- NEWS FEATURE
The lost art of looking at plants

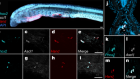
3D imaging offers new views of these antirrhinum (snapdragon) buds. Credit: Karen Lee/Xana Rebocho/John Innes Centre
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 553, 396-398 (2018)
doi: https://doi.org/10.1038/d41586-018-01075-5
References
Staedler, Y. M. et al. J. Exp. Bot. 69, 525–535 (2017).
Richardson, A., Rebocho, A. B. & Coen, E. Plant Cell 28, 2079–2096 (2016).
Dornbusch, T., Michaud, O., Xenarios, I. & Fankhauser, C. Plant Cell 26, 3911–3921 (2014).
Zhang, G-Q. et al. Nature 549, 379–383 (2017).
Vlad, D. et al. Science 343, 780–783 (2014).
Ranjan, A., Townsley, B. T., Ichihashi, Y., Sinha, N. R. & Chitwood, D. H. PLoS Genet. 11, e1004900 (2015).
Watanabe, K. et al. Sci. Rep. 7, 10028 (2017).
Kui, L. et al. Front. Plant. Sci. 7, 2036 (2017).