This is a transcript of the 21st June 2018 edition of the weekly Nature Podcast. Audio files for the current show and archive episodes can be accessed from the Nature Podcast index page, which also contains details on how to subscribe to the Nature Podcast for FREE, and has troubleshooting top-tips. Send us your feedback to podcast@nature.com.
[Jingle]
Host: Benjamin Thompson
Welcome back to the Nature Podcast. This week, we’ll be learning about the ethics of AI algorithms and we’ll be finding out about the underlying causes of weight loss in pancreatic cancer.
Host: Shamini Bundell
Plus, we’ll be hearing about super small self-assembling silica structures.
Host: Benjamin Thompson
Sounds super.
Host: Shamini Bundell
Certainly does. This is the Nature Podcast for the 21st June 2018. I’m Shamini Bundell.
Host: Benjamin Thompson
And I’m Benjamin Thompson.
[Jingle]
Host: Shamini Bundell
First up this week, Noah Baker has been looking very, very closely at a paper about mesoporous silica. In part, because he’s a diligent journalist of course, but mostly because the materials it describes are very, very small.
Interviewer: Noah Baker
If a lump of material has holes in it you’d call it porous. If those pores were between 5 and 20 nanometres across, you’d call it mesoporous. OK, maybe you wouldn’t but material scientists would. Mesoporous materials, because of all those tiny holes, have a really high surface area and that makes them great for things like absorbing gases or filtering air and water. They can come in all shapes and sizes, from blocks you can hold in your hands to microscopic particles. But how small can mesoporous materials get? What would they look like and how would you even see something that tiny? Well, that’s what I’m looking at this week – the story of how scientists made and imaged a new and exquisite ultra-small mesoporous particle out of silica. Uli Wiesner from Cornell University in the States led the team. Here he is with a bit of background on how to make a mesoporous silica material.
Interviewee: Uli Wiesner
It’s actually a fairly easy process. The only thing you need is water, typically, then a silica precursor and then you also need a template over which the silica will condense and then form the mesoporous structure.
Interviewer: Noah Baker
The template molecule, in this case surfactants, in other words soap, can then be washed or burned out after the silica structure has formed around them, leaving behind pores. This relatively simple technique was first published in Nature in 1992 and it was very popular. In fact, the paper went on to become one of Nature’s most cited papers ever. Here’s Nature editor Claire Hansell. She focuses on chemistry and materials science.
Interviewee: Claire Hansell
No one can ever quite predict citations or the impact a paper’s going to have, but as it happens, mesoporous silica as a material has really taken off in all sorts of areas. It’s become an incredibly important material in its own right.
Interviewer: Noah Baker
Using this protocol, scientists have made all kinds of mesoporous silica structures, including Uli. He and his team decided to go small.
Interviewee: Uli Wiesner
We are interested in extremely small silica nanoparticles in general below, say, 10 nanometres.
Interviewer: Noah Baker
Uli hopes that such ultra-small particles could be used in cancer treatments as a vehicle to deliver drugs or in diagnostic tests. One of Uli’s molecules is actually the focus of a stage two clinical trial. One of the key advantages of such tiny particles in a medical setting is that they’re excreted in the urine through the kidneys, unlike many other materials which are processed through the liver. The liver route takes longer and can lead to more side effects. Anyway, it was while exploring the synthesis of particles like this that Uli discovered his latest ultra-small particle. He stumbled across it while trying to image tiny silica rings.
Interviewee: Uli Wiesner
That got us to look into this in much more detail. We started to use a technique which is well-known in the biological area called cryo-electron microscopy.
Interviewer: Noah Baker
Cryo-electron microscopy is usually used to snap pics of biological tiny things like proteins. Here’s Claire again.
Interviewee: Claire Hansell
In cryo-electron microscopy they’re flash freezing the samples, and then you just go in and take pictures of as many of them as you can find. And some of them will be lying flat on the grid, some of them will be rotated slightly relevant to each other, etc. You take enough of those that you can be sure you’ve covered all the angles, literally, of the particle and then reconstruct the 3D structure from that.
Interviewer: Noah Baker
Now, Uli was convinced that these were rings and that they were regular and symmetrical. Sure enough, he and his team confirmed that with cryo-electron microscopy, but that isn’t all they saw. Alongside the rings, there was something else.
Interviewee: Uli Wiesner
We looked at these rings and then we realised, hmm… they are not only rings in the soap if you like, but there were also other structures which seem to have more arms than just a ring.
Interviewer: Noah Baker
Uli and his team wanted to know what they were, and so they invested a bit of time.
Interviewee: Uli Wiesner
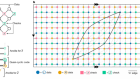
We manually identified and cut out about 20,000 cryo-electron microscopy images, which took us quite a long time, and then what emerged was actually a so-called dodecahedron.
Interviewer: Noah Baker
The dodecahedral cages each formed around a single surfactant template raised some tantalising possibilities. It could be conceived that ultra-small particles with an inside and an outside could carry and protect drugs within them, while simultaneously carrying binding molecules on their surface, helping target particular tissues. But, applications like this are a long way off, and we’ll have to wait and see if they ever even come to fruition. For now though, imaging the nanocages is a feat in itself.
Interviewee: Uli Wiesner
It’s a beautiful, highly symmetric cage structure, which we were absolutely elated about because we, you know, from all possible structures that could have been in there, that it turned out to be the dodecahedron was just a real, real, big surprise.
Host: Shamini Bundell
That was Uli Wiesner from Cornell University in the United States, and before that you heard from Nature editor Claire Hansell. And if you’re just aching to catch a glimpse of these teensy dodecahedra for yourself, head over to the Nature video channel. There is a very enlightening vid there which should tickle your fancy.
Interviewer: Benjamin Thompson
Whether we’re aware of them or not, artificial intelligences play a big part in our lives. Many of these systems rely on a branch of AI called machine learning, which as its name suggests, allows computers to learn, often from large data sets. These systems and the algorithms that underpin them are used all over the place, including in the public sector as Rhema Vaithianathan, co-director of the Centre for Social Data Analytics at Auckland University of Technology in New Zealand, explains.
Interviewee: Rhema Vaithianathan
So societally, I think they’re being used in a huge number of places in criminal justice, in education, in hospitals to try to figure out how to give services, how to make clinical decisions, you know, how to identify that people might become unemployed, so it’s being used in a lot of areas.
Interviewer: Benjamin Thompson
While machines learning algorithms potentially offer many benefits, there are also concerns. If machines are making decisions that could hugely affect someone’s life, how can we be sure that those decisions aren’t biased or prejudiced? This week, Nature has a Feature article looking algorithm accountability. But how could bias get introduced to an algorithm-based system in the first place?
Interviewee: Rhema Vaithianathan
I mean, often it’s, in my use cases it’s there because of the data that we’re bringing into the algorithms. Often those data come out of systems which themselves might have had human bias introduced. And so, the algorithm could correct for those biases, but they could also exacerbate those.
Interviewer: Benjamin Thompson
And this is one of the ways that bias might be introduced. A machine learning algorithm can be trained by showing it many previous examples. So, any existing bias found in the data it learns from could be repeated or amplified. Unfortunately, countering this is tricky, as a lot of these algorithms are part of closed proprietary systems, so knowing how any biases have come about can be difficult. But a closed system isn’t necessarily the only way. Back in August 2016, Allegheny County, in the US state of Pennsylvania launched the algorithm-based Allegheny Family Screening Tool, designed to help call-centre staff assess whether a child is at risk of abuse. Rhema led the team hired to develop it.
Interviewee: Rhema Vaithianathan
So, the algorithm is simply a predictive analytic tool that grabs all the relevant data systems. It has been trained on an outcome which is whether the child is going to be removed from home in the subsequent two years, and it offers to that person at that call-centre, a number from 1 to 20, where 20 says this child is in the 5% most likely to end up with being removed from home in the two years following the call, and 1 means they’re in the lowest 5% likelihood.
Interviewer: Benjamin Thompson
Unlike many algorithm-based systems, the Allegheny Family Screening Tool was designed from the outset to be open. Meetings were held with researchers, officials, and the local community to talk about the system and how it might affect them. It’s also being independently assessed and the system is open to public scrutiny.
Interviewee: Rhema Vaithianathan
We’re very passionate about transparency in this type of use, because I think we don’t actually know what the right answer is often, like what is the fairest algorithm, what does fairness mean? But those are all decisions that really needed a kind of a conversation with the community and the agency and the researchers, and only transparency can give you that. I think when you’re in these sorts of situations where you really have a diversity of opinions and all of them are just as valid in terms of how comfortable people are, how fair they think they are, what they mean by fairness, it’s really important for that to come out through a community conversation.
Interviewer: Benjamin Thompson
Defining what fairness actually is, is very complex, so being able to have these conversations, and knowing what an algorithm is doing is important. But what about closed systems, where algorithms aren’t open to scrutiny? Here’s Solon Barocas from Cornell University in the US.
Interviewee: Solon Barocas
There’s a fairly active community of researchers who are trying to develop methods to audit these kinds of systems from the outside. Of course, the challenge with that is that not all such systems have a public-facing component where you can submit this kind of information to see what the output looks like. So, a really good example of this is like we might want to be able to audit from the outside one of these credit-scoring models. But it’s not really possible to submit fake information to creditors to then see how they would respond to these different kinds of applicants. In contrast, you know, trying to audit the way that online advertising works is a lot more tractable because you can develop these sort of fake profiles online and then see how online advertisers treat those users.
Interviewer: Benjamin Thompson
Others researchers are taking different approaches, releasing open-source software or starting companies that offer algorithm auditing services. Facebook and Microsoft both announced this year that they’ll be developing tools to help detect bias as well. And it’s not just researchers – politicians are beginning to take notice. For example, the UK Parliament’s Science and Technology Committee looked at algorithm bias in a recent report and the French President Emmanuel Macron spoke about the issue when announcing a national AI strategy. However, there’s still a way to go. Algorithm-based systems are being used all over the place today, with little or no way to test whether they have any biases. So, what needs to be done as new tools are developed to make sure they’re as fair as possible? Solon has some ideas.
Interviewee: Solon Barocas
So, I think, you know, really one is to recognise that there’s no neutral way to learn from historical data, and the people who are developing these tools need to be extremely thoughtful and sensitive to what exactly is being recorded in the data that they’re using to train these systems. And increasingly, I think that the community is aware of that. What to do about these problems is going to be, I think, an ongoing debate and the goal here should be to have a debate in a more accessible way that includes a broader range of stakeholders. At the same time, I think there needs to be more work to kind of situate what this is doing and a broader debate around sort of you know what is a just institution, what is the appropriate way to think about fair employment or a fair process in the criminal justice system. So, my answer is like there’s a role here for these systems to be made more fair and themselves to make these institutions themselves more fair. But we need to keep an eye on what role they play in a larger push to kind of reform these fundamental institutions in society.
Interviewer: Benjamin Thompson
That was Solom Barocas. You also heard from Rhema Vaithianathan. You can find the Feature about algorithm fairness over at nature.com/news.
Host: Shamini Bundell
Later in the show we’ve got the News Chat, where we’ll be discussing the potential discovery of a host of new human genes. But before that, Adam Levy is here with this week’s Research Highlights.
[Jingle]
Interviewer: Adam Levy
Physicists have peered into the core of a super heavy element. Nobelium’s nucleus has a whopping 102 protons and is extremely unstable, making it hard to study. But researchers have now used lasers to catch a glimpse of nobelium nuclei. They studied how often the beam knocked electrons from the atoms at different light frequencies. This allowed the team not only to calculate the nuclear radius, but also its shape. They found that nobelium’s nucleus is shaped more like a rugby ball than the spherical nuclei of lighter elements. Peep inside that paper in Physical Review Letters.
[Jingle]
Interviewer: Adam Levy
If you’ve ever stuck a finger in the air to test the wind, then you’ve got something in common with crab spiders. Many tiny spiders catch a lift with the wind by sending out special silk threads. But scientists have been puzzled by how relatively large spiders, some up to 6 millimetres long, manage this impressive feat called ‘ballooning’. So, researchers watched how crab spiders took off. They saw that the spiders raised a front leg in the air to check if wind speeds and temperature were suitable for ballooning. The spiders then spun several specialised silk threads up to 4 metres long in a triangular sheet. Find out more in PLoS Biology.
[Jingle]
Host: Shamini Bundell
Next up, reporter Geoff Marsh has been looking into a Nature paper about weight loss in pancreatic cancer.
Interviewer: Geoff Marsh
Weight loss and tissue wasting are early warning signs of several different cancers. In pancreatic cancer, this wasting can come several months before the actual diagnosis. But it’s not just an early warning sign. This muscle and fat wasting phenomenon has long been thought to reduce a patient’s life expectancy. Keeping patients in good shape is key to helping the body withstand the effects of chemotherapy and surgery, for example. A study out this week has now used a mouse model of pancreatic cancer to shed light on the functions that go awry in this disease. As is often the case with science, the findings have raised more questions than they’ve answered. For a bit of context, I first called Matthias Löhr from the Karolinska Institutet in Sweden. His research specialises in the function and cancers of the pancreas. He didn’t work on this study but has written an accompanying News and Views article.
Interviewee: Matthias Löhr
First of all, weight loss is an early sign in several cancers, even prior to the diagnosis of a cancer. And there’s something also special with pancreatic cancer, at least the way we looked at it in the past, in that the gland is producing a juice, 1.5 litres on average per day, and then it’s squeezing out this liquid together with the bile and help digest the food. Most of the pancreatic cancer is actually located in the head of the pancreas and are then obstructing the duct, which is leading then to weight loss and tissue loss because you cannot digest food the way you could have done it without the tumour.
Interviewer: Geoff Marsh
So that was one of the theories for how pancreatic cancer causes this fat and muscle wasting. But the matter was by no means settled. To better understand the mechanism behind this wasting phenomenon, I spoke to Laura Danai, first author on the current study.
Interviewee: Laura Danai
So, in this paper we were looking at this fat and muscle wasting that happens with pancreatic cancer. And we wanted to ask whether pancreatic cancer cells themselves are causing this effect, or if it’s something about them growing in the pancreas. And so, what we did is that we took pancreatic cancer cells and wild-type ‘normal’ mice, and we injected these pancreatic cancer cells either under the skin, or we perform a surgery where we exposed the pancreas and injected the cancer cells into the pancreas. We did various iterations of this experiment, but what we found was that the cells that are in the pancreas promote this wasting phenomenon, whereas the cells that were under the skin did not. So, this has suggested that it’s not something that the pancreatic cancer cells do themselves, but rather that it’s something about growing in the pancreas that’s causing this effect.
Interviewer: Geoff Marsh
So, what does this tell you then? When you had the tumour in the pancreas but there wasn’t necessarily an obstructed duct, it still led to muscle and fat wasting.
Interviewee: Laura Danai
We do know that the exocrine part of the pancreas, the part that helps you break down the food, is somehow involved. And we know that from the diet experiments that we did, where we had mice with pancreatic cancer and either fed them a controlled diet, or a diet that was supplemented with these pancreatic enzymes. And when the mice had this diet that was supplemented with pancreatic enzymes, they had less of the fat wasting.
Interviewer: Geoff Marsh
But then it didn’t actually play out very well for the mice themselves, did it?
Interviewee: Laura Danai
No. So, if you are improving this wasting effect that we see with pancreatic cancer, you would expect that you would also improve survival. And what we actually found was that the mice that had this diet that was supplemented with pancreatic enzymes and they had less wasting, actually had lower survival or died faster.
Interviewer: Geoff Marsh
Matthias and Laura both told me that this is counterintuitive, given that doctors caring for patients with pancreatic cancer, the world over, supplement pancreatic enzymes to patients as a standard of care. This kind of tissue wasting is widely believed to reduce life expectancy, so the team decided to look at some clinical data for humans.
Interviewee: Laura Danai
What ended up happening is that as we were getting the mouse data together, we talked to the collaborators and they had been gathering data for humans for quite a while. They actually had expected to see that the more wasting, the worse survival, and they did not observe that. And so, this was kind of, we both had independently kind of come up with this data and so we were able to join forces.
Interviewer: Geoff Marsh
What do you think that this paper suggests about whether or not doctors should be trying to reverse the muscle and fat wasting in cancer patients with these supplements?
Interviewee: Laura Danai
So, assuming you could translate mouse studies to human studies, I would say that it may not be as beneficial to try to improve the wasting, because it may lead to worse survival and that wasting is maybe the body’s own way to kind of contain the cancer.
Interviewer: Geoff Marsh
Laura was careful to point out that this is still speculation and that she’s a scientist, not a doctor. So, I posed the same question to Matthias Löhr, who we heard from earlier, for his perspective as a qualified doctor who works with these patients.
Interviewee: Matthias Löhr
This is a very critical question because for all we know the cancer patient should be in the best possible physical shape. That is to say for the time being, we would of course try to really get every patient the best nutritional support and try to avoid the tissue wasting, because we know this is also linked to ability to withstand, or tolerate rather, chemotherapy, for instance. Having said this, conversely, if you have tissue wasting that does not automatically mean that you will die earlier than those who don’t have tissue wasting, which maybe should lead us to be a little bit more relaxed on the nutritional status of these patients.
Host: Shamini Bundell
That was Matthias Löhr talking with Geoff Marsh. You also heard from Laura Danai. You could read her paper and Matthias’ News and Views article over at nature.com/nature.
Interviewer: Benjamin Thompson
Right then listeners, finally in this week’s show its time for the News Chat and I’m joined here in the studio by Nisha Gaind, one of the News Editors here at Nature. Hi Nisha.
Interviewee: Nisha Gaind
Hi Ben.
Interviewer: Benjamin Thompson
Okay then for our first story today we’re going to be looking at genes and this is a story that started at the turn of the millennium.
Interviewee: Nisha Gaind
Yeah, this goes back to the year 2000 when researchers were still trying to figure out how many genes were in the human genome and estimates ranged from the tens of thousands to the hundreds of thousands. Almost two decades later, scientists still don’t really have an answer to that question despite the fact that they have much more data and they say it’s a knowledge gap that hampers efforts to spot disease-related mutations in genes.
Interviewer: Benjamin Thompson
Alright then Nisha, what’s happened this week then to try and get a more accurate number of genes then?
Interviewee: Nisha Gaind
So, we’ve got a new study published that is one of the latest attempts to try and plug this gap and it uses data from hundreds of human tissue samples, and it found almost 5,000 genes that haven’t previously been spotted. About 1,200 of those 5,000 new genes carry instructions for making proteins, so that takes the tally of protein coding genes to more than 21,000.
Interviewer: Benjamin Thompson
And who’s done this research?
Interviewee: Nisha Gaind
The study was posted on the bioRxiv preprints server and was led by Steven Salzberg who’s a computational biologist at Johns Hopkins University in Baltimore. And Salzberg’s team used data from Genotype-Tissue Expression Project or GTEx.
Interviewer: Benjamin Thompson
Oh, okay, well listeners we have covered GTEx in the podcast before so I’d have a look out for that. But Nisha, in this study what have they been looking for?
Interviewee: Nisha Gaind
The researchers wanted to identify genes that encode proteins and those that don’t, but still serve an important role in cells. And what they found was 21,306 protein-coding genes and 21,856 non-coding genes.
Interviewer: Benjamin Thompson
So how exactly does this compare to numbers that other teams have found before?
Interviewee: Nisha Gaind
So, what’s interesting about that finding is that those number of genes is many more than are including in even the two most widely used databases of human genes. For instance, GENCODE which is a European-run database has about 20,000 protein-coding genes and 16,000 non-coding genes. Another one called RefSeq which is a US-run database again has about 20,000 protein-coding genes but 18,000 non-coding genes.
Interviewer: Benjamin Thompson
So, a fair discrepancy then but do we know any reasons as to why this is?
Interviewee: Nisha Gaind
Yes, there are some suggested reasons. For instance Kim Pruitt who’s a genome researcher in the US says that the difference is probably in part due to the fact that Salzberg’s team analysed a much larger volume of data. And one other difference between the GENCODE and the RefSeq databases and the latest count is that those two databses rely on manual curation and that means that a person has to review evidence for each gene and then make a final determination. By contrast, Salzberg’s group relied solely on a computer programme to sift the data.
Interviewer: Benjamin Thompson
So, what still needs to be done then?
Interviewee: Nisha Gaind
So, these sorts of findings need to be validated by other independent teams but there are a few confounding factors that make this problem difficult in the first place. One of them is the fact that the definition of what a gene is itself is imprecise and changing. Biologists used to see genes as sequences that code for proteins, but then it became clear that some non-coding molecules have important roles in cells as well. So, judging which are important and which should be deemed genes is controversial and that could also explain some of the discrepancies in this latest work.
Interviewer: Benjamin Thompson
Right then Nisha, let’s move on to our second story and it concerns the World Cup which, of course, is a rather large international football or soccer tournament that’s going on right now in Russia. The eyes of the world are on it, but maybe not everybody is delighted about it.
Interviewee: Nisha Gaind
That’s right, we’ve got an interesting story out of Russia about how some science is being affected by some government decrees that have been put in place because of the World Cup.
Interviewer: Benjamin Thompson
And what decrees are they then?
Interviewee: Nisha Gaind
So, because of this international spectacle there are several security and counter-terror measures that have been enacted by the Russian government. And one of those is affecting the way that Russian labs can procure radioactive reagents that they urgently need for research, and that affects some types of molecular biology and biochemistry.
Interviewer: Benjamin Thompson
Right, so inadvertently then they’ve been caught up in this new policy.
Interviewee: Nisha Gaind
That’s right, it’s a temporary policy that has suspended the sale and transport of hazardous, chemical and biological substances for two months. Now, the decree only applies to cities that are actually hosting matches but many of those including Moscow are also big research hubs.
Interviewer: Benjamin Thompson
And what are the researchers being affected saying?
Interviewee: Nisha Gaind
Well they say that ordering these sorts of research materials can be difficult in the summer anyway, and this World Cup is making the situation worse. At one lab because of the World Cup and the summer break it means that the deliveries of these essential agents might not come until the Autumn and that means bad disruption including disrupting PhD students who are right in the middle of their thesis work.
Interviewer: Benjamin Thompson
Well let’s hope that resolved quickly then after the tournament. Nisha, thank you so much for joining us and listeners, for more on the latest science news don’t forget to head over to nature.com/news.
Host: Shamini Bundell
Well, that’s it for this week’s show but don’t forget we’ve got a video on Noah’s story from earlier about tiny self-assembling silica structures so you can see them for yourself. That’s at youtube.com/NatureVideoChannel. I’m Shamini Bundell.
Host: Benjamin Thompson
And I’m Benjamin Thompson. Thanks for listening everyone, see you all next time.
[Jingle]

 Read the paper: Altered exocrine function can drive adipose wasting in early pancreatic cancer
Read the paper: Altered exocrine function can drive adipose wasting in early pancreatic cancer
 New human gene tally reignites debate
New human gene tally reignites debate
 Nature Podcast: A new episode every week
Nature Podcast: A new episode every week