- NEWS AND VIEWS
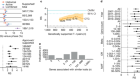
Genetic clues can be used to predict whether early-stage cancer will form an invasive tumour
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 566, 336-337 (2019)
doi: https://doi.org/10.1038/d41586-019-00567-2
References
Teixeira, V. H. et al. Nature Med. https://dx.doi.org/10.1038/s41591-018-0323-0 (2019).
Hoadley, K. A. et al. Cell 158, 929–944 (2014).
Campbell, J. D. et al. Nature Genet. 48, 607–616 (2016).
Casasent, A. K., Edgerton, M. & Navin, N. E. J. Pathol. 241, 208–218 (2017).
Martincorena, I. et al. Science 348, 880–886 (2015).
Martincorena, I. et al. Science 362, 911–917 (2018).
Yokoyama, A. et al. Nature 565, 312–317 (2019).

 Read the paper: Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions
Read the paper: Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions
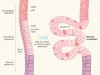 Origins in the oesophagus
Origins in the oesophagus
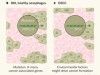 Mutations differ in normal and cancer cells of the oesophagus
Mutations differ in normal and cancer cells of the oesophagus