Abstract
Aggressive NK-cell leukemia (ANKL) is a rare form of NK cell neoplasm that is more prevalent among people from Asia and Central and South America. Patients usually die within days to months, even after receiving prompt therapeutic management. Here we performed the first comprehensive study of ANKL by integrating whole genome, transcriptome and targeted sequencing, cytokine array as well as functional assays. Mutations in the JAK-STAT pathway were identified in 48% (14/29) of ANKL patients, while the extracellular STAT3 stimulator IL10 was elevated by an average of 56-fold (P < 0.0001) in the plasma of all patients examined. Additional frequently mutated genes included TP53 (34%), TET2 (28%), CREBBP (21%) and MLL2 (21%). Patient NK leukemia cells showed prominent activation of STAT3 phosphorylation, MYC expression and transcriptional activities in multiple metabolic pathways. Functionally, STAT3 activation and MYC expression were critical for the proliferation and survival of ANKL cells. STAT signaling regulated the MYC transcription program, and both STAT signaling and MYC transcription were required to maintain the activation of nucleotide synthesis and glycolysis. Collectively, the JAK-STAT pathway represents a major target for genomic alterations and IL10 stimulation in ANKL. This newly discovered JAK/STAT-MYC-biosynthesis axis may provide opportunities for the development of novel therapeutic strategies in treating this subtype of leukemia.
Similar content being viewed by others
Introduction
Aggressive NK-cell leukemia (ANKL) results from the systemic neoplastic proliferation of natural killer (NK) cells. ANKL shows an aggressive clinical course with features that overlap with the advanced stage of natural killer/T-cell lymphoma (NKTCL). Unlike NKTCL, ANKL is a rare form of NK cell neoplasm and is more prevalent among people from Asia and Central and South America1. To date, less than 300 cases of ANKL have been described in the English literature worldwide.
Patients with ANKL manifest a fulminant clinical course, including high fever, pancytopenia, hepatosplenomegaly, multiple organ failure, cytokine storm and hemophagocytic lymphohistiocytosis, and usually die within days to months, even after receiving prompt therapeutic management1,2,3. Because of the rarity of the disease, the treatment strategy for ANKL has largely been extrapolated from other lymphoid neoplasms. However, the response to conventional chemotherapy routinely adopted in non-Hodgkin lymphoma and lymphoblastic leukemia is poor3,4. Thus, a better understanding of the pathogenesis of ANKL is critically needed to develop more effective therapies. The Epstein-Barr virus (EBV) sequence is nearly always detected in the neoplastic tissues and peripheral blood (PB) of ANKL patients, and is proposed to contribute to the pathogenesis of this disease1,2,3. Although chromosomal gains and losses were previously shown to be associated with ANKL5, no recurrent genetic aberrations in ANKL have been confirmed3. Here, we performed the first comprehensive genomic analysis of ANKL (Supplementary information, Figure S1). We identified and functionally validated the IL10-JAK/STAT-MYC signaling axis involved in the pathogenesis of ANKL. Further, the activation of this pathway led to significant metabolic reprogramming of ANKL in nucleotide synthesis and glycolysis. Finally, by elucidating the functional dependency of ANKL on the identified oncogenic pathway, we provide insights into the development of novel therapeutic strategies for treating this deadly type of leukemia.
Results
Genomic landscape of ANKL and alterations in the JAK-STAT pathway
To explore the genetic aberrations underpinning the pathogenesis of ANKL, whole-genome sequencing (WGS) was performed on purified CD56+CD3− NK cells and their paired normal granulocytes (Supplementary information, Figure S2) from a discovery cohort of 8 patients (Supplementary information, Table S1) with newly diagnosed ANKL (Figure 1 and Supplementary information, Tables S2, S3, S4). Using GVC-SNV in the GVC (Genomic Variant Caller) utility6, we identified 483 candidate somatic single-nucleotide variations (SNVs) in coding regions, including 311 non-silent SNVs (289 missense and 22 nonsense) and 9 splice-site mutations (Figure 1 and Supplementary information, Table S2). We identified a mean of 40 (range 16-59) somatic non-synonymous mutations per sample, which was comparable to NKTCL7 and multiple types of solid tumors8 and higher than diffuse large B-cell lymphoma (DLBCL)9 and acute myelogenous leukemia (AML)10 (Supplementary information, Figure S3). A high level of genomic EBV sequences was detected in leukemic NK cells but not in isolated patient granulocytes (Supplementary information, Figure S4).
Circos plots of 8 ANKL patients. Each Circos plot depicts the genomic location, depth distributions of normal and tumor cells (green track), somatic copy number gains (red track) and somatic copy number alterations (red, copy gain; blue, copy loss). Somatic non-synonymous mutations are marked by gene names (red points, mutation allele frequencies); somatic structural rearrangements are marked by orange lines.
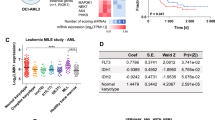
To predict the functional consequences of the ANKL genomic alterations, we looked for mutation-enriched pathway(s) using KEGG and IPA analysis as well as the HotNet algorithm, which constructs gene networks according to protein-protein interactions11. Remarkably, all three strategies consistently pointed to JAK-STAT cytokine signaling (including JAK2, JAK3, STAT3 and STAT5B) as the most significantly enriched pathway when analyzing 313 mutated genes discovered in the WGS of 8 ANKL patients (Figure 2A; Supplementary information, Figures S5 and S6). To validate this finding, we used AmpliSeq for targeted sequencing in a cohort of 29 ANKL patients (Supplementary information, Data S1). In addition to major components of the JAK-STAT pathway, we also included known oncogenes and tumor suppressors mutated in the discovery cohort as well as those frequently mutated in NKTCL (Figure 2B). Mutations in the JAK-STAT pathway, namely STAT3, STAT5B, STAT5A, JAK2, JAK3, STAT6, SOCS1, SOCS3 and PTPN11, were mutually exclusive and collectively recurrent (48%) in the combined discovery and validation cohort (n = 29). Additional frequently mutated genes included TP53 (34%), TET2 (28%), CREBBP (21%) and MLL2 (21%) (Figure 2B; Supplementary information, Figures S7, S8 and Table S5). Notably, we found that genes frequently mutated in NKTCL, such as DDX3X (20%) and BCOR (32%)7,12, were mutated in ANKL patients with a lower frequency (7% and 3%, respectively; Figure 2B and Supplementary information, Table S6). In addition, we observed that other commonly mutated genes in NKTCL (namely MLL2, ARID1A, EP300, ASXL3 and MGA) were also altered in ANKL. Interestingly, these genes were preferentially found in patients harboring mutations in the JAK/STAT pathway, but not those carrying TET2 mutations in ANKL. Together, these results showed that a similar set of genes related to the JAK/STAT pathway and epigenetic regulation were mutated in both ANKL and NKTCL. Although the observed mutational pattern of ANKL was different from that of NKTCL (Figure 2B and Supplementary information, Table S6)7,12, it cannot be ascertained whether such differences are related to the nature of the neoplasms or only reflecting the variations among different cohorts. Moreover, the majority (7 out of 8) of the identified STAT3 and STAT5B mutations are located in the Src homology 2 (SH2) domain (Supplementary information, Figure S7), which mediates STAT protein dimerization and was reported to be the hot spot containing activating mutations in T-cell large granular lymphocytic leukemia (T-LGL)13,14, NKTCL7,15, T-cell acute lymphoblastic leukemia (T-ALL)16, gamma delta T-cell lymphoma (γδTCL)15, T cell prolymphocytic leukemia (T-PLL)17 and inflammatory hepatocellular adenomas (IHCAs)18. Two cases carried the previously characterized activating mutation Y640F, which results in constitutively phosphorylated and transcriptionally active STAT3 protein in functional analyses14,18. In addition, an activating mutation located in STAT5 SH2 domain has recently been found in one ANKL patient by Sanger sequencing19. Although JAK2 chromosomal translocation was reported in ALL20, no rearrangements involving JAK-STAT genes were identified in ANKL.
Mutations in the JAK-STAT pathway, elevated IL10 level and activation of STAT3 in ANKL patients. (A) Enriched canonical pathways analyzed by Ingenuity Pathway Analysis (IPA) (P-value ≤ 0.01) of the 313 genes with somatic mutations detected in the WGS data. The orange dot represents the ratio value of the number of mutated genes over the total number of genes in the pathway. (B) Gene mutations identified by WGS in 8 subjects and targeted sequencing in an additional 21 subjects with ANKL. The percentage of subjects with mutations is listed on the right. The 34 genes examined are classified into the categories indicated on the left. Fold changes in the plasma IL10 levels relative to healthy donors for the indicated 6 cases (Figure 2C) are also shown. (C) The relative plasma levels of 20 inflammatory cytokines in 8 ANKL patients and 3 healthy donors. Statistical analysis was performed using Student's t-test or Student's t-test with Welch's correction. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001. The correlation between the levels of plasma IL10 protein and IL10 mRNA in sorted NK leukemia cells is shown in the inset. (D) The immunohistochemical staining of CD56 and phosphorylated STAT3 (pSTAT3) in bone marrow biopsy specimens. Representative images of samples from controls, JAK-STAT mutant ANKL cases and JAK-STAT wild-type ANKL cases are shown (left). Scale bar, 20 μm. The semi-quantitative immunoreactivity histological scores (H scores) of pSTAT3 staining for 10 ANKL and 21 control samples are shown and compared (P < 0.0001; right). JAK-STAT mutant (red), wild-type (blue) and mutation status unknown (black) cases are indicated. Bars represent the mean ± SD. Statistical analysis was performed using the Mann-Whitney U-test.
In addition to genomic mutations, extracellular ligand binding of cytokines and growth factors can lead to constitutive activation of JAK-STAT signaling. The overwhelming production of cytokines, referred to as the “cytokine storm”, is a prominent clinical feature in ANKL patients. Intrigued by the potential functional involvement of the cytokine storm in JAK-STAT signaling, we screened a panel of 20 inflammatory cytokines for their presence in patient plasma. The known JAK-STAT upstream stimulator IL1021 was dramatically elevated by an average of 56-fold (P < 0.0001, ranging from 7- to 98-fold) in ANKL patients compared to healthy controls (Figure 2C). Consistently, CD56-positive NK cells were abundant in the bone marrow of ANKL patients as shown by immunohistochemistry analysis (Figure 2D). Phosphorylation of STAT3, indicative of STAT3 activation, was significantly higher in the bone marrow cells of both JAK-STAT wild-type and mutated ANKL cases compared to controls (Figure 2D and Supplementary information, Table S7).
Deregulated transcriptome and aberrations in the MYC expression program and metabolic pathways
To identify key transcriptional hubs whose expression levels are regulated by the identified genetic mutations and/or remarkably elevated IL10, transcriptomic sequencing was performed on flow cytometry-sorted malignant or normal CD56+CD3− NK cells (Supplementary information, Figure S2) from the bone marrow of 7 ANKL patients (Supplementary information, Table S1) and 4 healthy donors, respectively. We used an integrative analysis tool, TieDIE (Tied Diffusion through Interacting Events), to search for interlinking genes connecting genomic and transcriptomic data22. Notably, this unbiased genome-wide analysis constructed a network in which the increased expression of the transcription factor MYC in ANKL is under the control of mutations in the JAK and STAT genes (Figure 3A). In support of this finding, we identified 7 activated regulators, including MYC and its interacting proteins MAX, MNT and SIN3A, along with SATB1, HOXD10 and HIC1 (Z-score > 2, P < 0.05) in the entire deregulated ANKL transcriptome, using IPA analysis of upstream regulators (Figure 3B). Consistently, the expression level of MYC was significantly higher in ANKL patients compared to healthy controls (Figures 2D and 3F; Supplementary information, Figure S9). The ANKL patient transcriptome was characterized by two known MYC-driven expression signatures23,24, but lacked the reported STAT3-associated inflammatory gene signature21 (Figure 3C).
JAK/STAT-MYC and metabolic signatures in ANKL patients. (A) Integrated network analysis of genomic alterations and transcriptomic variations with TieDIE. The figure shows four subnetworks predicted by TieDIE22. The largest subnetwork (upper left, shaded area) connects JAK and STAT mutations to the transcriptionally upregulated gene MYC. Diamonds indicate somatic SNVs detected using WGS data. The colors in the diamonds and circles indicate increased (red) or decreased (blue) expression of the gene relative to healthy donors. Solid lines indicate transcriptional regulation, and dotted lines indicate post-transcriptional regulation. (B) Seven transcriptional regulators (diamonds) were predicted to be activated in isolated patient NK leukemia cells (Supplementary information, Data S1). Their potential downstream targets are shown as circles. The colors in the diamonds and circles indicate increased (red) or decreased (blue) expression of the gene relative to healthy donors. The red and blue lines represent known activating or inhibitory effects, respectively, between the regulator and its targets. Black lines indicate unknown effects. (C) The graph shows GSEA of previously characterized MYC and STAT3 inflammatory signatures in primary ANKL leukemia cells. P-values were calculated by permutation test with a threshold of P < 0.05. (D) The graph shows the percentage of enriched pathways for each of the six indicated KEGG functional categories in ANKL and various tumors (P < 0.05). Differentially upregulated genes in comparison to normal controls were used for the enrichment analysis. The transcriptome data were from this study (ANKL) or retrieved from public data (Supplementary information, Data S1). (E) The graph shows the enriched metabolic pathways that revealed increased (red) or decreased (blue) transcriptional activities in ANKL, a MYC-inducible cell line (P493) and additional tumors with known metabolic features (P < 0.01). The pathways are classified into the categories indicated at the left. The color code indicates the level of significance. Lists of the enriched metabolic pathways are provided in Supplementary information, Table S12. (F) The bar plot (top) shows the mRNA expression level of MYC in ANKL patients and healthy donors. Asterisks indicate the activation status of MYC based on IPA analysis (P < 0.05, Z-score ≥ 2). The heat map (middle) shows the mRNA expression levels of nucleotide synthesis- and glycolysis-related genes that were highly expressed in ANKL patients (FDR < 0.1). MYC target genes were displayed in red font. The scale bar represents the level of gene expression. The MYC CNV, point mutations in the JAK-STAT pathway (JAK-STAT SNV) and fold change of IL10 expression are shown at the bottom of the graph. One case was found to contain amplification of the MYC locus (Supplementary information, Figure S21). BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CNV, copy number variation; COAD, colon adenocarcinoma; DLBCL_OxPhos, diffuse large B-cell lymphoma with oxidative phosphorylation signature; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; NES, normalized enrichment score; PRAD, prostate adenocarcinoma; T-ALL, T-cell acute lymphoblastic leukemia; THCA, thyroid carcinoma.
Consistent with the role of MYC as a master regulator in cellular metabolism25,26,27, analysis of the MYC signature genes that were upregulated in ANKL showed functional enrichment in the nucleotide synthesis, protein metabolism and cellular compound metabolic process pathways (Supplementary information, Figure S10). To explore whether metabolic aberration is a common feature in the entire transcriptome, we analyzed differentially transcribed genes. We identified 778 upregulated and 1 788 downregulated genes in isolated patient NK leukemic cells compared to normal controls (|Fold change| > 2 and FDR < 0.1) (Supplementary information, Tables S8 and S9). Surprisingly, the majority (13 out of 20, 65%) of the pathways enriched in upregulated genes involved cellular metabolism (P < 0.05; Figure 3D and Supplementary information, Table S10), including purine and pyrimidine synthesis, glycolysis, the tricarboxylic acid (TCA) cycle, oxidative phosphorylation, amino acid metabolism and lipid metabolism (P < 0.01; Figure 3E and Supplementary information, Table S12). A similar set of metabolic pathways were activated upon MYC induction in a well-characterized MYC-inducible system (Figure 3E and Supplementary information, Table S12)28. In contrast, the same analyses of transcriptome data could only detect a limited number of upregulated metabolic pathways in T-ALL (glycolysis, nucleotide synthesis and amino acid metabolism), lung cancer (LUAD: glycolysis, nucleotide synthesis and amino acid metabolism), kidney cancer (KIRC: glycolysis and nucleotide synthesis) and OxPhos-DLBCL (oxidative phosphorylation and amino acid metabolism; Figure 3E and Supplementary information, Table S12), which are largely consistent with previous characterizations of these tumors using isotope-tracing and functional assays29,30,31,32. Furthermore, signal transduction (including NOTCH signaling), specific cancer-related pathways and immune function-related processes all comprised downregulated genes in ANKLs (Supplementary information, Table S11). Interestingly, aberrant expression signatures in cellular metabolism, such as nucleotide synthesis and glycolysis pathways, showed similar patterns across JAK-STAT-mutated and wild-type samples (Figure 3F). Irrespective of the JAK-STAT mutation status, significantly higher levels of IL10 and MYC were observed in all of the 7 cases examined (Figure 3F). These data suggest that metabolic aberration in nucleotide synthesis, glycolysis and additional pathways is a prominent feature of the ANKL transcriptome.
Functional analysis of STAT3 signaling, and MYC expression and metabolic aberration in nucleotide synthesis and glycolysis
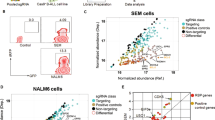
To determine the functional relevance of our integrative genomic analysis, we used two ANKL patient-derived cell lines, KHYG-1 and NK-9233,34, which lacked JAK-STAT mutations, as indicated by our exon sequencing analysis (data not shown). Functionally, IL10 could stimulate STAT3 phosphorylation (Figure 4A and Supplementary information, Figure S11A) and significantly promote the in vitro proliferation of these cells (Figure 4B). Conversely, inhibition of STAT3 signaling by Stattic, a STAT3 phosphorylation inhibitor35, significantly decreased cell viability and triggered apoptosis in ANKL cells (Figure 4C and 4D; Supplementary information, Figure S11B and S11C). These findings suggested that ANKL cells are dependent on STAT3 signaling for survival and are strongly responsive to IL10 for further induction of STAT3 phosphorylation and enhanced proliferation.
The STAT3-MYC axis is critical for the proliferation and survival of ANKL cells. Western blotting was performed to determine the protein expression levels of STAT3, phosphorylated STAT3 (pSTAT3) and MYC in KHYG-1 cells treated with the indicated concentrations of IL10 (A) or the STAT3 inhibitor Stattic (C). (B) Relative cell viability of IL10- or IL18-treated KHYG-1 or NK-92 cells. The data represent the mean ± SD. Statistical analysis was performed using Student's t-test. *P < 0.05, **P < 0.01 and ***P < 0.001. (D) Relative cell viabilities of Stattic-treated KHYG-1 or NK-92 cells. The data are shown as the mean ± SD. (E) The colony numbers of KHYG-1 cells treated with DMSO as a control (CON) or with the indicated concentrations of JQ1 (left). The data represent the mean ± SD. Statistical analysis was performed using Student's t-test; **P < 0.01 and ***P < 0.001. Representative images of the colonies are shown on the right. NS, not significant.
Consistent with the genomic analysis connecting the JAK-STAT pathway and MYC expression (Figure 3A), activation of STAT3 signaling by IL10 led to the significantly increased expression of MYC in KHYG-1 and NK-92 ANKL cells (Figure 4A; Supplementary information, Figures S11A and S12A). Inhibition of STAT3 phosphorylation downregulated MYC expression (Figure 4C; Supplementary information, Figures S11B and S12B). Furthermore, treatment with the known MYC inhibitor JQ1 effectively inhibited the proliferation and colony-forming capacities of ANKL cells (Figure 4E and Supplementary information, Figure S13). These data demonstrate that IL10-STAT3 signaling regulates MYC expression and the MYC-associated transcription program in ANKL.
To explore whether STAT3 signaling-driven MYC expression regulates downstream expression program and the observed metabolic activation in ANKL, we carried out transcriptome sequencing on two ANKL cell lines. While the reported STAT3-associated inflammatory expression program21 was not affected, the two known MYC-driven signatures23,24 were significantly induced or suppressed by IL10 or Stattic, respectively, in both ANKL cell lines (Figure 5A and Supplementary information, Figure S14). Among the 6 activated metabolic pathways in primary ANKL cells (Figure 5B and Supplementary information, Figure S15), only transcriptional activities in nucleotide synthesis and glycolysis positively and consistently responded to the modulation of STAT3 activity by IL10 and Stattic in KHYG-1 and NK-92 cells (Figure 5B and Supplementary information, Figure S15). Treatment with the MYC inhibitor JQ1, similar to the STAT3 inhibitor Stattic, resulted in the suppression of nucleotide synthesis and glycolysis in ANKL cells (Figure 5B and Supplementary information, Figure S15).
The STAT3-MYC axis is responsible for the metabolic activation in ANKL. (A) GSEA of previously characterized MYC and STAT3 inflammatory signatures in an ANKL cell line (KHYG-1) treated with IL10, Stattic or JQ1. P < 0.05 by permutation test. (B) Enriched metabolic pathways that had increased (red) or decreased (blue) transcriptional activity in isolated NK leukemia cells (ANKL patient NK cells, PNKC) and KHYG-1 (1) and NK-92 (2) cells treated with the indicated agents. Only pathways with contrasting changes between JAK-STAT-activating (IL10) and JAK-STAT-repressing (Stattic) conditions are shown (Supplementary information, Data S1). The color code indicates the level of significance. A complete list of the enriched pathways is provided in Supplementary information, Figure S15. (C) Integrative analysis of transcriptional and metabolic data revealed emerging drug targets in patient with ANKL. The graph shows the purine (left) and pyrimidine metabolism pathway (right). Upregulated genes (FDR < 0.1) in ANKL patients or in ANKL cell lines (KHYG-1 and NK-92) treated with IL10 were marked in red color. The inverted triangles indicate inhibition by Stattic or JQ1 (light blue). GSEA, gene-set enrichment analysis; NES, normalized enrichment score.
Discussion
By integrating genomic, transcriptomic and metabolic analyses as well as functional assays, we showed that the overproduction of plasma IL10 together with mutations in the JAK-STAT pathway are the primary factors fueling cellular metabolism in ANKL leukemia cells (Figure 6). Identified mutations such as STAT3 Y640F could stimulate STAT3 phosphorylation and downstream MYC activation (Supplementary information, Figure S16). Importantly, aberrant STAT3 signaling was also observed in JAK-STAT mutation-negative cases (Figure 2D). JAK-STAT-mutated and non-mutated patient NK leukemia cells showed a similar expression pattern in MYC-driven programs and metabolic activation (Figure 3F). These data suggest that independent of genetic mutations, STAT3 signaling can be activated by overproduced IL10 or additional unknown mechanisms. Interestingly, IL10 treatment preferentially stimulated the JAK/STAT wild-type cells (Figure 4A and Supplementary information, Figure S17), but not the STAT3 Y640F-mutant ANKL cells to increase pathway activity and cellular proliferation (Supplementary information, Figure S18), suggesting that IL10 upregulation plays a more prominent role in the JAK-STAT non-mutated patients. Mechanistically, the prevalent overexpression of IL10 is likely induced by EBER136,37, an EBV-encoded RNA that is highly expressed in patient NK leukemia cells (Supplementary information, Figures S19 and S20), and a positive feedback due to the STAT pathway activation21. Consistent with our findings, activation of the JAK-STAT pathway can occur without mutations in the JAK and STAT genes in a subset of T-LGL patients13. Together, targeting STAT3 signaling and nucleotide metabolism may offer new treatment strategies for both JAK-STAT wild-type and mutated ANKL patients.
A convergent model for the pathogenesis of ANKL. IL10 stimulates STAT3 signaling in both JAK-STAT-mutated and wild-type patients. The JAK-STAT pathway, EBV-derived RNA and additional mechanisms (not shown) contribute to the high level of plasma IL10 expression. Activated STAT3 signaling leads to the MYC-driven transcription program that sustains aberrant activation of nucleotide synthesis and glycolysis. Mutations in JAK-STAT pathway or TET2 genes cooperate with the JAK/STAT-MYC-biosynthesis axis and accelerate leukemogenesis in ANKL. Known inhibitors were tested to show the functional importance of specific target genes or pathways.
Additional mutated genes may also play important roles in the pathogenesis of ANKL, particularly for JAK-STAT wild-type patients. Interestingly, TET2 mutations were preferentially enriched in JAK-STAT non-mutated patients (3 TET2 mutated cases in JAK-STAT mutation-positive group vs 7 TET2 mutated cases in JAK-STAT mutation-negative group). All three reported loss-of-function TET2 mutations were in the JAK-STAT mutation-negative group (in a combined cohort of 39 patients, data not shown)38,39. Functional cooperation between STAT signaling and loss of TET2 has been recently reported in several studies of hematological malignancies40,41,42. In combination with STAT pathway activation, loss of TET2 sustains myeloproliferative neoplasms (MPNs) and accelerates cancer development in transgenic mouse models and patients41,42,43. Mechanistically, STAT signaling promotes cellular proliferation, while loss of TET2 stimulates self-renewal of leukemia stem cells. It is worth noting that the level of STAT pathway activation is a critical determinant in the disease phenotype. STAT pathway alone, when activated at an appropriate level, is a “weak” oncogene that gives rise to an MPN with low penetrance and delayed onset. A more drastic disease phenotype can be obtained by further loss of TET240. These observations are consistent with our model in that early stage STAT activation induced by IL10 overexpression is not sufficient to fully transform NK cells, secondary event (JAK/STAT or TET2/TP53 mutations) are needed to drive a full-blown disease (Figure 6).
The STATs-MYC-biosynthesis axis central to the molecular pathogenesis of ANKL is unique and has not been recognized previously in any other neoplasms. In multiple myeloma, T-LGL and various solid tumors (breast, prostate, ovarian, brain and pancreatic cancers), JAK-STAT signaling promotes pro-oncogenic inflammatory pathways by activating an expression signature that controls cell growth and survival (cyclin D1, MCL-1 and FGF2), angiogenesis (VEGFA), inflammatory factors (CXCL12, CCL2) and migration or invasion (MMP2 and MMP9)44. Consistent with this, activation of the pro-oncogenic inflammatory signature was evident in T-LGL. However, ANKL cells did not activate this inflammation program and possessed a distinct MYC expression signature (Figures 3C and 5A; Supplementary information, Figure S14). As a master regulator of cellular metabolism, MYC activation often results from gene amplification/translocation and upstream signaling pathways including NOTCH, WNT-β-catenin and sonic hedgehog-Gli45. The only evidence that JAK-STAT can activate MYC comes from an in vitro study reporting that the MYC promoter was bound by phosphorylated STAT3 (pSTAT3)46. ANKL is therefore the first human disease in which STAT3 signaling acts upstream to directly activate the MYC transcription program.
The fatal behavior and poor response of ANKL to current chemotherapy makes it of paramount importance to search for novel therapeutics. Our findings suggest that targeted therapies against IL10, JAK-STAT signaling, MYC or nucleotide synthesis/glycolysis hold great promise for ANKL, and thus, future investigations of these pathways are warranted (Figures 5C and 6). Recently, therapeutic utility of MYC inhibition has been revealed in NK-cell neoplasms, suggesting that MYC activation may represent a common mechanism in the pathogenesis of ANKL and NKTCL47. The hypothesis that nucleotide synthesis should be exploited for treatment is also supported by current clinical evidence. Specifically, regimens containing antimetabolites such as L-asparaginase, MTX and gemcitabine have proven to be the most effective for disseminated NKTCL as well as for limited cases of ANKL4,48, suggesting new treatment opportunities for refractory NK neoplasms by targeting the vulnerability in nucleotide synthesis.
Materials and Methods
Patients and sample description
Thirty-nine ANKL patients from 10 medical centers in China were included in this study. The main clinical characteristics are shown in Supplementary information, Table S1. For WGS, CD56+CD3− NK cells and matched CD33+CD14− granulocytes from 8 ANKL patients were isolated by fluorescence-activated cell sorting (FACS). For transcriptome analysis, CD56+CD3− NK cells from 7 ANKL patients and 4 healthy donors were isolated by FACS. The sorting strategy is shown in Supplementary information, Figure S2. AmpliSeq targeted sequencing of 36 mutated genes was performed in 29 ANKL patients. The study was approved by the Institutional Review Board. Informed consent was obtained from each individual in accordance with the principles expressed in the Declaration of Helsinki.
Whole genome and transcriptome sequencing, and data analyses
We performed WGS for CD56+CD3− NK cells and matched CD33+CD14− granulocytes from 8 ANKL patients. We conducted a genome-wide analysis of somatic SNVs, copy number variations and structural variations using GVC utilities, including GVC-SNV6, GVC-CNV and GVC-SV6. We performed RNA sequencing of both patient and cell line samples. Differentially expressed genes were identified using DESeq49 and DEGseq50 software. The pathways significantly enriched in differentially expressed genes were analyzed with the DAVID51 and IPA tools. Enrichment analysis of the gene signatures was performed using GSEA software52. The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive53 in BIG Data Center54, Beijing Institute of Genomics (BIG), Chinese Academy of Sciences, under accession numbers PRJCA000224, PRJCA000224 that are publicly accessible at http://bigd.big.ac.cn/gsa. Detailed library construction and data analysis are described in Supplementary information, Data S1.
Integrative analysis of genomic and transcriptomic data by TieDIE
To construct biological networks, we used an integrative method called TieDIE for the genomic and transcriptional data. TieDIE22 (Tied Diffusion through Interacting Events) is an integrative analysis method that employs a network diffusion approach to connect genetic perturbations (e.g., somatic mutations) with transcriptional changes. We used 313 genes with non-synonymous mutations identified in 8 ANKL patients as the upstream of the input and 778 transcriptionally upregulated genes as the downstream of the input. The obtained networks using default parameters were drawn using Cytoscape (Figure 3A).
Significantly mutated network identified by HotNet2
We identified significantly mutated sub-networks using HotNet211, which uses a directed heat diffusion model to calculate the significance of mutations in individual genes and the local topology of interactions among the encoded proteins to overcome the limitations of pathway-based enrichment statistics. All 313 genes with non-synonymous SNVs and 24 filtered genes from CNA-containing regions were used in the analysis. The filtered genes must be in consistent CNAs of at least 5 patients, and the FPKM values of the gene must be larger than 0.1. The networks are shown in Supplementary information, Figure S6.
Mutation validation in ANKL patients using AmpliSeq
Ion Torrent AmpliSeq was used for targeted sequencing of 28 mutated genes identified in the WGS analysis of 8 ANKL patients and 8 frequently mutated genes in NKTCL7,12. Based on ultrahigh-multiplex PCR, Ion AmpliSeq technology requires as little as 10 ng of input DNA from bone marrow smear samples to target sets of genes. The percentages of leukemia cells in bone marrow detected by flow cytometry of all included cases were between 8.0% and 76.0%, with an average of 29.4%. Only cases with over 10% leukemia cells were included. We designed primers for 36 target genes (Supplementary information, Table S5) for deep sequencing. We collected 34 samples from ANKL patients and 2 granulocyte samples in total. Five ANKL samples with targeted rates < 90% due to degradation were removed.
Cytokine chip array analysis
A custom-designed RayBio Sandwich-based Antibody Array (RayBiotech, Norcross, GA, USA) was spotted with specific antibodies against 20 inflammatory cytokines (IL-1α, IL-1, IL-2, IL-4, IL-6, IL-10, IL-12, IL-13, IL-15, IL-18, MIP-1α, TNFα, IFN-γ, TGF-β, CD163, IL-8, IFN-α, IFN-β, MCP-1, IL-3). The plasma cytokine levels were measured according to the manufacturer's instructions. Slides were scanned using an InnoScan 300 Microarray Scanner (Innopsys, France) at 532 nm with a resolution of 10 μm. All assays were performed in duplicate. Statistical analysis was performed with Student's t-test or Student's t-test with Welch's correction using GraphPad Prism 6.0c. Statistical significance was defined as P < 0.05.
Immunohistochemistry
The immunohistochemical staining of CD56, STAT3 phosphorylation (pSTAT3) and MYC in the bone marrow biopsy specimens were performed in ANKL and control samples, respectively. Lymphoma patients without bone marrow involvement were used as controls. The method of immunohistochemistry is described in Supplementary information, Data S1.
Cell lines
KHYG-1 was obtained from the Japanese Collection of Research Bioresources (JCRB) cell bank. NK-92 was obtained from the American Type Culture Collection (ATCC). YT was a gift from Dr Kai Fu at the University of Nebraska Medical Center. All cell lines were authenticated by short tandem repeat (STR) analysis and tested negative for mycoplasma contamination by R&D MycoProbe Mycoplasma Detection Kit. The cell culture methods are described in Supplementary information, Data S1.
Cell viability assays
Cells were seeded in 96-well plates in triplicate at a density of 1 × 104 cells/100 μL/well. For assays with Stattic (Sigma, Taufkirchen, Germany), the cells were treated with DMSO or the indicated concentrations of Stattic for 16 h. For assays with IL10 and IL18 (Life Technologies, USA), ANKL cell lines were pre-treated with a base dose of IL2 overnight and then stimulated with the indicated concentrations of IL10 or IL18 for 24 h. Cell viability was assayed with Cell Counting Kit-8 reagent (Dojindo, Japan), and the relative cell viability was calculated by normalizing to the vehicle control. The half-maximal inhibitory concentration (IC50) values were calculated by non-linear regression curve fitting using GraphPad Prism 6.0c. For all assays, at least three independent experiments were performed.
Colony-forming assays
KHYG-1 cells were seeded in 24-well plates at a density of 200 cells/500 μL/well and cultured in RPMI-1640, 1% methylcellulose (Sigma, Taufkirchen, Germany) and 10% FBS supplemented with 500 IU/mL IL2. The cells were maintained at 37 °C in a 5% CO2 and a > 95% humidity atmosphere for 14 days. Colony numbers were counted using a microscope equipped with a 2× objective and a 10× ocular eyepiece. All assays were performed in triplicate.
Statistical analyses
The Mann-Whitney U-test and Student's t-test were used for comparison of the results and were calculated using R and GraphPad Prism 6.0c software. Selections of all statistical analysis methods meet the assumptions of the tests. Equality of variances between the groups was statistically compared.
Author Contributions
JZ, QW and Liang H designed the project. NW, YT and XM performed cytokine array analyses and cell sorting. YT, NW, Hui L, XH, LS, LZ and MX performed cell line assays. Liang H, YT, JZ and QW mainly analyzed the experiment data. DL, SL, JW and Lingtong H performed sequencing data analyses. LC, LD, XZ, PS, JH, SM and JY assisted sequencing data analyses. SL and Lingtong H performed the validation of variations. DW, YL, KZ and ZS collected clinical data. Hongs Z, QL, Hongy Z, JL, JJ, LF, WZ, JC and XD provided clinical samples and relevant information. GQ, Hud L, XL, GH and DM provided support for the project. QW, JZ, YT, DL, SL and Liang H wrote the manuscript.
Competing Financial Interests
The authors declare no competing financial interests.
References
Chan J, Jaffe ES, Ralfkiaer E . Aggressive NK-cell leukaemia. In: Swerdlow S, Campo E, Harris N, eds. WHO Classification of Tumors of Heamatopoietic and Lymphoid Tissue. Lyon: International Agency for Research on Cancer, 2008:276–277.
Chan JK, Sin VC, Wong KF, et al. Nonnasal lymphoma expressing the natural killer cell marker CD56: a clinicopathologic study of 49 cases of an uncommon aggressive neoplasm. Blood 1997; 89:4501–4513.
Suzuki R, Suzumiya J, Nakamura S, et al. Aggressive natural killer-cell leukemia revisited: large granular lymphocyte leukemia of cytotoxic NK cells. Leukemia 2004; 18:763–770.
Suzuki R . Treatment of advanced extranodal NK/T cell lymphoma, nasal-type and aggressive NK-cell leukemia. Int J Hematol 2010; 92:697–701.
Nakashima Y, Tagawa H, Suzuki R, et al. Genome-wide array-based comparative genomic hybridization of natural killer cell lymphoma/leukemia: different genomic alteration patterns of aggressive NK-cell leukemia and extranodal Nk/T-cell lymphoma, nasal type. Genes Chromosomes Cancer 2005; 44:247–255.
Ling S, Hu Z, Yang Z, et al. Extremely high genetic diversity in a single tumor points to prevalence of non-Darwinian cell evolution. Proc Natl Acad Sci USA 2015; 112:E6496–E6505.
Jiang L, Gu ZH, Yan ZX, et al. Exome sequencing identifies somatic mutations of DDX3X in natural killer/T-cell lymphoma. Nat Genet 2015; 47:1061–1066.
Kandoth C, McLellan MD, Vandin F, et al. Mutational landscape and significance across 12 major cancer types. Nature 2013; 502:333–339.
Pasqualucci L, Trifonov V, Fabbri G, et al. Analysis of the coding genome of diffuse large B-cell lymphoma. Nat Genet 2011; 43:830–837.
Cancer Genome Atlas Research N. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 2013; 368:2059–2074.
Leiserson MD, Vandin F, Wu HT, et al. Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat Genet 2015; 47:106–114.
Dobashi A, Tsuyama N, Asaka R, et al. Frequent BCOR aberrations in extranodal NK/T-cell lymphoma, nasal type. Genes Chromosomes Cancer 2016; 55:460–471.
Jerez A, Clemente MJ, Makishima H, et al. STAT3 mutations unify the pathogenesis of chronic lymphoproliferative disorders of NK cells and T-cell large granular lymphocyte leukemia. Blood 2012; 120:3048–3057.
Koskela HL, Eldfors S, Ellonen P, et al. Somatic STAT3 mutations in large granular lymphocytic leukemia. N Engl J Med 2012; 366:1905–1913.
Kucuk C, Jiang B, Hu X, et al. Activating mutations of STAT5B and STAT3 in lymphomas derived from γδ-T or NK cells. Nat Commun 2015; 6:6025.
Bandapalli OR, Schuessele S, Kunz JB, et al. The activating STAT5B N642H mutation is a common abnormality in pediatric T-cell acute lymphoblastic leukemia and confers a higher risk of relapse. Haematologica 2014; 99:e188–e192.
Lopez C, Bergmann AK, Paul U, et al. Genes encoding members of the JAK-STAT pathway or epigenetic regulators are recurrently mutated in T-cell prolymphocytic leukaemia. Br J Haematol 2016; 173:265–273.
Pilati C, Amessou M, Bihl MP, et al. Somatic mutations activating STAT3 in human inflammatory hepatocellular adenomas. J Exp Med 2011; 208:1359–1366.
Gao LM, Zhao S, Liu WP, et al. Clinicopathologic characterization of aggressive natural killer cell leukemia involving different tissue sites. Am J Surg Pathol 2016; 40:836–846.
Roberts KG, Li Y, Payne-Turner D, et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N Engl J Med 2014; 371:1005–1015.
Yu H, Pardoll D, Jove R . STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer 2009; 9:798–809.
Paull EO, Carlin DE, Niepel M, Sorger PK, Haussler D, Stuart JM . Discovering causal pathways linking genomic events to transcriptional states using Tied Diffusion through Interacting Events (TieDIE). Bioinformatics 2013; 29:2757–2764.
Ji H, Wu G, Zhan X, et al. Cell-type independent MYC target genes reveal a primordial signature involved in biomass accumulation. PLoS One 2011; 6:e26057.
Perna D, Faga G, Verrecchia A, et al. Genome-wide mapping of Myc binding and gene regulation in serum-stimulated fibroblasts. Oncogene 2012; 31:1695–1709.
Dang CV . MYC, metabolism, cell growth, and tumorigenesis. Cold Spring Harb Perspect Med 2013; 3:a014217.
Stine ZE, Walton ZE, Altman BJ, Hsieh AL, Dang CV . MYC, metabolism, and cancer. Cancer Discov 2015; 5:1024–1039.
Martinez-Outschoorn UE, Peiris-Pages M, Pestell RG, Sotgia F, Lisanti MP . Cancer metabolism: a therapeutic perspective. Nat Rev Clin Oncol 2016; 14:11–31.
Loven J, Orlando DA, Sigova AA, et al. Revisiting global gene expression analysis. Cell 2012; 151:476–482.
Caro P, Kishan AU, Norberg E, et al. Metabolic signatures uncover distinct targets in molecular subsets of diffuse large B cell lymphoma. Cancer Cell 2012; 22:547–560.
Li B, Qiu B, Lee DS, et al. Fructose-1,6-bisphosphatase opposes renal carcinoma progression. Nature 2014; 513:251–255.
Sellers K, Fox MP, Bousamra M, et al. Pyruvate carboxylase is critical for non-small-cell lung cancer proliferation. J Clin Invest 2015; 125:687–698.
Palomero T, Lim WK, Odom DT, et al. NOTCH1 directly regulates c-MYC and activates a feed-forward-loop transcriptional network promoting leukemic cell growth. Proc Natl Acad Sci USA 2006; 103:18261–18266.
Yagita M, Huang CL, Umehara H, et al. A novel natural killer cell line (KHYG-1) from a patient with aggressive natural killer cell leukemia carrying a p53 point mutation. Leukemia 2000; 14:922–930.
Gong JH, Maki G, Klingemann HG . Characterization of a human cell line (NK-92) with phenotypical and functional characteristics of activated natural killer cells. Leukemia 1994; 8:652–658.
Schust J, Sperl B, Hollis A, Mayer TU, Berg T . Stattic: a small-molecule inhibitor of STAT3 activation and dimerization. Chem Biol 2006; 13:1235–1242.
Samanta M, Iwakiri D, Takada K . Epstein-Barr virus-encoded small RNA induces IL-10 through RIG-I-mediated IRF-3 signaling. Oncogene 2008; 27:4150–4160.
Kitagawa N, Goto M, Kurozumi K, et al. Epstein-Barr virus-encoded poly(A)(-) RNA supports Burkitt's lymphoma growth through interleukin-10 induction. EMBO J 2000; 19:6742–6750.
Papaemmanuil E, Gerstung M, Malcovati L, et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 2013; 122:3616–3627.
Odejide O, Weigert O, Lane AA, et al. A targeted mutational landscape of angioimmunoblastic T-cell lymphoma. Blood 2014; 123:1293–1296.
Vainchenker W, Plo I . TET2 loss, a rescue of JAK2V617F HSCs. Blood 2015; 125:212–213.
Chen E, Schneider RK, Breyfogle LJ, et al. Distinct effects of concomitant c expression and Tet2 loss in mice promote disease progression in myeloproliferative neoplasms. Blood 2015; 125:327–335.
Kameda T, Shide K, Yamaji T, et al. Loss of TET2 has dual roles in murine myeloproliferative neoplasms: disease sustainer and disease accelerator. Blood 2015; 125:304–315.
Ortmann CA, Kent DG, Nangalia J, et al. Effect of mutation order on myeloproliferative neoplasms. N Engl J Med 2015; 372:601–612.
Miklossy G, Hilliard TS, Turkson J . Therapeutic modulators of STAT signalling for human diseases. Nat Rev Drug Discov 2013; 12:611–629.
Conacci-Sorrell M, McFerrin L, Eisenman RN . An overview of MYC and its interactome. Cold Spring Harb Perspect Med 2014; 4:a014357.
Kiuchi N, Nakajima K, Ichiba M, et al. STAT3 is required for the gp130-mediated full activation of the c-myc gene. J Exp Med 1999; 189:63–73.
Selvarajan V, Osato M, Nah GS, et al. RUNX3 is oncogenic in natural killer/T-cell lymphoma and is transcriptionally regulated by MYC. Leukemia 2017; 31:2219–2227.
Wang L, Wang ZH, Chen XQ, et al. First-line combination of gemcitabine, oxaliplatin, and L-asparaginase (GELOX) followed by involved-field radiation therapy for patients with stage IE/IIE extranodal natural killer/T-cell lymphoma. Cancer 2013; 119:348–355.
Anders S, Huber W . Differential expression analysis for sequence count data. Genome Biol 2010; 11:R106.
Wang L, Feng Z, Wang X, Wang X, Zhang X . DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010; 26:136–138.
Huang DW, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4:44–57.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102:15545–15550.
Wang Y, Song F, Zhu J, et al. GSA: Genome Sequence Archive. Genomics Proteomics Bioinformatics 2017; 15:14–18.
BIG Data Center Members. The BIG Data Center: from deposition to integration to translation. Nucleic Acids Res 2017; 45:D18–D24.
Acknowledgements
We thank all the faculties and staffs in the Clinical and Laboratory Unit of the Department of Hematology, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology for their clinical and technical support; the Core Genomic Facility of Beijing Institute of Genomics, Chinese Academy of Sciences for the construction and sequencing of libraries; Drs Qing Li and Jiguang Wang for critical reading and valuable comments on the manuscript; Dr Kai Fu for providing the cell line YT. This study was supported by the National Natural Science Foundation of China (81570196 to JZ, 81425003 to QW, 81670152 to Liang H, 81600120 to NW, 81300410 to DW, 81500100 to YL and 81400122 to KZ), the National Key Basic Research Program of China (2014CB542001 to QW), the Key Program of the National Natural Science Foundation of China (81230052 to JZ), the Key Research Program of the Chinese Academy of Sciences (Precious Medicine Research in Chinese Population; KJZD-EW-L14-3 to QW), and the National High Technology Research and Development Program of China (863 program; 2012AA02A507 to JZ and 2014AA020532 to Liang H).
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Strategies for the integrative analysis of ANKLs. (PDF 401 kb)
Supplementary information, Figure S2
Isolation of granulocytes, leukemia NK cells and normal NK cells. (PDF 387 kb)
Supplementary information, Figure S3
Frequency of somatic SNV of ANKL and 40 additional cancer types. (PDF 493 kb)
Supplementary information, Figure S4
EBV reads number of leukemia NK cells and granulocytes from 8 ANKL patients. (PDF 421 kb)
Supplementary information, Figure S5
KEGG pathway enrichment analysis of protein-altering somatic SNVs. (PDF 438 kb)
Supplementary information, Figure S6
Functional enrichment analysis of genomic alterations with HotNet2. (PDF 373 kb)
Supplementary information, Figure S7
STAT3 and STAT5B mutations identified in ANKL patients. (PDF 541 kb)
Supplementary information, Figure S8
TET2 mutations identified in ANKL patients. (PDF 471 kb)
Supplementary information, Figure S9
Semi-quantitative immunoreactivity histological scores of MYC staining in bone marrow biopsy of ANKL and control samples. (PDF 422 kb)
Supplementary information, Figure S10
Functional enrichment map for MYC-signature genes that were upregulated in ANKLs compared to healthy donors. (PDF 856 kb)
Supplementary information, Figure S11
The effect of IL10 or a STAT3 inhibitor (Stattic) on the apoptosis of ANKL cell lines. (PDF 457 kb)
Supplementary information, Figure S12
The effect of IL10 and the STAT3 inhibitor (Stattic) on the mRNA expression of MYC in ANKL cell lines. (PDF 470 kb)
Supplementary information, Figure S13
The rate of EdU incorporation in JQ1-treated ANKL cell lines. (PDF 404 kb)
Supplementary information, Figure S14
Gene-set enrichment analysis (GSEA) of known MYC and STAT3 signatures. (PDF 523 kb)
Supplementary information, Figure S15
Enrichment of metabolic pathways in primary ANKL leukemia cells and ANKL cell lines. (PDF 480 kb)
Supplementary information, Figure S16
The effect of STAT3 Y640F mutant on the phosphorylation of STAT3 and mRNA expression of MYC target gene. (PDF 583 kb)
Supplementary information, Figure S17
The effect of IL10 on the mRNA expression of MYC and MYC target genes in KHYG-1 cell line transfected with STAT3 Y640F mutant. (PDF 584 kb)
Supplementary information, Figure S18
The effect of IL10 on STAT3 phosphorylation, MYC expressionand the proliferation of STAT3 Y640F-mutant ANKL cell line YT. (PDF 505 kb)
Supplementary information, Figure S19
Quantitative analysis of the Epstein-Barr virus (EBV) load in ANKL patients and healthy donors with whole-transcriptome sequencing (WTS) data. (PDF 351 kb)
Supplementary information, Figure S20
Expression of EBV-encoded small RNAs in primary ANKL leukemia cells. (PDF 400 kb)
Supplementary information, Figure S21
A large gain across MYC and an inter-chromosomal translocation detected by GVC-CNV and GVC-SV in ANKL No.19. (PDF 456 kb)
Supplementary information, Table S1
Patient characteristics of ANKL. (PDF 633 kb)
Supplementary information, Table S2
SNVs identified in ANKL NK leukemia cells. (PDF 412 kb)
Supplementary information, Table S3
CNVs identified in ANKL NK leukemia cells. (PDF 6359 kb)
Supplementary information, Table S4
SVs identified in ANKL NK leukemia cells. (PDF 3596 kb)
Supplementary information, Table S5
AmpliSeq targeted sequencing results of ANKLs. (PDF 630 kb)
Supplementary information, Table S7
H-score of phosphorylated STAT3 and corresponding STAT mutation status of ANKL cases. (PDF 369 kb)
Supplementary information, Table S8
Differentially upregulated genes in ANKL leukemia cells in comparison with normal controls. (PDF 730 kb)
Supplementary information, Table S9
Differentially downregulated genes in ANKL leukemia cells in comparison with normal controls. (PDF 1385 kb)
Supplementary information, Table S10
KEGG pathway enrichment analysis of upregulated genes in ANKL leukemia cells. (PDF 548 kb)
Supplementary information, Table S11
KEGG pathway enrichment analysis of downregulated genes in ANKL leukemia cells. (PDF 398 kb)
Supplementary information, Table S12
IPA metabolic pathways analysis of ANKL and tumors that have known metabolic features. (PDF 1531 kb)
Supplementary information, Table S13
Sequencing depth information of 8 WGS ANKL patients. (PDF 430 kb)
Supplementary information, Table S14
Primers for quantitative RT-PCR (PDF 319 kb)
Supplementary information, Data S1
Materials and methods (PDF 1056 kb)
Rights and permissions
About this article
Cite this article
Huang, L., Liu, D., Wang, N. et al. Integrated genomic analysis identifies deregulated JAK/STAT-MYC-biosynthesis axis in aggressive NK-cell leukemia. Cell Res 28, 172–186 (2018). https://doi.org/10.1038/cr.2017.146
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2017.146
Keywords
This article is cited by
-
Initial diagnosis of extranodal NK/T-cell lymphoma in pericardial fluid with concomitant hemophagocytic lymphohistiocytosis (HLH)
Journal of Hematopathology (2024)
-
Outcome of aggressive B-cell lymphoma with TP53 alterations administered with CAR T-cell cocktail alone or in combination with ASCT
Signal Transduction and Targeted Therapy (2022)
-
The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms
Leukemia (2022)
-
Directly targeting c-Myc contributes to the anti-multiple myeloma effect of anlotinib
Cell Death & Disease (2021)
-
Genomic and transcriptomic landscapes of Epstein-Barr virus in extranodal natural killer T-cell lymphoma
Leukemia (2019)