- NEWS AND VIEWS
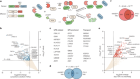
Enzymes engineered to trap reaction intermediates
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 565, 28-29 (2019)
doi: https://doi.org/10.1038/d41586-018-07569-6
References
Wang. L., Brock, A., Herberich, B. & Schultz, P. G. Science 292, 498–500 (2001).
Huguenin-Dezot, N. et al. Nature 565, 112–117 (2019).
Payne, J. A., Schoppet, M., Hansen, M. H. & Cryle, M. J. Mol. Biosyst. 13, 9–22 (2016).
Cheng, Y. Q. ChemBioChem. 7, 471–477 (2006).
Olsen, S. K., Capili, A. D., Lu, X., Tan, D. S. & Lima, C. D. Nature 463, 906–912 (2010).
Bachovchin, D. A. & Cravatt, B. F. Nature Rev. Drug Discov. 11, 52–68 (2012).
Lentz, C. S. et al. Nature Chem. Biol. 14, 609–617 (2018).
Gulick, A. M. & Aldrich C. C. Nat. Prod. Rep. 35, 1156–1184 (2018).
Ye, Z. & Williams, G. J. Biochemistry 53, 7494–7502 (2014).
Backus K. M. et al. Nature 534, 570–574 (2016).

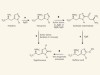
 Read the paper: Trapping biosynthetic acyl-enzyme intermediates with encoded 2,3-diaminopropionic acid
Read the paper: Trapping biosynthetic acyl-enzyme intermediates with encoded 2,3-diaminopropionic acid
 The surprising history of an antioxidant
The surprising history of an antioxidant
 A toxin that fuels metabolism
A toxin that fuels metabolism