- NEWS
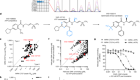
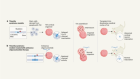
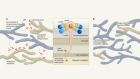
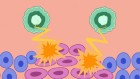
Nigeria’s largest Lassa fever outbreak sparked by rats
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
doi: https://doi.org/10.1038/d41586-018-07024-6
References
Siddle, K.J. et al. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa1804498 (2018).