- NEWS AND VIEWS
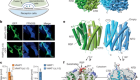
DNA tags used to image sugar-bearing proteins on cells
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 561, 38-40 (2018)
doi: https://doi.org/10.1038/d41586-018-06092-y
References
Varki, A. Glycobiology 27, 3–49 (2017).
Li, S. et al. Angew. Chem. Int. Edn https://doi.org/10.1002/anie.201807054 (2018).
Miyawaki, A. Nature Rev. Mol. Cell Biol. 12, 656–668 (2011).
Hori, Y. & Kikuchi, K. Curr. Opin. Chem. Biol. 17, 644–650 (2013).
Haga, Y. et al. Nature Commun. 3, 907 (2012).
Lin, W., Du, Y., Zhu, Y. & Chen, X. J. Am. Chem. Soc. 136, 679–687 (2014).
Belardi, B. et al. Angew. Chem. Int. Edn 52, 14045–14049 (2013).
Doll, F. et al. Angew. Chem. Int. Edn 55, 2262–2266 (2016).
Laughlin, S. T. & Bertozzi, C. R. Nature Protoc. 2, 2930–2944 (2007).
Wu, N., Bao, L., Ding, L & Ju, H. Angew. Chem. Int. Edn 55, 5220–5224 (2016).
Hui, J. et al. Angew. Chem. Int. Edn 56, 8139–8143 (2017).

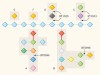 A wine-induced breakdown
A wine-induced breakdown
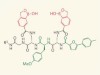 Sticking to sugars
Sticking to sugars