- TECHNOLOGY FEATURE
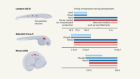
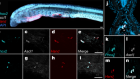
Stem-cell culture moves to the third dimension
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 558, 329-331 (2018)
doi: https://doi.org/10.1038/d41586-018-05380-x
References
Sato, T. et al. Nature 459, 262–265 (2009).
van de Wetering, M. et al. Cell 161, 933–945 (2015).
Wang, G. et al. Nature Med. 20, 616–623 (2014).
Huh, D. et al. Science 328, 1662–1668 (2010).
Miller, P. G. & Shuler, M. L. Biotechnol. Bioeng. 113, 2213–2227 (2016).
Edington, C. D. et al. Sci. Rep. 8, 4530 (2018).
Benam, K. H. et al. Cell Syst. 3, 456–466 (2016).

 ‘Organs-on-chips’ go mainstream
‘Organs-on-chips’ go mainstream
 Q&A: Hans Clevers
Q&A: Hans Clevers
 The boom in mini stomachs, brains, breasts, kidneys and more
The boom in mini stomachs, brains, breasts, kidneys and more