Abstract
Subtelomeres consist of sequences adjacent to telomeres and contain genes involved in important cellular functions, as subtelomere instability is associated with several human diseases. Balancing between subtelomere stability and plasticity is particularly important for Trypanosoma brucei, a protozoan parasite that causes human African trypanosomiasis. T. brucei regularly switches its major variant surface antigen, variant surface glycoprotein (VSG), to evade the host immune response, and VSGs are expressed exclusively from subtelomeres in a strictly monoallelic fashion. Telomere proteins are important for protecting chromosome ends from illegitimate DNA processes. However, whether they contribute to subtelomere integrity and stability has not been well studied. We have identified a novel T. brucei telomere protein, T. brucei TRF-Interacting Factor 2 (TbTIF2), as a functional homolog of mammalian TIN2. A transient depletion of TbTIF2 led to an elevated VSG switching frequency and an increased amount of DNA double-strand breaks (DSBs) in both active and silent subtelomeric bloodstream form expression sites (BESs). Therefore, TbTIF2 plays an important role in VSG switching regulation and is important for subtelomere integrity and stability. TbTIF2 depletion increased the association of TbRAD51 with the telomeric and subtelomeric chromatin, and TbRAD51 deletion further increased subtelomeric DSBs in TbTIF2-depleted cells, suggesting that TbRAD51-mediated DSB repair is the underlying mechanism of subsequent VSG switching. Surprisingly, significantly more TbRAD51 associated with the active BES than with the silent BESs upon TbTIF2 depletion, and TbRAD51 deletion induced much more DSBs in the active BES than in the silent BESs in TbTIF2-depleted cells, suggesting that TbRAD51 preferentially repairs DSBs in the active BES.
Similar content being viewed by others
Introduction
The ends of linear chromosomes form specialized nucleoprotein complexes termed telomeres, which often consist of simple repetitive TG-rich sequences (e.g., (TTAGGG)n in vertebrates and Trypanosoma brucei) and associated proteins. Sequences located next to telomeres are subtelomeres. In contrast to previous beliefs that subtelomeres are barren regions of no importance, they contain genes involved in important cellular functions, and instability of human subtelomere regions is often associated with diseases. For example, the double homeobox 4 (DUX4) gene locates in the D4Z4 repeat array at human 4q35 subtelomere1. Drastically reduced D4Z4 repeat number can lead to abnormal DUX4 transcription and is associated with facioscapulohumeral muscular dystrophy (FSHD)2. Submicroscopic deletion of subtelomeric 6p25 and 9q has been recognized as clinically identifiable syndromes3,4. In addition, some OR genes encoding olfactory receptors are located at subtelomeres, and changes in subtelomeres contribute to the diversity of the OR gene family5.
Subtelomeres are particularly important for a number of microbial pathogens that undergo antigenic variation, including T. brucei that causes human African trypano-somiasis, Plasmodium falciparum that causes malaria, Pneumocystis jirovecii that causes pneumonia, and Borrelia burgdorferi that causes Lyme disease6. These pathogens regularly switch their major surface antigen to evade the host immune response, and genes encoding variant surface antigens are expressed exclusively (or sometimes in P. falciparum) from subtelomeric regions in a strictly monoallelic fashion6. Therefore, plasticity at subtelomeric regions is necessary for these microbial pathogens. However, maintaining a relatively stable subtelomere is also important, as unstable subtelomeres can lead to loss of functional surface antigen genes. How do these pathogens maintain a delicate balance between stability and plasticity at their subtelomeres is not well known.
Maintenance of subtelomere stability is challenging because subtelomeres often consist of DNA blocks duplicated on multiple chromosomes and are highly dynamic with very heterogeneous sequences, sizes, and copy numbers7. Human subtelomeres are hot spots of interchromosomal recombination and segmental duplications8. A high level of nucleotide divergence among Saccharomyces yeasts has been observed at the subtelomeres9. Similarly in T. brucei, subtelomeres have been considered fragile sites and undergo DNA recombination frequently10.
It is well known that proteins associated with the telomere are essential for maintaining the telomere stability11. However, whether telomere proteins directly contribute to subtelomere stability has not been well studied. “Shelterin” is a six-member complex that associates with the vertebrate telomere12. In the complex, TRF1 and TRF2 bind the duplex TTAGGG repeats13,14,15, and the TPP1/POT1 heterodimer binds the telomeric single-stranded 3′ overhang16. As a core component of Shelterin, TIN2 interacts with TRF1, TRF2, and TPP1 directly17,18,19,20,21. Shelterin components are essential for preventing the natural chromosome ends from being processed as double-strand breaks (DSBs)12. For example, TRF2 suppresses ATM activation and POT1 inhibits ATR activation at the telomere22. In addition, telomere proteins are important for telomeric DNA replication. Deletion of TRF1 results in replication fork stalling in telomere repeats and fragile telomeres23, and TRF2 coordinates with its interacting factor Apollo, a 5′ exonuclease, to relieve topological stress during telomere replication24. X. laevis TRF2 and fission yeast TAZ1 that binds the duplex telomere DNA are also required for DNA replication at the telomere25,26. Mammalian TIN2 also interacts directly with SA1, a telomere-specific cohesin subunit27,28, and HP1γ29, which is required to establish/maintain telomere cohesion and maintain proper telomere length29. Presumably, replication stress or improper sister telomere pairing resulting from dysfunctional telomere proteins can affect subtelomere stability, but this has not been shown in microbial pathogens such as T. brucei.
T. brucei is the causal agent of human African trypanosomiasis, which is fatal without treatment. Bloodstream form (BF) T. brucei stays in extracellular spaces inside the mammalian host and regularly switches its major surface antigen, variant surface glycoprotein (VSG), to evade the host immune response30. T. brucei has more than 2 000 VSG genes and pseudogenes31, but VSGs are exclusively expressed from BF expression sites (BESs), which are polycistronic transcription units at subtelomeres32. VSG is the last gene in any BES, located less than 2 kb from the telomere and 40-60 kb downstream of the BES promoter33. T. brucei has multiple BESs (15-20 in the lister 427 strain used in this work), but only one is fully active, expressing a single type of VSG at any time34,35.
VSG switching has several major pathways36. In in situ switching, an originally silent BES is expressed, while the originally active BES is silenced without DNA rearrangements. Other major VSG switching pathways are homologous recombination (HR) mediated. In crossover (CO) events, part of the active BES, including the VSG gene, changes place with that of a silent BES without losing any genetic information. In gene conversion (GC) events, a silent VSG is copied into the active BES to replace the active VSG gene, resulting in the loss of the originally active VSG and duplication of the newly active VSG. GC is the preferred mechanism of VSG switching36. It can encompass only the VSG gene and its neighboring regions (VSG GC) or the entire BES (ES GC). Several proteins play important roles in VSG switching. RAD51 binds to the single-stranded 3′ overhang at DNA DSBs following 5′ resection and promotes strand invasion during HR37. Deletion of TbRAD51 or TbRAD51-3 leads to decreased VSG switching frequencies38,39. TbBRCA2 is similarly required for efficient VSG switching40. In contrast, deletion of TOPO3 and RMI1 leads to more frequent VSG switching41,42. Recent studies showed that inducing DSBs in the active BES increases the VSG switching rate, and different VSG switching mechanisms are used depending on the DSB position10,43. As high as 25% of DSBs downstream of the active VSG result in the loss of the entire active BES10, and an accompanying in situ switch can give rise to ES loss coupled with in situ switchers (ES loss + in situ). Interestingly, DSBs in silent BESs seldom lead to VSG switching10. However, how DSBs in the active and silent BESs lead to different outcomes is not well known.
T. brucei is transmitted by the tsetse fly (Glossina spp.). In the midgut of the insect host, procyclic form (PF) T. brucei expresses procyclins instead of VSG. Upon reaching the salivary glands, T. brucei differentiates into the metacyclic form, reacquires infectivity, and expresses metacyclic VSGs (mVSGs) from metacyclic ESs (MESs), which are monocistronic transcription units located in subtelomeric regions44. mVSGs are silenced soon after T. brucei cells are transmitted into the mammalian host.
We have shown that a telomere protein, TbRAP1, plays an essential role in silencing subtelomeric VSGs45,46. However, it is not known whether telomere proteins influence VSG switching, although extremely short telomeres (∼1.5 kb) elevate VSG switching frequencies by ∼10-fold47. In this study, we identified a novel T. brucei telomere protein, T. brucei TRF-Interacting Factor 2 (TbTIF2), that interacts with TbTRF and is functionally homologous to mammalian TIN2. A transient depletion of TbTIF2 by RNAi led to an increase in VSG switching frequency where most switchers arose from GC events, indicating that telomere proteins are indeed important for VSG switching regulation and contribute to subtelomere stability. Depletion of TbTIF2 increased subtelomeric DSBs, indicating that TbTIF2 contributes directly to subtelomere integrity. More TbRAD51 associated with telomeric and subtelomeric chromatin following TbTIF2 depletion, and this increase was, surprisingly, much stronger at the active BES than at silent ones. Deletion of TbRAD51 further increased subtelomeric DSB levels in TbTIF2-depleted cells, particularly in the active BES, suggesting that TbRAD51-mediated DSB repair in the active BES is the underlying mechanism of the increased VSG switching and that the choice of DSB repair mechanisms may be influenced by transcriptional status of the DSB site.
Results
T. brucei TIF2 is a telomeric protein
We have previously identified T. brucei TRF as a telomere-binding protein48. We have also identified TbRAP1 as a TbTRF-interacting factor and have shown that TbRAP1 is essential for silencing of subtelomeric BES-linked and metacyclic VSG genes45,46. To further investigate telomere functions in antigenic variation, we attempted to identify additional TbTRF-interacting factors.
In order to pull down the TbTRF protein complex specifically, we established a TbTRF single-allele knockout PF strain with the remaining endogenous TbTRF allele tagged with an N-terminal FLAG-HA-HA (F2H) epitope. This strain grows normally (Supplementary information, Figure S1A), indicating that the sole F2H-TbTRF allele is functional. We immunoprecipitated (IP) the whole cell extract with the FLAG monoclonal antibody M2 (Sigma), eluted the IP product with the FLAG peptide, and immunoprecipitated the eluate with the HA monoclonal antibody 12CA5 (MSKCC monoclonal AB core). The final IP product was separated on a polyacrylamide gel (Supplementary information, Figure S1B) and analyzed by mass spectrometry. WT cells without the F2H tag were treated exactly the same way and proteins identified in the final IP product by mass spectrometry were considered as background contaminants. In addition to TbTRF, we identified a novel protein (TriTrypDB ID: Tb427.03.1560) in the TbTRF IP product.
To verify the interaction between Tb427.03.1560 and TbTRF, we first performed yeast two-hybrid analysis. Full-length Tb427.03.1560 and TbTRF interact much more strongly than TbTRF homodimerization (Figure 1A and Supplementary information, Figure S1C), confirming that Tb427.03.1560 interacts with TbTRF directly when analyzed in yeast. We therefore named this protein T. brucei TRF-Interacting Factor 2 (TbTIF2; TbRAP1 is TbTRF-interacting factor 145). Sequence analysis of TbTIF2 did not identify any functional domains. However, alignment of TbTIF2 with known telomere proteins showed that it is weakly homologous to TIN2 (14% sequence identity between TbTIF2 and human TIN2; Supplementary information, Figure S1D). T. brucei telomere proteins exhibit significant similarities to their vertebrate homologs only within functional domains. The identity between TbTRF and vertebrate TRFs is 18%-22% in the Myb domain and 9%-12% in the TRFH domain, respectively48. The identity between TbRAP1 and vertebrate RAP1s is 8%-23% within various functional domains45. In addition, yeast two-hybrid analysis using various fragments of TbTIF2 and TbTRF showed that the N-terminal half of TbTIF2 interacts with the TRFH domain of TbTRF (Figure 1A), which is partially conserved with the situation in mammalian cells, where the N-terminal half of TIN2 interacts with a short peptide in the linker region of TRF2, while a short stretch towards the C-terminal half of TIN2 interacts with the TRFH domain of TRF149. Therefore, TbTIF2 is likely a functional homolog of vertebrate TIN2.
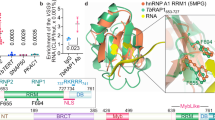
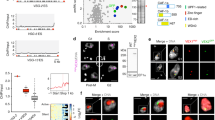
TbTIF2 interacts with TbTRF and associates with telomeres. (A) TbTIF2 and TbTRF interact in yeast two-hybrid analysis. Top, different Gal4 Activation Domain (GAD)-fused TbTIF2 fragments (starting and ending amino acids of each fragment are listed in the parentheses) were tested for their interaction with LexA-fused full-length TbTRF or just LexA (empty). The expression of GAD-TbTIF2 fragments was examined by western blotting (Supplementary information, Figure S1C). Middle, different LexA-fused TbTRF fragments were tested for their interaction with the GAD-fused full-length TbTIF2. Control experiments testing the interaction between LexA-TbTRF and GAD alone were performed previously48. β-galactosidase activities, shown as average (calculated from at lease four independent tests) ± standard deviations, reflect the expression of the reporter LacZ gene resulting from the interaction between the LexA- and GAD-fused proteins. A summary of the interaction between TbTIF2 and TbTRF is shown at the bottom. (B) TbTIF2 and TbTRF interact in vivo. Whole Cell Extract (WCE) was prepared from SM/TbTIF2-F2H. The soluble fraction of the lysate (soluble lysate) was cleared with protein G beads (IP input) and Immunoprecipitated with TbTRF antibody 126148, HA antibody 12CA5, or no antibody. IP supernatant (sup) and IP product (pellet) were subsequently analyzed by western blotting using HA antibody F-7 (Santa Cruz Biotechnology, Inc.) or chicken anti-TbTRF antibody 60645. (C) Immunofluorescence analysis of SM/TbTIF-F2H cells using 12CA5 and 1261. DAPI stains DNA in both the nucleus (larger blue circle) and the kinetoplast (small blue dot). (D) ChIP analysis in SM/TbTIF2-F2H cells using F-7, 1261, or mouse IgG (M-IgG). ChIP products were hybridized with a TTAGGG or a 50 bp repeat probe. Representative slot blots are shown in Supplementary information, Figure S1F. The blots were exposed to a phosphorimager and results were quantified by ImageQuant. Average was calculated from at least three independent experiments. In this and following figures, error bars represent standard deviation. Numbers next to the brackets indicate P-values (unpaired t-tests) between different experiments. P < 0.05 is considered to be significant.
To further characterize TbTIF2, we established T. brucei strains that carry a C-terminally F2H-tagged TbTIF2 at its endogenous locus and found that TbTIF2 and TbTRF co-immunoprecipitated in both BF (Figure 1B) and PF (Supplementary information, Figure S1E) cells when we used either TbTRF antibody or HA antibody for IP, further confirming that TbTIF2 and TbTRF interact in vivo.
Immunofluorescence using an HA antibody and a TbTRF antibody (served as a marker for the telomere) in TbTIF2-F2H-containing cells showed that TbTIF2 and TbTRF co-localized in the nucleus (Figure 1C), indicating that TbTIF2 is located at the telomere. Chromatin IP (ChIP) with an HA antibody in these cells showed that telomeric DNA was significantly enriched in the ChIP product (Figure 1D and Supplementary information, Figure S1F), while the 50 bp repeat DNA located upstream of BES promoters was not (Figure 1D and Supplementary information, Figures S1F and S2A). These results indicate that TbTIF2 is a component of T. brucei telomere complex.
TbTIF2 is essential for cell viability and mildly affects VSG silencing at some telomeres
To examine the functions of TbTIF2, we established TbTIF2 RNAi strains in a BF strain that express T7 polymerase and Tet repressor and allow inducible expression of the RNAi construct (SM,50). One endogenous TbTIF2 allele was also tagged with F2H in these cells. Since these cells express VSG2, we named them 2/TIF2i. Northern blotting using the TbTIF2 probe and western blotting using an HA antibody showed that the levels of TbTIF2 RNA (Supplementary information, Figure S2B) and TbTIF2 protein (Figure 2A) were substantially decreased following induction of TbTIF2 RNAi by doxycycline. TbTIF2-depleted cells experienced a growth arrest within 24 h (Figure 2B), indicating that TbTIF2 is essential for cell proliferation. FACS analysis showed that depletion of TbTIF2 led to a decrease in G1 cells (2C) and an increase in G2/M (4C) and polyploidy cells (6C and 8C) (Supplementary information, Figure S3). This is similar to the situation in mouse cells, where deletion of TIN2 leads to polyploidization resulting from endoreduplication51.
TbTIF2 is essential for cell survival. (A) Induction of TbTIF2 RNAi led to a decreased TbTIF2 protein level. Whole cell extracts were prepared from 2/TIF2i clone B9 cells at different time points after induction of TbTIF2 RNAi. HA antibody F-7 and EF-2 antibody (as a loading control, Santa Cruz Biotechnologies) were used in western blotting analyses. (B) Depletion of TbTIF2 by RNAi led to a growth arrest in T. brucei cells. Growth curves for SM/TbTIF2-F2H and 2/TIF2i cells in the presence (+) or the absence (−) of doxycycline (Dox). Average population doublings were calculated from three independent cultures. (C) TbTIF2 depletion had mild effects on subtelomeric VSG silencing. mRNA levels for several BES-linked VSGs and mVSGs were estimated by qRT-PCR at different time points after induction of TbTIF2 RNAi. The fold changes in mRNA levels were calculated from three independent experiments. 0 h values are all equal to “1” but not shown. Changes significantly different from that of rRNA are indicated by asterisks. ***P < 0.001 (unpaired t-tests).
To explore whether TbTIF2 is involved in regulation of subtelomeric VSG expression, we performed quantitative RT-PCR (qRT-PCR) using primers specific to several BES-linked VSGs. In 2/TIF2i cells where normally only VSG2 is active, we observed an ∼10-fold derepression of VSG14 but no significant derepression (consistent, >2-fold increase of mRNA level) of other BES-linked VSGs upon depletion of TbTIF2 (Figure 2C). VSG14 resides in BES8, which is the shortest BES (∼10 kb) in our lab strain, containing only ESAG7, 70 bp repeats, and VSG14, while most other BESs are 40-60 kb long (Supplementary information, Figure S2A)33. BES10 is the second shortest BES (25-30 kb) in this strain33, and we observed occasional, though not consistent derepression of VSG15 that resides in BES10 (Figure 2C). Thus, TbTIF2 may affect BES promoters that are close to telomeres. Because MESs are monocistronic transcription units whose promoters are usually only 5 kb upstream of the telomere (Supplementary information, Figure S2A), we tested whether mVSGs are affected by TbTIF2 depletion. We examined the mRNA levels of mVSG531, 639, and 39752 and found that mVSG531 and mVSG397 were derepressed by 3-4-fold upon depletion of TbTIF2 (Figure 2C). In an independent TbTIF2 RNAi strain, S/TIF2i (see below), we observed the same VSG derepression pattern: VSG14 was derepressed by 6-7-fold, while mVSG539 and mVSG397 were derepressed by 2-3-fold upon TbTIF2 depletion (Supplementary information, Figure S2C). Therefore, depletion of TbTIF2 led to a mild derepression of certain subtelomeric VSGs.
TbTIF2 suppresses VSG switching
To test whether TbTIF2 affects subtelomeric VSG switching, we introduced the inducible TbTIF2 RNAi construct into the SM-derived HSTB261 strain, which was designed for VSG switching analysis (Supplementary information, Figure S4)42. We obtained two independent S/TIF2i clones (“S” stands for “switching”), in which one endogenous TbTIF2 allele was tagged with F2H.
HSTB261 cells carry a blasticidin resistance (BSD) marker immediately downstream of the active BES promoter and a puromycin resistance (PUR) gene fused with the Herpes simplex virus thymidine kinase (TK) gene between the 70 bp repeats and the active VSG2 gene in the same BES (Supplementary information, Figure S4). All VSG switchers are expected to lose the expression of TK and become resistant to ganciclovir (GCV), a nucleoside analogue, allowing easy selection of VSG switchers. Because TbTIF2 is essential for cell survival, we recovered switchers after only transient TbTIF2 depletions. Inducing TbTIF2 RNAi in S/TIF2i cells for 30 h resulted in a transient growth arrest (Figure 3A) and a temporary decrease in TbTIF2 protein level for ∼24 h (Figure 3B). The recovered cells experienced a growth arrest again upon reinduction of TbTIF2 RNAi (Supplementary information, Figure S5A), indicating that these cells did not lose the TbTIF2 RNAi construct or the RNAi mechanism.
Transient depletion of TbTIF2 led to an elevated VSG switching frequency. (A) Induction of TbTIF2 for 30 h led to a growth arrest for ∼24 h. S/TIF2i clone A14 and S/v cells were cultured without doxycycline, with doxycycline for 30 h, or with doxycycline throughout the experiment. Average population doublings were calculated from four independent experiments to plot the growth curve. (B) The TbTIF2 protein level decreased for ∼24 h when TbTIF2 RNAi was induced for 30 h in S/TIF2i cells. Western blotting analyses were performed using HA antibody F-7 and EF-2 antibody. S/v cells were treated the same way as a control. (C) VSG switching frequencies in S/TIF2i and control cells. Cells were cultured with doxycycline for 30 h or without doxycycline as indicated. Average switching frequencies were calculated from 4-12 independent assays. P-values (unpaired t-tests) between S/v and other cells are shown as numbers on top of the corresponding columns. (D) VSG switching pathways in S/TIF2i and control cells. Percent of each VSG switching pathway of total events was plotted. Listed on top of each column is the number of switchers analyzed. Detailed switcher characterization results are listed in Supplementary information, Tables S3-S6.
We performed the switching assay as reported previously42. The parental HSTB261 cells, control cells carrying an empty vector (S/v), and two independent S/TIF2i clones (A12 & A14) were incubated with or without doxycycline for 30 h, washed extensively, and allowed to recover for 2-3 days. All cells were cultured for the same number of population doublings to allow a fair comparison of switching frequencies among different strains. Subsequently, cells were plated in the presence of GCV to select for switchers and in the absence of GCV to determine the plating efficiencies. Plating efficiencies (Supplementary information, Figure S5B) were used to normalize the final VSG switching frequencies.
We found that a transient depletion of TbTIF2 led to a 4.2–5.8-fold increase in VSG switching frequency when compared to S/v cells, while uninduced S/TIF2i and parental cells exhibited similar VSG switching frequencies to S/v cells (Figure 3C). To confirm that this effect was specifically due to depletion of TbTIF2, we performed a complementation analysis. An inducible expression vector containing FLAG-HA (FH)-tagged TbTIF2 was transfected into S/TIF2i A14 cells. Culturing S/TIF2i + TbTIF2-FH cells in the presence of doxycycline for 30 h resulted in a decrease in the endogenous TbTIF2 mRNA level (Supplementary information, Figure S5C, left) and a simultaneous increase in TbTIF2-FH mRNA (Supplementary information, Figure S5C, left) and protein (Supplementary information, Figure S5C, right) levels. This induction did not cause cell growth arrest (Supplementary information, Figure S5D). The VSG switching frequency in these cells was comparable to that in S/v cells (Figure 3C), indicating that the elevated VSG switching frequency was specifically due to depletion of TbTIF2. Therefore, our observations indicate that TbTIF2 suppresses VSG switching. The bulk telomeres in S/TIF2i cells are 15 kb long on average, and depletion of TbTIF2 did not result in a dramatic telomere shortening within the time frame of the switching experiment (Supplementary information, Figure S5E). Therefore, increased VSG switching following TbTIF2 depletion was not due to extremely short telomeres.
VSG switching can occur through several pathways including in situ switching, crossover (CO), VSG and ES gene conversion (GC), and ES loss + in situ switching. By examining marker genotypes and antibiotic resistance phenotypes in the recovered switchers, we can determine the mechanism of a given switching event (Supplementary information, Figure S4). However, this assay cannot differentiate between ES GC and ES loss + in situ events. These can only be differentiated by pulsed-field gel electrophoresis (PFGE) of intact chromosomes followed by Southern analysis. The majority of switchers in TbTIF2-depleted and control cells (∼90%) arose from VSG GC and ES GC/ES loss + in situ events (Figure 3D), suggesting that GC is the predominant mechanism of VSG switching under these conditions. ES GC/ES loss + in situ is most popular (60%-75%) in control cells and became even more prevalent when TbTIF2 was transiently depleted (> 80%; Figure 3D), indicating that there were more subtelomeric gene rearrangements when TbTIF2 was depleted. Therefore, TbTIF2 is important for subtelomere stability. To verify each VSG switching mechanism in several representative switchers, we determined their active VSGs by RT-PCR using a spliced leader primer (common to all T. brucei mRNAs) and a VSG C-terminal primer (common to VSGs) followed by sequencing analysis. PFGE of intact chromosomes from these switchers followed by Southern analyses using probes specific for VSG2, BSD, and the newly active VSG confirmed all predicted switching mechanisms (Supplementary information, Figure S6). ES loss + in situ switchers were present in both the parental strain and TbTIF2-depleted cells (Supplementary information, Figure S6; data not shown), indicating that this mechanism is not unique in TbTIF2 RNAi cells.
Depletion of TbTIF2 increased DSBs in subtelomeric BESs
Because most VSG switchers arose from HR-mediated events, we wondered whether depletion of TbTIF2 led to more DSBs in subtelomeric BESs. We performed a Ligation-Mediated PCR (LMPCR) assay to directly detect DSBs along BESs10,43. In this assay, a double-stranded adaptor with a blunt end and a staggered end was ligated to genomic DNA at the DSB followed by PCR using locus-specific primers and Southern analysis (Figure 4A). Blunt-ended broken DNA can be ligated to the blunt end of the adaptor directly, while staggered broken DNA ends can be converted to blunt ends by T4 DNA polymerase before ligation. The staggered end of the adaptor and single-stranded DNA breaks are not expected to be ligated in this assay53.
Depletion of TbTIF2 led to increased amount of DSBs at subtelomeric BES regions. (A) The principle of LMPCR. LMPCR analyses were performed in S/TIF2i clone A14 (B-D) and S/v (E) cells. The LMPCR products were hybridized with VSG2 (B), VSG21 (C), Tb427.9.9970 (which encodes a putative small nuclear RNA gene activation protein 50, SNAP50) (D), and 70 bp repeat (E) probes. In each panel of this figure and Figure 6, the Ethidium bromide (EtBr)-stained LMPCR products are shown at the top, the Southern blot result is shown in the middle, and the PCR products using primers specific to TbRAP1 are shown at the bottom as a loading control. The amounts of input genomic DNA, either treated (+) or untreated (−) with T4 DNA polymerase, were marked on top of each lane. Molecular weight markers (in kb unless otherwise indicated) are labeled on the left. (F) The signal intensities of LMPCR products (treated with T4 DNA polymerase) were quantified by ImageQuant from Southern blots exposed to phosphorimagers. The fold changes were calculated by dividing post TbTIF2 depletion value by preinduction value. The average was calculated from at least three independent experiments. Error bars represent standard deviation. P-values (unaired t-tests) were calculated using Graphpad Prism between values at each subtelomeric locus and that at Tb427.9.9970. Asterisks indicate significant difference. *P < 0.05; **P < 0.01; ***P < 0.001.
Following induction of TbTIF2 RNAi for 24 h, we detected substantially more LMPCR products at BES promoter (Supplementary information, Figure S7A) and 70 bp repeat regions (Supplementary information, Figure S7B), indicating that TbTIF2 is important for subtelomere integrity. Since all BESs have nearly identical promoter and 70 bp repeat sequences33, these results did not distingusih whether DSBs are in the active or silent BESs. We therefore used specific probes to detect DSBs at active or silent single-copy gene loci. Because LMPCR is semiquantitative, it is unfair to compare the absolute amount of DSBs at different loci due to possible different PCR amplification efficiencies and different probe hybridization efficiencies. We therefore estimated the change of DSB amount before and after depletion of TbTIF2 at individual locus, then compared whether TbTIF2 has similar effects at different loci. To avoid variation that resulted from different RNAi inductions, we performed LMPCR at two pairs of active and silent loci using genomic DNA isolated from the same induction experiments. We observed increased amount of LMPCR products following TbTIF2 depletion at the active VSG2 (Figure 4B) and the silent VSG21 (Figure 4C) loci. ΨBES1 and ΨBES11 are unique VSG pseudogenes in the active VSG2-containing BES1 and the silent VSG16-containing BES11, respectively (Supplementary information, Figure S7I, top)33,45. We observed increased LMPCR products at ΨBES1 (Supplementary information, Figure S7C) and ΨBES11 (Supplementary information, Figure S7D) loci, too. Quantification of LMPCR signals (in ImageQuant) from at least three sets of independent experiments showed similar fold of increase at all four loci (Figure 4F), indicating that TbTIF2 depletion led to similar increases in DSB levels in both active and silent BESs.
Subsequently, we performed the LMPCR analysis in an independent TbTIF2 RNAi strain (9/TIF2i) that expresses VSG9 and carries BSD at the active BES promoter and PUR at the silent VSG2-containing BES promoter (Supplementary information, Figure S7I, bottom)45. Again, we detected more LMPCR products at both loci after depletion of TbTIF2 (Supplementary information, Figure S7E and S7F), indicating that this phenotype was not specific for any particular VSG expresser. In contrast, approximately the same levels of LMPCR products were detected at a random single-copy chromosomal locus (Tb427.9.9970) before and after TbTIF2 depletion (Figure 4D and 4F), suggesting that TbTIF2 influences DSB amount primarily at subtelomeric regions. In most cases, blunt-ended DSBs represented a small fraction of all breaks (Figure 4 and Supplementary information, Figure S7, compare + and −T4 DNA polymerase). TbTIF2 depletion induced both blunt and stagger ended DSBs. As a control, S/v cells were examined the same way and we did not detect elevated LMPCR products at 70 bp repeats (Figure 4E), the ΨBES1 locus (Supplementary information, Figure S7G), or BES promoters (Supplementary information, Figure S7H). The increased DSB amount in TbTIF2 depleted cells could result from increased DSB formation or decreased DSB repair. Our current assay cannot differentiate between these two possibilities. Nevertheless, our data demonstrate that TbTIF2 plays an important role in subtelomere integrity maintenance.
We also examined whether depletion of TbTIF2 altered chromatin structure by the Formaldehyde-Assisted Isolation of Regulatory Elements (FAIRE) analysis54. In this assay, chromatin is fixed by formaldehyde. After sonication, nucleosome-free DNA fragments are enriched in the phenol-chloroform extracted fraction. We compared the FAIRE-extracted DNA before and after TbTIF2 depletion at several subtelomeric VSG loci, BES promoters, and a random chromosomal locus (Tb427.2.2440) in S/TIF2i, 2/TIF2i, and S/v cells. We only observed a mild change in FAIRE-extracted DNA before and after depletion of TbTIF2 at some VSG loci (Supplementary information, Figure S5F). Thus, TbTIF2 depletion does not affect chromatin structure significantly, suggesting that such changes are unlikely the main reason for elevated VSG switching frequency.
Depletion of TbTIF2 significantly increases association of TbRAD51 with the active BES
TbRAD51 and some of its paralogues are required for most VSG switching events38,39. Since TbTIF2 depletion elevated the DSB amount at BESs, we hypothesized that more TbRAD51 might associate with BESs. To test this, we first performed immunofluorescence using the TbRAD51 antibody. TbRAD51 forms distinct foci in cell nuclei in response to DNA damage55. Indeed, in WT SM/TbTIF2-F2H cells treated with phleomycin, we observed large, bright TbRAD51 foci in the nucleus (Supplementary information, Figure S8A, right). In 2/TIF2i cells treated with doxycycline, we observed similar distinct TbRAD51 foci in the nucleus (Supplementary information, Figure S8B, right), confirming that depletion of TbTIF2 led to an elevated amount of DSBs. In contrast, in untreated 2/TIF2i cells (Supplementary information, Figure S8B, left) or in WT cells treated with or without doxycycline (Supplementary information, Figure S8A, middle and left), TbRAD51 did not form distinct large foci in the nucleus but gave a hazy staining pattern. A significant increase in the number of cells that have distinct TbRAD51 foci was seen when TbTIF2 was depleted (Figure 5A). Therefore, depletion of TbTIF2 resulted in a DNA damage response-like phenotype where TbRAD51 forms distinct foci in the nucleus.
Depletion of TbTIF2 led to increased TbRAD51 association with telomeric and subtelomeric chromatin. (A) Quantification of cells with punctate nuclear TbRAD51 foci at various time points after induction of TbTIF2 RNAi in S/TIF2i A14 cells. Number of cells counted was indicated in the parentheses. Averages were calculated from three independent experiments. Numbers on columns indicate P-values (unpaired t-tests) compared to the 0 h value. (B, C) ChIP analysis using the TbRAD51 antibody or rabbit IgG. ChIP products and input DNA were hybridized with either a TTAGGG or a 50 bp repeat probe in Southern blotting, which were exposed to a phosphorimager. The hybridization signal intensity was quantified by ImageQuant, and averages were calculated from four independent experiments (B). ChIP products were analyzed by qPCR using primer pairs specific to various loci (Supplementary information, Table S1). Averages were calculated from 5-10 independent experiments (C). In both B and C, numbers on top of brackets indicate P-values (unpaired t-tests) comparing TbRAD51 ChIP values before and after TbTIF2 depletion.
To further determine whether TbRAD51 exhibited an increased association with telomeres and subtelomeres in TbTIF2-depleted cells, we performed ChIP analysis using the TbRAD51 antibody in S/TIF2i cells. Depletion of TbTIF2 led to an increase in the association of TbRAD51 with TTAGGG and 50 bp repeats, suggesting that TbTIF2 dysfunction led to more DSBs at telomeres and subtelomeres (Figure 5B). Our T. brucei strain has nearly 250 telomeres due to its ∼100 minichromosomes in addition to 11 pairs of megabase chromosomes and a few intermediate chromosomes. However, there are only nearly 20 BESs. The level of increase in TbRAD51 association with TTAGGG and 50 bp repeats is similar (Figure 5B), suggesting that TbRAD51 might be recruited to individual BESs at higher concentration than to individual telomeres that do not have any adjacent BES. To achieve better resolution, we performed ChIP experiments using the TbRAD51 antibody followed by qPCR using primers specific to BES promoters, PUR and VSG2 in the active BES, and VSG21, VSG16, and ΨBES11 in silent BESs (Supplementary information, Figure S7I, top)45. We observed a significant increase in the association of TbRAD51 with BES promoters, PUR, and VSG2 (Figure 5C), confirming that TbRAD51 associated more with subtelomeric BESs upon TbTIF2 depletion. Surprisingly, this increase was mostly restricted in the active BES, since the association of TbRAD51 with the silent VSG21 was not significantly changed (Figure 5C). Although more TbRAD51 associated with the silent VSG16 and ΨBES11, the increase was not as dramatic as that at the active VSG2 locus (Figure 5C). Since we observed similar increases in DSB amounts at both active and silent BESs upon depletion of TbTIF2 (Figure 4F), this observation suggests that TbRAD51 preferentially associates with DSBs in the active BES. No significant increase was observed for the association of TbRAD51 with the chromosome internal tubulin array or the Tb427.9.9970 locus (Figure 5C), supporting our conclusion that depletion of TbTIF2 does not induce DSBs at chromosome internal loci.
Subtelomeric DSBs in TbTIF2-depleted cells are further increased by TbRAD51 disruption
Upon depletion of TbTIF2, DSB levels increased at subtelomeric BESs, as did association of TbRAD51 with the active BES, indicating that TbRAD51 associates with and repairs DSBs in the active BES, which can lead to VSG switching, while DSBs in silent BESs may not be repaired by TbRAD51-mediated HR. To test this hypothesis, we deleted both alleles of TbRAD51 in S/TIF2i cells and tried to perform the VSG switching assay. However, these cells were less healthy than the S/TIF2i cells, and even a transient induction of TbTIF2 RNAi resulted in an irreversible cell growth arrest, preventing us from carrying out switching assays.
We therefore performed LMPCR in the S/TIF2i/TbRAD51Δ cells. Similar to S/TIF2i cells, depletion of TbTIF2 increased DSB levels at BES promoters (Supplementary information, Figure S9A), 70 bp repeats (Supplementary information, Figure S9B), active ΨBES1 (Figure 6A), silent ΨBES11 (Figure 6B), active VSG2 (Supplementary information, Figure S9C), and silent VSG21 (Supplementary information, Figure S9D) loci. More strikingly, the amount of DSBs at subtelomeric BESs were much higher when TbRAD51 was deleted than when it was not (Figure 6 and Supplementary information, S9), indicating that loss of TbRAD51 exacerbated the increased DSB phenotype in TbTIF2-depleted cells. To avoid variations between different inductions and LMPCR experiments, we always induced S/TIF2i and S/TIF2i/TbRAD51Δ cells at the same time using exactly the same conditions. By quantifying the signal intensity of LMPCR products, we found that after depletion of TbTIF2, the amount of DSBs in the TbRAD51Δ background was 2.7-fold and 4.5-fold of that in the TbRAD51 wild-type background at the active VSG2 and ΨBES1 locus, respectively, but only 1.6-fold and 1.5-fold at the silent VSG21 and ΨBES11 locus, respectively (Figure 6C). This suggests that DSBs in the active BES were preferentially repaired by TbRAD51-dependent mechanism.
More subtelomeric DSBs persisted when TbRAD51 was deleted in TbTIF2-depleted cells. The LMPCR products from S/TIF2i A14 and S/TIF2i/TbRAD51Δ C6 cells were hybridized with the ΨBES1 (A) or ΨBES11 (B) probe. (C) The signal intensities of LMPCR products (treated with T4 DNA polymerase) were quantified by ImageQuant from Southern blots exposed to phosphorimagers. The fold changes were calculated by dividing post TbTIF2 depletion value in S/TIF2i/ΔTbRAD51 cells by that in S/TIF2i cells on the same gel. The average was calculated from at least three independent experiments. Error bars represent standard deviation. P-values (unpaired t-tests) between pairs of loci were indicated.
Discussion
Telomere proteins and antigenic variation
Our current and previous studies showed that T. brucei telomere proteins play important roles in antigenic variation, monoallelic VSG expression and VSG switching, which are critical pathogenesis mechanisms in T. brucei45,46. Among the three known T. brucei telomere proteins, TbTIF2 suppresses VSG switching but only mildly affects BES promoters that are close to telomeres. This is different from TbRAP1, depletion of which led to dramatic subtelomeric VSG derepression45,46. In contrast, TbTIF2's effect on VSG silencing is similar to that of TbTRF, which does not affect BES-linked VSG silencing45. Although both TbTIF2 and TbRAP1 interact with TbTRF, the interaction between TbTIF2 and TbTRF is much stronger than that between TbRAP1 and TbTRF based on yeast two-hybrid and co-IP analyses (this study and45). Our observations suggest that TbTIF2 may function in the same pathway as TbTRF but not TbRAP1. Further studies of functions of TbTRF and TbRAP1 in VSG switching would shed more light on the interplay between T. brucei telomere proteins in antigenic variation.
Telomere proteins in subtelomere integrity and stability maintenance
In this study, we found that depletion of TbTIF2 increased DSBs at subtelomeres and elevated recombination events that lead to VSG switching, demonstrating that TbTIF2, a telomere protein, is important for maintaining subtelomere integrity and stability.
More TbRAD51 associates with telomeric DNA upon TbTIF2 depletion, indicating that TbTIF2 depletion led to more telomeric DSBs. This is consistent with the notion that telomere proteins are important for protecting chromosome ends11. Loss of mammalian TIN2 led to both chromatid-type and chromosome-type telomere fusions51. It is possible that TbTIF2 depletion results in similar telomere fusions and subsequent breakage-fusion-bridge cycle, which can result in DNA amplification and large terminal deletions57. Some of the ES loss + in situ switching that we observed might have been a consequence of the breakage-fusion-bridge cycle, although telomere fusions have not been reported in T. brucei. It was reported previously that a considerable percent (22%) of cells with DSBs downstream of the active VSG lose the whole active BES10. TbTIF2 depletion-induced DSBs were detected at subtelomeres including the region downstream of the active VSG2, which may also explain why we obtained ES loss + in situ switchers.
TbTIF2 may contribute to subtelomere stability through other means. First, TbTIF2 may be required for proper telomeric DNA replication, as some telomere proteins do in other organisms23,24,26. Fork stalling in the absence of TbTIF2 could increase DSB formation at the telomere vicinity, and increased DNA topological stress could also lead to more DSBs at subtelomeres. Second, TbTIF2 may play an important role in telomere cohesion formation/maintenance. Mammalian TIN2 interacts with SA127,28, an ortholog of SCC3 and a telomere-specific cohesin subunit that is important for sister telomere cohesion. TbTIF2 is a functional homolog of TIN2 and may have similar functions to TIN2. Defective sister telomere pairing due to TbTIF2 depletion could lead to low efficiency of DNA damage repair via the sister chromatid, resulting in more DSBs in subtelomeric BESs. This in turn could lead to loss of terminal chromosome fragments. In addition, a lack of sister telomere pairing is expected to allow more flexible pairing and HR between non-sister telomeres, which in turn may result in more frequent VSG switching. The predominant VSG switching pathway is ES GC/ES loss + in situ switching in TbTIF2-depleted cells, which is consistent with this hypothesis. Interestingly, partial depletion of TbSCC1 or other cohesin subunits TbSMC1 and TbSCC3 also led to premature dissociation of sister chromatids at the active BES and increased VSG switching frequency58, although in situ switching appears to be the preferred pathway in cohesion-defective cells, which is not exactly the same as in TbTIF2-depleted cells, where a significant portion of the VSG switchers arose from ES loss + in situ or ES GC switching. Our first model that TbTIF2 is important for telomeric DNA replication suggests that more DSB formation is the reason for more subtelomeric DSBs in TbTIF2-depleted cells, while the second model suggests that less DSB repair is the reason. However, both factors could contribute to the observed phenotypes. More investigations are necessary to further discern these two underlying mechanisms.
TbRAD51-dependent HR-mediated DNA damage repair and VSG switching
RAD51 is a major player in HR-mediated DNA damage repair37. VSG switching is mostly dependent on TbRAD5139, its paralogue TbRAD51-338, and its interacting factor BRCA240. In addition, HR-mediated GC events are the preferred mechanism for VSG switching in our lab strain39,42. Depletion of TbTIF2 led to more DSBs at subtelomeric BESs and an increased TbRAD51 association with subtelomeric chromatin. TbRAD51 disruption further increased subtelomeric DSBs in TbTIF2-depleted cells. These observations indicate that TbRAD51-mediated HR is a major pathway for repair of the elevated DSBs at subtelomeres. Repair of these induced DSBs presumably contributed to the increased VSG switching frequency in TbTIF2-depleted cells. In S/TIF2i/TbRAD51Δ cells, the DSBs induced by TbTIF2 depletion were not repaired due to the lack of TbRAD51, which is likely the reason for subsequent cell lethality.
TbTIF2 depletion led to increased TbRAD51 association with subtelomeric chromatin, which was more at active loci than at silent loci. In addition, we detected stronger increases of DSBs at the active loci than at the silent loci in S/TIF2i/TbRAD51Δ cells. The extra DSBs detected in the TbRAD51Δ background were presumably repaired by TbRAD51 in S/TIF2i cells. TbTIF2 depletion induced similar level of DSBs at the active and silent loci. Therefore, TbRAD51 preferentially repairs DSBs in the active BES. The active BES differs from silent BESs in that it is actively transcribed by RNA polymerase I59. In addition, the active BES forms an open chromatin structure depleted of most nucleosomes, while silent BESs are packed with nucleosomes60,61. Our data suggest that the transcriptional status and/or the chromatin structure of the DSB site may influence the choice of DSB repair mechanism. In yeast, two independent HR pathways are involved in telomere maintenance in telomerase-negative cells62: amplification of subtelomeric Y′ elements is RAD51-dependent, while amplification of telomeric repeats is RAD50-dependent. In T. brucei, DSBs at silent BESs can be repaired by TbRAD51-independent microhomology-mediated end joining10 or possibly other mechanisms such as TbRAD50-dependent pathway. However, more studies are necessary to test this hypothesis. A RAD50 DNA repair-like protein (Tb427tmp.01.0340) has been annotated in TriTrypDB, although its characterization has not been reported. Alternatively, DSBs in silent BESs are well tolerated and may not be repaired, as loss of BES following DSB induction in silent BESs has been observed frequently10.
Telomere protein evolution from protozoa to mammals
We have identified TbTIF2 as a functional homolog of TIN2. TbTIF2 does not have any telomere DNA binding domains, thus it relies on TbTRF to localize to the telomere, which is presumably essential for its function in subtelomere integrity and stability maintenance. It is worth noting that human TRF2 interacts with Apollo in the same way that human TRF1 interacts with TIN2. Apollo and TIN2 share a conserved TRF-interacting motif Y/F-X-L-X-P, where X can be any residue49. Interestingly, TbTIF2 has a Y-F-L-C-P motif at its C-terminus (Supplementary information, Figure S1D). However, yeast two-hybrid analysis showed that the C-terminal half of TbTIF2 did not interact with TbTRF. A careful examination of the TbTRF sequence showed that TbTRF lacks the conserved region in the TRFH domain of human TRF1 and TRF2 that is essential for recognizing the Y/F-X-L-X-P motif48, explaining why the C-terminus of TbTIF2 does not interact with TbTRF. Therefore, although the TIN2-TRF/TbTIF2-TbTRF interaction is conserved, the detailed interaction surfaces on the two protein pairs have changed through evolution. This makes the interaction domain between TbTRF and TbTIF2 an attractive target for anti-parasite agents, as the interaction is specific for T. brucei and not present in its host.
Materials and Methods
ChIP
ChIP was performed according to64. ChIP products were hybridized with a TTAGGG or a 50-bp repeat probes in slot blot analysis. Alternatively, qPCR using primers specific to various BES loci (Supplementary information, Table S1) was performed to detect unique sequences in ChIP products.
LMPCR
LMPCR were performed according to43. More details are described in Supplementary information, Data S1. Primers used in LMPCR are listed in Supplementary information, Table S2.
VSG switching assay
Switching assays were performed according to42. Additional details are described in Supplementary information, Data S1. Detailed switcher analyses are listed in Supplementary information, Tables S3-S6.
FAIRE
FAIRE was performed according to46 except that the amount of isolated DNA was estimated by qPCR using SsoAdvanced SYBR Green in a CFX96 Connect (Bio-Rad). The FAIRE result was calculated by dividing FAIRE-extracted DNA amount with the input DNA amount, and the fold changes in FAIRE results between TbTIF2-depleted and uninduced cells were plotted.
qRT-PCR
qRT-PCR was performed as described previously45 except that the samples were analyzed using SsoAdvanced SYBR Green in a CFX96 Connect (Bio-Rad). Primers used in qRT-PCR are described in the previous study45.
Yeast two-hybrid analyses
Yeast two-hybrid analyses were carried out as described in48.
Immunofluorescence
Immunofluorescence was carried out as described in48. Cell images were captured by a DeltaVision image restoration microscope (Applied Precision/Olympus), deconvoluted by using measured point spread functions, and edited with Photoshop.
References
Gabriels J, Beckers MC, Ding H, et al. Nucleotide sequence of the partially deleted D4Z4 locus in a patient with FSHD identifies a putative gene within each 3.3 kb element. Gene 1999; 236:25–32.
van der Maarel SM, Frants RR, Padberg GW . Facioscapulohumeral muscular dystrophy. Biochim Biophys Acta 2007; 1772:186–194.
DeScipio C . The 6p subtelomere deletion syndrome. Am J Med Genet C Semin Med Genet 2007; 145C:377–382.
Stewart DR, Kleefstra T . The chromosome 9q subtelomere deletion syndrome. Am J Med Genet C Semin Med Genet 2007; 145C:383–392.
Trask BJ, Friedman C, Martin-Gallardo A, et al. Members of the olfactory receptor gene family are contained in large blocks of DNA duplicated polymorphically near the ends of human chromosomes. Hum Mol Genet 1998; 7:13–26.
Li B . Telomere components as potential therapeutic targets for treating microbial pathogen infections. Front Oncol 2012; 2:156.
Mefford HC, Trask BJ . The complex structure and dynamic evolution of human subtelomeres. Nat Rev Genet 2002; 3:91–102.
Linardopoulou EV, Williams EM, Fan Y, Friedman C, Young JM, Trask BJ . Human subtelomeres are hot spots of interchromosomal recombination and segmental duplication. Nature 2005; 437:94–100.
Teytelman L, Eisen MB, Rine J . Silent but not static: accelerated base-pair substitution in silenced chromatin of budding yeasts. PLoS Genet 2008; 4:e1000247.
Glover L, Alsford S, Horn D . DNA break site at fragile subtelomeres determines probability and mechanism of antigenic variation in African trypanosomes. PLoS Pathog 2013; 9:e1003260.
Stewart JA, Chaiken MF, Wang F, Price CM . Maintaining the end: roles of telomere proteins in end-protection, telomere replication and length regulation. Mutat Res 2012; 730:12–19.
de Lange T . Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev 2005; 19:2100–2110.
Chong L, van Steensel B, Broccoli D, et al. A human telomeric protein. Science 1995; 270:1663–1667.
Broccoli D, Smogorzewska A, Chong L, de Lange T . Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nat Genet 1997; 17:231–235.
Bilaud T, Brun C, Ancelin K, Koering CE, Laroche T, Gilson E . Telomeric localization of TRF2, a novel human telobox protein. Nat Genet 1997; 17:236–239.
Xin H, Liu D, Wan M, et al. TPP1 is a homologue of ciliate TEBP-beta and interacts with POT1 to recruit telomerase. Nature 2007; 445:559–562.
Kim SH, Kaminker P, Campisi J . TIN2, a new regulator of telomere length in human cells. Nat Genet 1999; 23:405–412.
Houghtaling BR, Cuttonaro L, Chang W, Smith S . A dynamic molecular link between the telomere length regulator trf1 and the chromosome end protector TRF2. Curr Biol 2004; 14:1621–1631.
Ye JZ, Hockemeyer D, Krutchinsky AN, et al. POT1-interacting protein PIP1: a telomere length regulator that recruits POT1 to the TIN2/TRF1 complex. Genes Dev 2004; 18:1649–1654.
Ye JZ, Donigian JR, van Overbeek M, et al. TIN2 binds TRF1 and TRF2 simultaneously and stabilizes the TRF2 complex on telomeres. J Biol Chem 2004; 279:47264–47271.
Kim SH, Beausejour C, Davalos AR, Kaminker P, Heo SJ, Campisi J . TIN2 mediates functions of TRF2 at human telomeres. J Biol Chem 2004; 279:43799–43804.
Denchi EL, de Lange T . Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 2007; 448:1068–1071.
Sfeir A, Kosiyatrakul ST, Hockemeyer D, et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 2009; 138:90–103.
Ye J, Lenain C, Bauwens S, et al. TRF2 and apollo cooperate with topoisomerase 2alpha to protect human telomeres from replicative damage. Cell 2010; 142:230–242.
Muraki K, Nabetani A, Nishiyama A, Ishikawa F . Essential roles of Xenopus TRF2 in telomere end protection and replication. Genes Cells 2011; 16:728–739.
Miller KM, Rog O, Cooper JP . Semi-conservative DNA replication through telomeres requires Taz1. Nature 2006; 440:824–828.
Canudas S, Houghtaling BR, Kim JY, Dynek JN, Chang WG, Smith S . Protein requirements for sister telomere association in human cells. EMBO J 2007; 26:4867–4878.
Canudas S, Smith S . Differential regulation of telomere and centromere cohesion by the Scc3 homologues SA1 and SA2, respectively, in human cells. J Cell Biol 2009; 187:165–173.
Canudas S, Houghtaling BR, Bhanot M, et al. A role for heterochromatin protein 1gamma at human telomeres. Genes Dev 2011; 25:1807–1819.
Barry JD, McCulloch R . Antigenic variation in trypanosomes: enhanced phenotypic variation in a eukaryotic parasite. Adv Parasitol 2001; 49:1–70.
Berriman M, Ghedin E, Hertz-Fowler C, et al. The genome of the African trypanosome Trypanosoma brucei. Science 2005; 309:416–422.
de Lange T, Borst P . Genomic environment of the expression-linked extra copies of genes for surface antigens of Trypanosoma brucei resembles the end of a chromosome. Nature 1982; 299:451–453.
Hertz-Fowler C, Figueiredo LM, Quail MA, et al. Telomeric expression sites are highly conserved in Trypanosoma brucei. PLoS One 2008; 3:e3527.
Navarro M, Cross GAM . DNA rearrangements associated with multiple consecutive directed antigenic switches in Trypanosoma brucei. Mol Cell Biol 1996; 16:3615–3625.
Chaves I, Zomerdijk J, Dirks-Mulder A, Dirks RW, Raap AK, Borst P . Subnuclear localization of the active variant surface glycoprotein gene expression site in Trypanosoma brucei. Proc Natl Acad Sci USA 1998; 95:12328–12333.
Morrison LJ, Marcello L, McCulloch R . Antigenic variation in the African trypanosome: molecular mechanisms and phenotypic complexity. Cell Microbiol 2009; 11:1724–1734.
Holthausen JT, Wyman C, Kanaar R . Regulation of DNA strand exchange in homologous recombination. DNA Repair (Amst) 2010; 9:1264–1272.
Proudfoot C, McCulloch R . Distinct roles for two RAD51-related genes in Trypanosoma brucei antigenic variation. Nucleic Acids Res 2005; 33:6906–6919.
McCulloch R, Barry JD . A role for RAD51 and homologous recombination in Trypanosoma brucei antigenic variation. Genes Dev 1999; 13:2875–2888.
Hartley CL, McCulloch R . Trypanosoma brucei BRCA2 acts in antigenic variation and has undergone a recent expansion in BRC repeat number that is important during homologous recombination. Mol Microbiol 2008; 68:1237–1251.
Kim HS, Cross GAM . Identification of Trypanosoma brucei RMI1/BLAP75 homologue and its roles in antigenic variation. PLoS One 2011; 6:e25313.
Kim HS, Cross GAM . TOPO3alpha influences antigenic variation by monitoring expression-site-associated VSG switching in Trypanosoma brucei. PLoS Pathog 2010; 6:e1000992.
Boothroyd CE, Dreesen O, Leonova T, et al. A yeast-endonuclease-generated DNA break induces antigenic switching in Trypanosoma brucei. Nature 2009; 459:278–281.
Ginger ML, Blundell PA, Lewis AM, Browitt A, Gunzl A, Barry JD . Ex vivo and in vitro identification of a consensus promoter for VSG genes expressed by metacyclic-stage trypanosomes in the tsetse fly. Eukaryot Cell 2002; 1:1000–1009.
Yang X, Figueiredo LM, Espinal A, Okubo E, Li B . RAP1 is essential for silencing telomeric variant surface glycoprotein genes in Trypanosoma brucei. Cell 2009; 137:99–109.
Pandya UM, Sandhu R, Li B . Silencing subtelomeric VSGs by Trypanosoma brucei RAP1 at the insect stage involves chromatin structure changes. Nucleic Acids Res 2013; 41:7673–7682.
Hovel-Miner GA, Boothroyd CE, Mugnier M, Dreesen O, Cross GAM, Papavasiliou FN . Telomere length affects the frequency and mechanism of antigenic variation in Trypanosoma brucei. PLoS Pathog 2012; 8:e1002900.
Li B, Espinal A, Cross GAM . Trypanosome telomeres are protected by a homologue of mammalian TRF2. Mol Cell Biol 2005; 25:5011–5021.
Chen Y, Yang Y, van Overbeek M, et al. A shared docking motif in TRF1 and TRF2 used for differential recruitment of telomeric proteins. Science 2008; 319:1092–1096.
Wirtz E, Leal S, Ochatt C, Cross GAM . A tightly regulated inducible expression system for dominant negative approaches in Trypanosoma brucei. Mol Biochem Parasitol 1999; 99:89–101.
Takai KK, Kibe T, Donigian JR, Frescas D, de Lange T . Telomere protection by TPP1/POT1 requires tethering to TIN2. Mol Cell 2011; 44:647–659.
Kolev NG, Ramey-Butler K, Cross GAM, Ullu E, Tschudi C . Developmental progression to infectivity in trypanosoma brucei triggered by an rna-binding protein. Science 2012; 338:1352–1353.
Arudchandran A, Bernstein RM, Max EE . Single-stranded DNA breaks adjacent to cytosines occur during Ig gene class switch recombination. J Immunol 2004; 173:3223–3229.
Giresi PG, Lieb JD . Isolation of active regulatory elements from eukaryotic chromatin using FAIRE (Formaldehyde Assisted Isolation of Regulatory Elements). Methods 2009; 48:233–239.
Glover L, McCulloch R, Horn D . Sequence homology and microhomology dominate chromosomal double-strand break repair in African trypanosomes. Nucleic Acids Res 2008; 36:2608–2618.
Riethman H . Human telomere structure and biology. Annu Rev Genomics Hum Genet 2008; 9:1–19.
Murnane JP . Telomeres and chromosome instability. DNA Repair (Amst) 2006; 5:1082–1092.
Landeira D, Bart JM, Van Tyne D, Navarro M . Cohesin regulates VSG monoallelic expression in trypanosomes. J Cell Biol 2009; 186:243–254.
Gunzl A, Bruderer T, Laufer G, et al. RNA polymerase I transcribes procyclin genes and variant surface glycoprotein gene expression sites in Trypanosoma brucei. Eukaryot Cell 2003; 2:542–551.
Figueiredo LM, Cross GAM . Nucleosomes are depleted at the VSG expression site transcribed by RNA polymerase I in African trypanosomes. Eukaryot Cell 2010; 9:148–154.
Stanne TM, Rudenko G . Active VSG expression sites in Trypanosoma brucei are depleted of nucleosomes. Eukaryot Cell 2010; 9:136–147.
Chen Q, Ijpma A, Greider CW . Two survivor pathways that allow growth in the absence of telomerase are generated by distinct telomere recombination events. Mol Cell Biol 2001; 21:1819–1827.
Kim SH, Davalos AR, Heo SJ, et al. Telomere dysfunction and cell survival: roles for distinct TIN2-containing complexes. J Cell Biol 2008; 181:447–460 .
Siegel TN, Hekstra DR, Kemp LE, et al. Four histone variants mark the boundaries of polycistronic transcription units in Trypanosoma brucei. Genes Dev 2009; 23:1063–1076.
Vanhamme L, Poelvoorde P, Pays A, Tebabi P, Van Xong H, Pays E . Differential RNA elongation controls the variant surface glycoprotein gene expression sites of Trypanosoma brucei. Mol Microbiol 2000; 36:328–340.
Wang Z, Morris JC, Drew ME, Englund PT . Inhibition of Trypanosoma brucei gene expression by RNA interference using an integratable vector with opposing T7 promoters. J Biol Chem 2000; 275:40174–40179.
de Lange T, Shiue L, Myers RM, et al. Structure and variability of human chromosome ends. Mol Cell Biol 1990; 10:518–527.
Acknowledgements
We thank Dr Hee-Sook Kim and Dr George AM Cross for sending us the HSTB261 cells, the TbRAD51 knockout and Cre expression constructs, and VSG antibodies. We are very grateful to Dr Richard McCulloch for providing us with the TbRAD51 antibody, and Dr Keith Gull for providing tubulin antibodies. We thank The Rockefeller University proteomics resource center and particularly Dr Joseph Fernandez for carrying out the mass spectrometry analysis. We thank Dr Valentin Börner, Dr Aaron Severson, and Li lab members for their comments on the manuscript. We also thank Vishal Nanavaty for technical support. This work is in part supported by an NIH grant (AI066095), the CSU 2010 Faculty Research and Development award to B Li, and Center for Gene Regulation in Health and Disease at CSU.
Author information
Authors and Affiliations
Corresponding author
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Identification of TbTIF2 as a TbTRF-interacting factor. (PDF 169 kb)
Supplementary information, Figure S2
TbTIF2 depletion has mild effects on subtelomeric VSG silencing. (PDF 136 kb)
Supplementary information, Figure S3
FACS analysis of TbTIF2 depleted cells. (PDF 100 kb)
Supplementary information, Figure S4
Diagram depicting the principle of the VSG switching assay. (PDF 172 kb)
Supplementary information, Figure S5
TbTIF2 depletion does not affect telomere length or subtelomeric chromatin structure significantly. (PDF 13755 kb)
Supplementary information, Figure S6
Validating VSG switching mechanisms in selected switchers. (PDF 324 kb)
Supplementary information, Figure S7
LMPCR analyses in S/TIF2i clone A14 (A-D), 9/TIF2i Clone C3 (E-F), and S/v (G-H) cells. (PDF 443 kb)
Supplementary information, Figure S8
Depletion of TbTIF2 resulted in nuclear punctate TbRAD51 foci formation. (PDF 411 kb)
Supplementary information, Figure S9
LMPCR analyses in S/TIF2i clone A14 and S/TIF2i/TbRAD51Δ clone C6. (PDF 200 kb)
Supplementary information, Table S1
List of primers used in qPCR analysis in TbRAD51 ChIP experiments (PDF 12 kb)
Supplementary information, Table S2
Primers used in LMPCR analyses. (PDF 13 kb)
Supplementary information, Table S3
S/TIF2i Switcher phenotype and genotype characterization (PDF 108 kb)
Supplementary information, Table S4
Parent switcher phenotype and genotype characterization (PDF 27 kb)
Supplementary information, Table S5
S/v switcher phenotype and genotype characterization. (PDF 18 kb)
Supplementary information, Table S6
Phenotype and genotype characterization of switchers isolated from uninduced S/TIF2i A14 cells (PDF 14 kb)
Supplementary information, Data S1
Materials and Methods (PDF 111 kb)
Rights and permissions
About this article
Cite this article
Jehi, S., Wu, F. & Li, B. Trypanosoma brucei TIF2 suppresses VSG switching by maintaining subtelomere integrity. Cell Res 24, 870–885 (2014). https://doi.org/10.1038/cr.2014.60
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2014.60