Abstract
The spike glycoprotein (S) of recently identified Middle East respiratory syndrome coronavirus (MERS-CoV) targets the cellular receptor, dipeptidyl peptidase 4 (DPP4). Sequence comparison and modeling analysis have revealed a putative receptor-binding domain (RBD) on the viral spike, which mediates this interaction. We report the 3.0 Å-resolution crystal structure of MERS-CoV RBD bound to the extracellular domain of human DPP4. Our results show that MERS-CoV RBD consists of a core and a receptor-binding subdomain. The receptor-binding subdomain interacts with DPP4 β-propeller but not its intrinsic hydrolase domain. MERS-CoV RBD and related SARS-CoV RBD share a high degree of structural similarity in their core subdomains, but are notably divergent in the receptor-binding subdomain. Mutagenesis studies have identified several key residues in the receptor-binding subdomain that are critical for viral binding to DPP4 and entry into the target cell. The atomic details at the interface between MERS-CoV RBD and DPP4 provide structural understanding of the virus and receptor interaction, which can guide development of therapeutics and vaccines against MERS-CoV infection.
Similar content being viewed by others
Introduction
A novel coronavirus, recently named the Middle East respiratory syndrome coronavirus (MERS-CoV), was first identified in humans in the Middle East in 2012, and later in several European countries1,2,3. A sizable fraction (> 50%) of the infected patients developed severe respiratory illness and clinical symptoms similar to those seen during outbreak of severe acute respiratory symptom (SARS) caused by the coronavirus SARS-CoV in 20034. In particular, preliminary epidemiology studies suggest human-to-human transmission of this deadly virus, leading to global concern about the potential for a MERS pandemic5,6. Genetic and phylogenetic characterization has shown that MERS-CoV belongs to lineage C of the genus of betacoronavirus and is closely related to Tylonycteris bat coronavirus HKU4 and Pipistrellus bat coronavirus HKU5, although the direct source and reservoirs of MERS-CoV remain uncertain2,7,8. Like other coronaviruses, the MERS-CoV virion utilizes a large surface spike (S) glycoprotein for interaction with and entry into the target cell9,10. The S glycoprotein consists of a globular S1 domain at the N-terminal region, followed by membrane-proximal S2 domain, a transmembrane domain and an intracellular domain11,12,13. Determinants of cellular tropism and interaction with the target cell are within the S1 domain, while mediators of membrane fusion have been identified within the S2 domain11,12. Through co-purification with the MERS-CoV S1 domain, Raj and colleagues9 recently identified that dipeptidyl peptidase 4 (DPP4, also called CD26) functions as a cellular receptor for MERS-CoV. DPP4 does not share any sequence or structural similarities to the previously identified human coronavirus receptors, such as angiotensin-converting enzyme 2 (ACE2) for SARS-CoV14,15 and HCoV-NL6316, or aminopeptidase N (APN) for HCoV-229E17. Like ACE2 and APN, however, DDP4 is expressed on the surface of several cell types, including those found in human airways, and possesses ectopeptidase activity, although this enzymatic function does not appear to be essential for viral entry18. Sequence and modeling analysis of S glycoproteins from several human coronaviruses has revealed a potential receptor-binding domain (RBD) of MERS-CoV10,19. However, given the relatively low degree of homology between S glycoprotein sequences and mechanisms of interaction with distinct cell surface receptors, it is likely that there is a significant variability in structural features among respective RBD receptor pairs. Here, we report the crystal structure of MERS-CoV RBD in complex with human DPP4 extracellular domain at a resolution of 3.0 Å.
Results
Overall structure of the complex
MERS-CoV RBD (E367 to Y606) and soluble DPP4 extracellular domain (S39 to P766) with C-terminal histidine tag were expressed in Sf9 cells and purified by Ni-NTA and size-exclusion chromatography (Supplementary information, Figure S1). The two components were then mixed, and the complex was purified by size-exclusion chromatography (Supplementary information, Figure S1). Crystals belong to the P6122 space group with cell dimensions, a = b = 110.6 Å, c = 527.6 Å, α = β = 90°, γ = 120°. The structure was determined by the molecular replacement method and refined to a resolution of 3.0 Å with Rwork of 19.2% and Rfree of 24.1% (Supplementary information, Table S1). There is one complex of DPP4 extracellular domain with MERS-CoV RBD in the asymmetric unit. The final model consists of residues E382 to C585 of the MERS-CoV RBD, S39 to P766 of the DPP4 extracellular domain, and glycans N-linked to residues N85, N92, N150, N219, N229, N281, N321 and N520 of the DPP4 extracellular domain.
The DPP4 extracellular domain consists of an N-terminal eight-bladed β-propeller domain (S39 to D496) and a C-terminal α/β hydrolase domain (N497 to P766) (Figure 1). The β-propeller domain consists of eight blades, each made up of four antiparallel β-strands (Supplementary information, Figure S2). DPP4 utilizes the blades 4 and 5 to contact the MERS-CoV RBD (Supplementary information, Figure S2). The contact site is far from the hydrolase domain (Figure 1), which is consistent with the previous findings showing that the addition of DPP4 inhibitors sitagliptin, vildagliptin, saxagliptin or P32/98 does not block MERS-CoV entry9.
Overall structure of the complex. DPP4 extracellular domain consists of N-terminal eight-bladed β-propeller domain (green) and C-terminal α/β-hydrolase domain (orange). MERS-CoV RBD contains a core (cyan) and a receptor-binding subdomain (purple). The disulfide bonds are drawn as yellow sticks and the N-linked glycans are drawn as pink sticks.
Structure of MERS-CoV RBD and its comparison with SARS-CoV RBD
The MERS-CoV RBD contains a core subdomain and a receptor-binding subdomain (Figure 1). The core subdomain is a five-stranded antiparallel β sheet (β1, β2, β3, β4 and β9) with two short α helices in the connecting loops (Figure 2C and 2E). Three disulfide bonds, connecting C383 to C407, C425 to C478 and C437 to C585, reside in the core subdomain to maintain the fold (Figure 2E). The receptor-binding subdomain is a four-stranded antiparallel β sheet (β5, β6, β7 and β8), located between strands β4 and β9 of the core domain (Figure 2C and 2E). There is a long loop connecting β6 and β7 strands, which crosses perpendicularly to the β sheet (Figure 2C and 2E). The disulfide bond between C503 and C526 connects this loop with strand β5 (Figure 2E), thereby providing structural support for the β sheet to contact DPP4. Similarly, SARS-CoV RBD also contains the core and receptor-binding subdomains (Figure 2D and 2F)14. Although MERS-CoV and SARS-CoV have low amino acid sequence homology in the RBD (Supplementary information, Figure S3), their core subdomains are structurally similar with an r.m.s.d. of 2.0 Å for 95 aligned Cα atoms. However, clear differences exist in the receptor-binding subdomain between MERS-CoV and SARS-CoV. The former contains 84 residues (V484 to L567, Figure 2A), forming a four-stranded antiparallel β sheet (Figure 2C and 2E), whereas the latter has 68 amino acids (T425 to Q492, Figure 2B), forming a long extended loop with two short antiparallel β strands and one disulfide bond between C467 and C474 (Figure 2D and 2F).
Structural comparison between MERS-CoV RBD and SARS-CoV RBD. Domain structures of MERS-CoV S1 (A) and of SARS-CoV S1 (B). (C) Structure of MERS-CoV RBD. The receptor-binding subdomain is colored in purple and the core subdomain is colored in cyan. (D) Structure of SARS-CoV RBD (PDB code 2AJF). The receptor-binding subdomain is colored in purple and the core subdomain is colored in wheat. (E) Schematic illustration of MERS-CoV RBD topology. β strands are drawn as arrows and α helices are drawn as cylinders. The disulfide bonds are drawn as yellow sticks. (F) Schematic illustration of SARS-CoV RBD topology. β strands are drawn as arrows and α helices are drawn as cylinders. The disulfide bonds are drawn as yellow sticks.
Binding interface
The MERS-CoV RBD binds to the DPP4 β-propeller domain with a buried surface of ∼2 550 Å2. At the interface, a total of 14 residues of MERS-CoV contact with 15 residues of the DPP4 with a distance cutoff of 3.6 Å (Supplementary information, Table S2). The interface consists of two major binding patches. In patch 1 (∼1 250 Å2 buried surface), the C-terminal end of the long loop connecting the β6 and β7 strands contacts the blade 4 of DPP4 (Figure 3A). MERS-CoV acidic residues E536, D537 and D539 at the C-terminal end of the loop form a negatively charged surface, and D539 has salt-bridge interaction with DPP4 basic residue K267 (Figure 3B). MERS-CoV residue Y499 forms hydrogen bond with the DPP4 residue R336 in this patch (Figure 3B). In patch 2 (∼1 300 Å2 buried surface), there is a slightly concaved outer surface at the far end of the MERS-CoV receptor-binding subdomain, formed by the short β6 strand, C-terminal parts of β5 and β7 strands, N-terminal part of β8 strand and the β5-β6 linking loop (Figure 3A). This concaved outer surface accommodates a linker containing a short α helix between blade 4 and blade 5 of DPP4 (Figure 3A). In this patch, a hydrophobic core is found, consisting of MERS-CoV RBD residues L506, W553 and V555, and DPP4 residues L294 and I295, surrounded by hydrophilic MERS-CoV RBD residues D510, E513 and Y540, and DPP4 residues H298, R317 and Q344 (Figure 3C). Among the peripheral hydrophilic residues, MERS-CoV RBD residues D510 and E513 form salt-bridge and hydrogen-bonding interactions with DPP4 residues R317 and Q344, respectively (Figure 3C).
Binding interface. (A) DPP4 contacts the MERS-CoV RBD with its blades 4 and 5 in the N-terminal eight-bladed β-propeller domain. Patch 1 is centered around the C-terminal end of the long linker connecting β6 and β7 strands in MERS-CoV RBD. Patch 2 has a gently concaved outer surface in MERS-CoV RBD that contacts a linker containing a short α helix between blades 4 and 5 of DPP4. Amino acid interactions in patch 1 (B), and in patch 2 (C).
MERS-CoV critical residues for DPP4 binding and viral entry
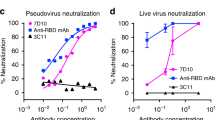
To study the biological relevance of these residues at the interface between MERS-CoV RBD and DPP4, we generated a series of mutant MERS-CoV RBD proteins and characterized their impact on binding activity to DPP4 and the entry efficiency of pseudotyped viruses bearing the mutant S glycoprotein. As shown in Figure 4, mutations at several residues, either singly or in combination, resulted in a significant reduction in binding to DPP4 and lowered the efficiency of viral entry. In patch 1, for instance, a single-residue substitution of Y499A, expected to eliminate hydrogen bonding between Y499 and R336, significantly interrupted the binding between MERS-CoV RBD and DPP4, as well as hindered viral entry (Figure 4). Similarly, combined substitutions involving E536R, D537K and D539K, hypothesized to disrupt the native interaction with K267, resulted in profound reduction in the binding and viral entry levels (Figure 4). Furthermore, in patch 2, a single-residue substitution of either L506A or W553A, expected to alter the hydrophobic core (Figure 3C), significantly reduced both binding and viral entry efficiency (Figure 4). In particular, combined residue substitutions involving L506A, W553A and V555A resulted in even more profound reduction in viral entry efficiency (Figure 4), suggesting that the hydrophobic core formed between MERS-CoV RBD and DPP4 plays a critical role in mediating viral binding and entry into the target cells. Finally, a single-residue substitution of D510A that disrupts its salt-bridge interaction with DPP4 R317, or a change of E513A that disrupts its hydrogen-bonding interaction with DPP4 Q344, also hindered, to variable degrees, both binding and viral entry; while the impact of other mutations such as R511A was not significant (Figure 4).
Effect of residue substitution on MERS-CoV RBD binding to DPP4 (A) and entry efficiency of pseudotyped viruses (B). (A) SDS-PAGE analysis of co-purified complexes of wild-type or mutant forms of His-tagged RBD and untagged DPP4. The actual residue changes in the RBD are indicated above each lane. The DPP4 untagged serves as a negative control to exclude nonspecific binding of untagged DPP4 with Ni-NTA resin. (B) Entry efficiency of pseudotyped viruses bearing the wild-type and mutant forms of viral spike glycoprotein. The percentage of entry efficiency was calculated on the basis of luciferase activity of mutant viruses versus that of the wild-type virus. Soluble RBD (∼150 μg/ml) and DPP4 (∼150 μg/ml) were also tested for their inhibitory activity against wild-type virus. One irrelevant soluble protein with the same concentration was used as a negative control. Error bars represent SD of two replicate experiments.
Discussion
In summary, we report the crystal structure of MERS-CoV RBD in complex with the DPP4 extracellular domain and have identified several key residues in the S glycoprotein critical for viral binding and entry into target cells. In general, MERS-CoV RBD consists of two major subdomains, the core and receptor-binding subdomain. While the former shares a high degree of structural similarity to SARS-CoV RBD, the latter is notably divergent from known structures (Figure 2). This structural distinction helps to explain receptor specificities of MERS-CoV and SARS-CoV9,15. The high degree of structural similarity in the core subdomain, however, also suggests that there is a selection pressure for structural conservation or convergence in this region, despite profound sequence diversity.
In terms of receptor binding, MERS-CoV RBD specifically binds to blades 4 and 5 (Supplementary information, Figure S2) but not other blades in the β-propeller domain of DPP4, which could be explained by charge and shape complementarities at the binding interface (Supplementary information, Figure S4). Specifically, the β-propeller domain of DPP4 has three positively charged residues at the outer surface of blades 4 and 5 (K267, R336 and R317), which interact with the negatively charged residues on the surface of RBD (D510, E536, D537 and D539) (Supplementary information, Figure S4). Surrounded by these charge-charge interactions, the short α helix between blades 4 and 5 of DPP4 docks into the hydrophobic concaved surface of RBD (Supplementary information, Figure S4). These structural features suitable for RBD binding are only observed in the outer surface of blades 4 and 5 of DPP4. Furthermore, it is also clear that the enzymatic site of DPP4 is distant from MERS-CoV RBD-binding site (Figure 1), similar to the structure of ACE2 binding with SARS-CoV RBD14. This provides a structural explanation for previous findings showing that DPP4 enzymatic inhibitors do not block the viral entry of MERS-CoV9. However, the overall structural features of DPP4 are clearly distinct from those of ACE2. Such structural differences, in addition to their potential differences in expression levels and tissue distribution, is expected to play a critical role in determining cell tropism as well as pathogenesis of MERS-CoV and SARS-CoV in vivo. Of note, small degree of sequence variation has been identified in the contact residues among DPP4 from different mammals, which may help to determine the cell susceptibility as well as the host range of MERS-CoV (Supplementary information, Figure S5). Nevertheless, our report of structural features of MERS-CoV RBD in complex with the DPP4 has provided atomic understanding of virus and receptor interaction, and will guide development of therapeutics and vaccines against MERS-CoV infection.
Materials and Methods
Expression and purification of human DPP4
Human DPP4 extracellular domain (residues 39-766) with an N-terminal Hemolin signal peptide for secretion and a C-terminal 6× His tag for purification was inserted into pFastBac-Dual vector (Invitrogen). The construct was transformed into bacterial DH10Bac component cells, and the extracted bacmid was then transfected into Sf9 cells using Cellfectin II Reagent (Invitrogen). The low-titer viruses were harvested and then amplified to generate high-titer virus stock, which was used to infect 2 L Sf9 cells at a density of 2 × 106 cells/ml. The supernatant of cell culture containing the secreted DPP4 was harvested 72 h after infection, concentrated and buffer-exchanged to HBS (10 mM HEPES, pH 7.2, 150 mM NaCl). DPP4 was captured by nickel (Ni)-charged resin (GE Healthcare) and eluted with 500 mM imidazole in Tris buffer (50 mM Tris, pH 8.8, 300 mM NaCl). DPP4 was then purified by gel filtration chromatography using the Superdex 200 column (GE Healthcare) pre-equilibrated with Tris buffer (25 mM Tris, pH 8.8, 30 mM NaCl). Fractions containing DPP4 were collected and applied directly to a pre-equilibrated RESOURCE Q column (GE Healthcare) and then eluted with a 30-1000 mM NaCl gradient in 25 mM Tris buffer (pH 8.8). Fractions containing DPP4 were finally purified with gel filtration chromatography using Superdex 200 column.
Identification and preparation of MERS-CoV RBD
A series of constructs encoding different fragments of MERS-CoV S1 protein were designed based on secondary structure prediction. Each fragment was cloned into the vector pFastBac-Dual (Invitrogen) with an N-terminal gp67 signal peptide and C-terminal His tag. All constructs were transformed into bacterial DH10Bac component cells to obtain recombinant bacmid and then transfected into Sf9 insect cells. Viruses were harvested after 7 days, amplified and tested for protein expression. One of the fragments (MERS-CoV RBD, residues 367-606) showed the highest level of protein expression and was able to be co-purified with untagged DPP4 using Ni-charged resin. This fragment was further produced in large quantity and purified using Ni-charged resin and HiLoad Superdex 75 Column (GE healthcare).
Crystallization and data collection
Purified RBD and DPP4 were mixed for 1 h and then loaded to Superdex 200 column. The complex was collected and concentrated to ∼10 mg/ml in HBS buffer (10 mM HEPES, pH 7.2, 150 mM NaCl) for crystallization. Crystals were successfully grown at room temperature using the sitting drop vapor diffusion method by mixing equal volumes of protein and reservoir solution containing 25% (w/v) polyethylene glycol 1500. Crystals were frozen in liquid nitrogen with cryoprotectant (well solution plus 20% (v/v) glycol) before data collection. Diffraction data were collected at the BL17U beam line of the Shanghai Synchrotron Research Facility (SSRF). Diffraction data were indexed, integrated and scaled with the program HKL200020.
Structural determination and refinement
The structure was determined by the molecular replacement method with PHASER21 in CCP4 suite22. The search models are DPP4 extracellular domain structure (PDB code 2G63) and SARS-CoV RBD structure deleting the receptor-binding loop (PDB code 2AJF). Density map improvement by atoms update and refinement was performed with ARP/wARP23, and automatic model extension of the missing receptor-binding subdomain of MERS-CoV was carried out with BUCCANNER24. At the final stages of model building and refinement, glycans were added based on the electron densities. Structure validation was performed with PROCHECK25. All structural figures were made with PyMol26.
Mutagenesis and Ni-NTA pull-down experiments
All MERS-CoV RBD mutants were generated by site-specific PCR mutagenesis and were confirmed by sequencing. Baculoviruses expressing these mutants with C-terminal His tag were prepared in the same way as wild-type MERS-CoV RBD. Sf9 cells were coinfected by baculovirus expressing untagged DPP4 and a MERS-CoV RBD mutant baculovirus. The supernatant of 2 ml cell culture in six-well plates was harvested 72 h after infection and buffer-exchanged to HBS (10 mM HEPES, pH 7.2, 150 mM NaCl). Protein was captured by adding 40 μl Ni-charged resin (GE Healthcare) and incubated at 4 °C for 3 h. After an extensive wash with Tris buffer (50 mM Tris, pH 8.8, 300 mM NaCl) containing 20 mM imidazole, the bound protein was eluted with 40 μl Tris buffer (50 mM Tris, pH 8.8, 300 mM NaCl) containing 500 mM imidazole. Twenty microlitres of eluted sample was mixed with 5 μl of 5× SDS-PAGE loading buffer and boiled at 100 °C for 5 min, and then loaded on 12% SDS-PAGE gel. The gel was stained with coomassie blue.
Pseudotyped virus and viral entry
Pseudotyped virus was generated by co-transfection of human immunodeficiency virus backbone expressing firefly luciferase (pNL43R-E-luciferase) and MERS-CoV spike glycoprotein expression vector (pcDNA3.1+; Invitrogen) into the 293T cells27. Viral supernatants were harvested 48 h later, normalized by p24 ELISA kit (Beijing Quantobio Biotechnology Co., Ltd., China) before infecting the target cells Huh7. The infected Huh7 cells were lysed at 48 h after infection and viral entry efficiency was quantified by comparing the luciferase activity between pseudotyped viruses bearing the mutant and wild-type MERS-CoV spike glycoprotein. The inhibitory effect of soluble RBD and DPP4 was analyzed by incubating with pseudotyped virus bearing the wild-type MERS-CoV spike glycoprotein before the infection.
PDB deposition
The coordinates and diffraction data have been deposited into the Protein Data Bank with accession code 4L72.
Accession codes
References
Bermingham A, Chand MA, Brown CS, et al. Severe respiratory illness caused by a novel coronavirus, in a patient transferred to the United Kingdom from the Middle East, September 2012. Euro Surveill 2012; 17:20290.
Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA . Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med 2012; 367:1814–1820.
de Groot RJ, Baker SC, Baric RS, et al. Middle East Respiratory Syndrome Coronavirus (MERS-CoV); Announcement of the Coronavirus Study Group. J Virol 2013; 87:7790–7792.
WHO. World Health Organization Global alert and response; Coronavirus infections. Available from: http://www.who.int/csr/disease/coronavirus_infections/en/
Chan JF, Li KS, To KK, Cheng VC, Chen H, Yuen KY . Is the discovery of the novel human betacoronavirus 2c EMC/2012 (HCoV-EMC) the beginning of another SARS-like pandemic? J Infect 2012; 65:477–489.
Memish ZA, Zumla AI, Al-Hakeem RF, Al-Rabeeah AA, Stephens GM . Family cluster of Middle East respiratory syndrome coronavirus infections. N Engl J Med 2013; 368:2487–2494
van Boheemen S, de Graaf M, Lauber C, et al. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. MBio 2012; 3:e00473–e00412.
Woo PCY, Lau SKP, Li KSM, Tsang AKL, Yuen K-Y . Genetic relatedness of the novel human group C betacoronavirus to Tylonycteris bat coronavirus HKU4 and Pipistrellus bat coronavirus HKU5. Emerg Microbes Infect 2012; 1:e35.
Raj VS, Mou H, Smits SL, et al. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 2013; 495:251–254.
Jiang S, Lu L, Du L, Debnath AK . A predicted receptor-binding and critical neutralizing domain in S protein of the novel human coronavirus HCoV-EMC. J Infect 2013; 66:464–466.
Weiss SR, Navas-Martin S . Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol Mol Biol Rev 2005; 69:635–664.
Du L, He Y, Zhou Y, Liu S, Zheng BJ, Jiang S . The spike protein of SARS-CoV: a target for vaccine and therapeutic development. Nat Rev Microbiol 2009; 7:226–236.
Masters PS . The molecular biology of coronaviruses. Adv Virus Res 2006; 66:193–292.
Li F, Li W, Farzan M, Harrison SC . Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science 2005; 309:1864–1868.
Li W, Moore MJ, Vasilieva N, et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003; 426:450–454.
Wu K, Li W, Peng G, Li F . Crystal structure of NL63 respiratory coronavirus receptor-binding domain complexed with its human receptor. Proc Natl Acad Sci USA 2009; 106:19970–19974.
Yeager CL, Ashmun RA, Williams RK, et al. Human aminopeptidase N is a receptor for human coronavirus 229E. Nature 1992; 357:420–422.
Gallagher T, Perlman S . Public health: broad reception for coronavirus. Nature 2013; 495:176–177.
Jiang S, Lu L, Du L, Debnath AK . Putative conformations of the receptor-binding domain in S protein of hCoV-EMC in complex with its receptor dipeptidyl peptidase-4. J Infect 2013; 67:156–158.
Otwinowski Z, Minor, W . Processing of X-ray diffraction data collected in oscillation mode. Method Enzymol 1997; 276:307–326.
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ . Phaser crystallographic software. J Appl Crystallogr 2007; 40:658–674.
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr D Biol Crystallogr 1994; 50:760–763.
Cohen SX, Ben Jelloul M, Long F, et al. ARP/wARP and molecular replacement: the next generation. Acta Crystallogr D Biol Crystallogr 2008; 64:49–60.
Cowtan K . The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr D 2006; 62:1002–1011.
Laskowski RA, MacArthur MW, Moss DS, Thornton JM . PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 1993; 26:283–291.
DeLano WL . Pymol Molecular Graphics System (DeLano Scientific, San Carlos, California, USA, 2002).
Shang H, Han X, Shi X, et al. Genetic and neutralization sensitivity of diverse HIV-1 env clones from chronically infected patients in China. J Biol Chem 2011; 286:14531–14541.
Acknowledgements
We thank Dr JW Wang for assistance with structural determination, and Dr JH He and other staff members at Shanghai Synchrotron Research Facility beam line BL17U for help with data collection. This work was supported by the Ministry of Science and Technology (2010CB912402 and 2011CB910502), the Ministry of Health (2012ZX10001009), the Fok Ying Tung Education Foundation to X Wang and the National Outstanding Youth Award (30825035) to L Zhang.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Gel filtration chromatography profiles of MERS-CoV RBD (a), DPP4 extracellular domain (b), and RBD-DPP4 complex (c). (PDF 185 kb)
Supplementary information, Figure S2
The N-terminal eight-bladed β-propeller domain of DPP4 contacts MERS-CoV RBD through blades 4 and 5. (PDF 232 kb)
Supplementary information, Figure S3
Structure-based sequence alignment of MERS-CoV and SARS-CoV RBDs. (PDF 194 kb)
Supplementary information, Figure S4
Electrostatic potential maps of DPP4 and MERS-CoV RBD. (PDF 167 kb)
Supplementary information, Figure S5
Sequence alignment of DPP4 eight-bladed β-propeller domains from different species. (PDF 1252 kb)
Supplementary information, Table S1
Data collection and refinement statistics (PDF 48 kb)
Supplementary information, Table S2
Contacting residues at the binding interface (d ≤ 3.6 Å) (PDF 26 kb)
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0
About this article
Cite this article
Wang, N., Shi, X., Jiang, L. et al. Structure of MERS-CoV spike receptor-binding domain complexed with human receptor DPP4. Cell Res 23, 986–993 (2013). https://doi.org/10.1038/cr.2013.92
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2013.92
Keywords
This article is cited by
-
(Pro)renin receptor mediates tubular epithelial cell pyroptosis in diabetic kidney disease via DPP4-JNK pathway
Journal of Translational Medicine (2024)
-
Isolation and characterization of a pangolin-borne HKU4-related coronavirus that potentially infects human-DPP4-transgenic mice
Nature Communications (2024)
-
Recombinant protein embedded liposome on gold nanoparticle based on LSPR method to detect Corona virus
Nano Convergence (2023)
-
Plant lectins as versatile tools to fight coronavirus outbreaks
Glycoconjugate Journal (2023)
-
Consolidating food safety measures against COVID-19: a review
Journal of the Egyptian Public Health Association (2022)