. But there was a hope that when Prime Minister Modi came to power in 2014 that because it was seen as being more open to testing and potentially approving these GM crops, that the whole pipeline might be restarted. So this could be a test, this particular crop, even though it's only one maybe not even major crop. It's been seen as a test.
Adam Levy: Do we have any timeline for when the Supreme Court would make a decision on this?
Davide Castelvecchi: So apparently, there is no deadline, and the court has given no indication of when it might reach a decision.
Adam Levy: Well, let's move on from this project which I suppose has been put on hold to a project which seems to be on the way to getting the green light, and that's a European Space Agency mission to launch a new probe. But this probe isn't going to the far reaches of the solar system, right?
Davide Castelvecchi: No, but it would go far enough away from the earth. It would be orbiting the sun at one of the so-called Lagrange points, which are kind of like a balancing act between the gravity of the sun and the earth. And the reason why they would choose this location is that it would be a good viewing point to see solar storms in the process of being ejected from the sun, but before they reach Earth and potentially cause havoc.
Adam Levy: Aren't there already probes which monitor space weather?
Davide Castelvecchi: Yes, there are. And there are probes that study in general – that study the sun – both built by NASA and ESA. But this one would be peculiar. It would be kind of unprecedented in that it would have this vantage point from which it can tell when a solar storm in particular the most dangerous type, which is called a coronal mass ejection, it would see it on the approach to Earth, and it would be able to give longer advance notice, for example, to grid operators, so that they can take precautions to avoid a lot of damage.
Adam Levy: Is that actually a big risk of these things? And would they cause really bad damage?
Davide Castelvecchi: These really extreme space weather events are not that uncommon. But they don't always get directed at Earth and the last one to when we can actually see it from Earth. So if there is a menacing sunspot, for example, something that looks like it could produce a big solar storm, we would be able to see it much longer in advance.
Adam Levy: So it seems like quite a reassuring thing to have then, this warning system about these huge solar storms. Do we know when it's actually going to be out there?
Davide Castelvecchi: So it's still at an early stage. So far, the Council of Ministers has approved the initial funding for R&D, which is about 30 million Euros. And it will certainly be several years before the final funding is approved, which is expected to cost about 450 million Euros. So we're talking about at least 2020 – it's not clear how long.
Adam Levy: Thank you for joining us, Davide. For more on those stories, http://Nature.com/news. And to stay up to date with all the latest science news, make sure to follow Nature News on Twitter, @NatureNews.
Noah Baker: And, while you're there, give the podcast a follow, too, @NaturePodcast. That's all for this week. I'm Noah Baker.
Adam Levy: And I'm Adam Levy.
Jingle