- NEWS AND VIEWS
An ‘on’ switch for proteins
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 569, 490-491 (2019)
doi: https://doi.org/10.1038/d41586-019-01394-1
References
Rost, B. R., Schneider-Warme, F., Schmitz, D. & Hegemann, P. Neuron 96, 572–603 (2017).
Dagliyan, O. & Hahn, K. M. Curr. Opin. Struct. Biol. 57, 17–22 (2019).
Wang, J. et al. Nature https://doi.org/10.1038/s41586-019-1188-1 (2019).
Arbely, E., Torres-Kolbus, J., Deiters, A. & Chin, J. W. J. Am. Chem. Soc. 134, 11912–11915 (2012).
Krall, N., da Cruz, F. P., Boutureira, O. & Bernardes, G. J. L. Nature Chem. 8, 103–113 (2016).
Chin, J. W. Annu. Rev. Biochem. 83, 379–408 (2014).

 Read the paper: Time-resolved protein activation by proximal decaging in living systems
Read the paper: Time-resolved protein activation by proximal decaging in living systems
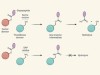 Enzymes engineered to trap reaction intermediates
Enzymes engineered to trap reaction intermediates
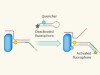 DNA tags used to image sugar-bearing proteins on cells
DNA tags used to image sugar-bearing proteins on cells