Abstract
Multipotent neural stem/progenitor cells hold great promise for cell therapy. The reprogramming of fibroblasts to induced pluripotent stem cells as well as mature neurons suggests a possibility to convert a terminally differentiated somatic cell into a multipotent state without first establishing pluripotency. Here, we demonstrate that Sertoli cells derived from mesoderm can be directly converted into a multipotent state that possesses neural stem/progenitor cell properties. The induced neural stem/progenitor cells (iNSCs) express multiple NSC-specific markers, exhibit a global gene-expression profile similar to normal NSCs, and are capable of self-renewal and differentiating into glia and electrophysiologically functional neurons. iNSC-derived neurons stain positive for tyrosine hydroxylase (TH), γ-aminobutyric acid, and choline acetyltransferase. In addition, iNSCs can survive and generate synapses following transplantation into the dentate gyrus. Generation of iNSCs may have important implications for disease modeling and regenerative medicine.
Similar content being viewed by others
Introduction
The successful establishment of induced pluripotent stem cells (iPSCs)1 raises the question of whether terminally differentiated cells can be directly converted to cells of other lineages by ectopic expression of defined factors. Although several recent studies have demonstrated such a possibility2,3,4,5,6, the reprogrammed cells obtained in those studies were mostly originated from dermal fibroblasts, which may not be the most ideal cells for cell-lineage-conversion studies due to the presence of multipotent precursor cells7. In addition, whether the trans-differentiated cells exert functions in vivo has not been fully explored2,6.
Sertoli cells are mesoderm-originated terminally differentiated cells that provide a nurturing microenvironment for spermatogenesis8. Sertoli cells can be cultured at high purity to avoid the heterogeneity problems inherent in mouse dermal fibroblast cultures, thus may serve as a good model to test the principles of trans-germ-layer differentiation and somatic-to-multipotent conversion.
We initially selected nine factors trying to convert mesoderm-derived terminally differentiated sertoli cells to neural lineages that have an ectodermal origin. We focused on the basic helix-loop-helix factors, a super family that plays critical roles in the development, differentiation, functional stability, and plasticity of neural cells. Ascl1, Neurog2 (Ngn2), Hes1, and Id1 belong to different subfamilies and are part of a reciprocal regulation network that is involved in the self-renewal, timing of differentiation, and cell fate specification of neural cells9,10,11. We also chose Pax6 and Brn2 for their close interaction with basic helix-loop-helix factors11,12. Sox2 is one of the four “Yamanaka factors” that is also highly expressed in the neuroepithelium of the developing central nervous system and plays critical roles in neural stem cells13. We also included c-Myc and Klf4 to facilitate the chromosome remolding process14,15. The nine reprogramming factors, Ascl1, Ngn2, Hes1, Id1, Pax6, Brn2, Sox2, c-Myc, and Klf4, are all expressed in normal neural stem/progenitor cells. These factors either perform significant roles in neural lineage development and specification, or are involved in the epigenetic regulation of the reprogramming process.
Here, we demonstrate that, without first establishing pluripotency, nine transcription factors can directly convert sertoli cells into multipotent neural stem/progenitor cells. The obtained iNSCs show a gene expression profile similar to normal neural stem cells (NSCs), and are functional both in vitro and in vivo.
Results
Derivation of induced neural stem/progenitor cells
Sertoli cells were obtained from 5-day-old pups by a modified procedure16, and tested for purity by staining with markers specific for sertoli cells or NSCs. Nearly all of the cells stained positive for Wt1 and Gata4, two specific markers of sertoli cells (Supplementary information, Figure S1A and S1B), while no expression of Olig2, Pax6, Sox2, or Doublecortin (Dcx) – all well documented neural lineage markers – was detected on sertoli cells (Supplementary information, Figure S1A and S1C). Sertoli cell culture was also negative for Ddx4 and 3β-HSD, specific markers of germ cells and leydig cells, respectively, as well as Oct4 and Nanog, two markers expressed by spermatogonial stem cells (Supplementary information, Figure S1A and S1C)17. To further confirm that sertoli cells are not able to transform/differentiate into neural cells in the culture medium that promotes the propagation or differentiation of NSCs, sertoli cells were cultured in N2B27 medium with or without basic fibroblast growth factor (bFGF) and epidermal growth factor (EGF) for 2 to 4 weeks. Under either condition, no cells of the neural lineage were detected, excluding the possibility of neural cell contamination in the culture.
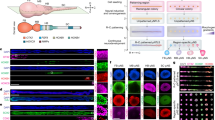
The sertoli cells isolated from neonatal pups normally go through a few cell cycles before completely losing proliferative capacity in about 2 weeks. We infected the cells that were in the last several divisions with retrovirus vectors expressing each of the nine transcription factors (Ascl1, Ngn2, Hes1, Id1, Pax6, Brn2, Sox2, c-Myc, and Klf4). After two rounds of infection/recovery, the cells were replated onto poly-D-lysine/laminin-coated plates in N2B27 medium with the addition of bFGF and EGF (Figure 1A). In about 3 days after replating, around twenty colonies would normally emerge per 1 × 105 cells plated, but only one to two of them turned out to be well-reprogrammed iNSC colonies. The low frequency was most likely due to the poor proliferative ability of sertoli cells and subsequently the low infection efficiency with retrovirus (Supplementary information, Figure S2). The well-reprogrammed colonies could be identified by the enhanced proliferative ability and bipolar shape, a typical NSC morphology that is distinct from sertoli cells and iPSCs (Figure 1B and 1C, and Supplementary information, Figure S3). The iNSC colonies could be expanded either as monolayer or floating spheres (Figure 1C and 1D). After about 1 month of propagation, the induced cells would reach sufficient number for characterization and other analyses. In order to refine the pool of factors required for induction, we tested 8-factor combinations with the removal of each individual factor. Surprisingly, most of the 8-factor combinations did not give rise to iNSC colonies, and the only dispensable factor was Sox2. PCR and southern blot results confirmed the genomic integration of the reprogramming factors (Supplementary information, Figure S4). We repeated the 8-factor tests three times and only the combination without Sox2 (Pax6, Ngn2, Hes1, Id1, Ascl1, Brn2, c-Myc, and Klf4) could generate iNSC colonies, implying a complicated reprogramming process between these two lineages.
Characterization of iNSCs. (A) The paradigm used for iNSCs induction from sertoli cells by defined factors. (B-D) Images of sertoli cells before infection (B), iNSCs in monolayer (C) and suspension cultures (D). (E-H) iNSCs were immunoreactive for Nestin (iNSC 1-9, red), Pax6 (iNSC 1-9 with exogenous Pax6 silenced, green), Olig2 (iNSC 1-9, red), and Sox2 (iNSC 12-8-1, an 8-factor-induced iNSC line omitting Sox2, red). (I) Few iNSCs (∼ 0.7%) expressed Doublecortin (green) in monolayer culture. (J, K) iNSCs cultured as spheres expressed neural markers Nestin (green, neural stem/progenitor cells) and Doublecortin (green, neural precursors). (L) iNSCs were immunoreactive for Mki67 (red). Higher magnification (3 × of the marked region in the original figure) was shown in the inset. (M) Karyotype analysis of iNSC1-9 (passage 15) showed normal ploidy and male origin. Blue, DAPI. Scale bars: 100 μm (B-D), 50 μm (E-L).
Characterization of iNSCs
We used iNSC 1-9, an established line that had been induced with nine factors, to characterize the NSC-related properties. iNSC 1-9 readily expressed specific neural stem cell markers that include Nestin (Figure 1E), Pax6 (Figure 1F), Olig2 (Figure 1G), and Sox2. Pax6 expression should be endogenous in that the exogenous Pax6 was found to have been silenced in iNSC 1-9 (Supplementary information, Figure S5). Endogenous Sox2 expression was also detected in the 8-factor-induced iNSCs (iNSC 12-8-1) where Sox2 has been removed from the pool of inducing factors (Figure 1H). Meanwhile, iNSCs expressed Olig2, another neural stem cell marker that has been documented previously18,19. iNSCs presented a morphology and doubling time very similar to those of normal NSCs. When cultured on non-coated plates, iNSCs formed floating aggregates (Figure 1D) that grew in size and formed secondary spheres upon dissociation. The majority of cells were Nestin- and Mki67-positive, and the Dcx staining revealed about 0.7% positive cells in the monolayer culture and about 3% positive cells in the sphere culture (Figure 1I-1L). After 15 passages, the iNSCs still retained a normal karyotype consistent with the male origin of sertoli cells (Figure 1M). All of these characteristics were equivalent to neural stem cells isolated directly from fetal or newborn mouse brains.
Global gene expression of iNSCs
Next, we compared the global gene expression pattern of early- and late-passage iNSCs with normal NSCs and sertoli cells by microarray analysis. Hierarchical clustering revealed that the global gene expression profile of iNSCs showed a higher degree of similarity to their normal NSC counterparts than to the sertoli cells (Figure 2A). Among the 1 757 differentially expressed genes between NSCs and sertoli cells, 1 062 genes were downregulated or upregulated in the same way as iNSCs versus sertoli cells (Figure 2B and Supplementary information, Table S1). Many genes important to the neural lineage, such as Egfr, Mki67, Sox11, Cdk2, Cdk4, Notch3, Robo1, Ezh2, Crmp1, Mll1, and Cdk5rap2, were significantly upregulated in NSCs and iNSCs compared with those in sertoli cells, whereas key sertoli cell genes, including Wt1, Dhh, Inhbb, Inhba, and Gata4, were significantly downregulated in iNSCs (Supplementary information, Figure S6). Real-time PCR assay showed that among the exogenous reprogramming genes, only Pax6 was silenced in the 9-factor iNSCs, and Id1 was silenced in the 8-factor (without Sox2) iNSCs (Supplementary information, Figure S5). Incomplete silencing of exogenous genes was also observed in a recent reprogramming study where fibroblasts were directly converted to cardiomyocytes3. It is still unknown whether sustained expression of the exogenous reprogramming factors is required to maintain the properties of iNSCs.
Whole-genome gene expression profile of iNSCs. (A) Hierarchical clustering analysis of global gene expression patterns of NSCs, iNSCs and sertoli cells. The “e-iNSC” and “l-iNSC” represented the early passage (13) and late passage (22) of iNSCs, respectively. NSC1 and NSC2 were neural stem cells isolated from newborn pup brains. NSC3 was isolated from E12.5 fetal brain. iNSC 1-9 represented a 9-factor iNSC line and iNSC 12-8-1 represented an 8-factor (without Sox2) iNSC line. Clusters were determined using the hclust package of R. (B) Genes differentially expressed between NSCs and sertoli cells. Group I and II were genes downregulated or upregulated in both iNSCs and NSCs vs. sertoli cells, respectively. Heat map was plotted using heatmap.2 function of R, and gene expressions were scaled to Z-scores.
Neuronal differentiation of iNSCs in vitro and in vivo
To verify that the iNSCs were capable of differentiating into neurons, about 1.0 × 104 cells of iNSC 1-9 at passage 10 to 18 were plated onto poly-D-lysine/laminin-coated coverslips in N2B27 medium with removal of growth factors (bFGF and EGF), and addition of BDNF and NT-3. At 2 weeks, ∼30% of the differentiated iNSCs stained positive for MAP2 and ∼6% positive for NeuN, two well-characterized neuronal markers. For a longer differentiation paradigm, we changed the ratio of N2 to B27 from 1:1 to 1:3 to support further differentiation and maturation of neurons18. After 3 weeks of differentiation, 2.8% of the neurons stained positive for TH, 33.9% positive for γ-aminobutyric acid (GABA) and 2.6% positive for choline acetyltransferase (ChAT) (Figure 3A-3C). The proportion of TH-positive neurons was increased to ∼10.2% after treatment with sonic hedgehog and fibroblast growth factor-8 (Figure 3D-3F). Meanwhile, TH-positive neurons co-expressed Pitx3, a mesencephalic dopaminergic neuronal marker, suggesting their midbrain identity (Figure 3G-3I). Moreover, synapsin staining suggested that functional synapses had formed by 3 weeks of differentiation (Figure 3J-3L). In addition, iNSCs could also differentiate into astrocytes and oligodendrocytes in vitro (Supplementary information, Figure S7A-S7C).
In vitro differentiation of iNSCs. (A-C) Neurons differentiated from iNSCs expressed MAP2 (red, A, B), NeuN (red, C), and specific neuronal markers TH (green, A), GABA (green, B), and ChAT (green, C) after a 3-week differentiation. (D-F) The proportion of TH-positive neurons was significantly increased after treatment with sonic hedgehog and fibroblast growth factor-8. TH (red), MAP2 (green). (G-I) TH-positive neurons (red) co-labeled with mesencephalic dopaminergic neuronal marker Pitx3 (green). (J, K) Triple staining for MAP2 (white), TH (red), and Synapsin (green) of iNSC-derived neurons after a 3-week differentiation. (L) Higher magnification of the region marked in J. Blue, DAPI. Scale bars: 50 μm (A-I), 20 μm (J, K), and 10 μm (L).
To explore whether iNSCs can function in vivo, we introduced GFP-labeled iNSCs into the dentate gyrus (DG) of hippocampus, a native neurogenic area in adult mouse brain. C57BL/6 mice 5-6 weeks of age received grafts bilaterally and were sacrificed 2-4 weeks post transplantation for graft analysis (Figure 4A). Consistent with a previous report20, a bolus of donor cells was observed at the injection site within the DG. Many cells had migrated from the injection site to populate the neighboring subgranular zone and granular cell layer (Figure 4B and 4C). At 3 weeks following transplantation, NeuN-positive grafted cells were detected (Figure 4C and Supplementary information, Figure S7D and S7E), suggesting that some iNSC-derived neurons had matured in vivo at this time point. To explore whether the grafted cells had established functional connections with host neurons, we stained the sections for Synapsin and the result suggested that some grafted cells had received extensive presynaptic innervations from other neurons 4 weeks post transplantation (Figure 4D-4F and Supplementary information, Movie S1). We examined 38 hippocampi (19 mice) 4-6 weeks following transplantation of syngeneic iNSCs and did not observe tumorigenesis, indicating that iNSCs may provide a potential source for cell replacement therapy.
In vivo transplantation of iNSCs. (A) Schematic representation of iNSCs transplantation. (B) Overview of GFP+ cells in DG two weeks after grafting of iNSCs. GFP (green), Doublecortin (white). (C) A GFP+ cell (green) co-localized with NeuN (red) was shown three weeks after transplantation. (D-F) A GFP+ cell (green) co-labeled with Synapsin (red) four weeks post transplantation (a merged image of 3 consecutive scans with an interval of 0.5 μm). Blue, DAPI. Scale bars: 50 μm (B), 20 μm (C), 10 μm (D-F).
To examine whether iNSC-derived neurons possess functional membrane properties, we performed whole-cell recording 2 or 3 weeks following in vitro differentiation. The resting membrane potentials of iNSC-derived neurons ranged from −55 mV to −75 mV. Following 2 weeks of differentiation, some cells exhibited spontaneous action potentials, which could be blocked by tetradotoxin (TTX), a specific inhibitor of sodium ion channels (Figure 5A). When recording in the voltage-clamp mode, we observed inward fast inactivating currents and outward currents, which may correspond to the opening of voltage-dependent Na+ and K+ channels, respectively. The inward current could be blocked and recovered by addition and removal of TTX (Figure 5B). Mature action potentials could be elicited by depolarizing the membrane in the current-clamp mode in a quarter of tested cells at 2 weeks (25%, n = 12, Figure 5C). At 3 weeks, more neurons (57%, n = 14) fired mature action potentials (Figure 5D) evoked by current injection, which were sensitive to TTX treatment. Furthermore, iNSC-derived neurons showed spontaneous EPSCs under the whole-cell voltage-clamp mode 3 weeks after differentiation, which could be blocked by bath application of CNQX and CPP, two inhibitors of glutamate receptors (Figure 5E)21, further demonstrating that functional synapses had formed by this time point. The above results indicate that iNSC-derived neurons exhibit functional membrane properties of mature neurons in vitro.
Electrophysiology of iNSC-derived neurons. (A) A recorded neuron co-labeled with Neurobiotin (green) and MAP2 (red). iNSC-derived neurons exhibited spontaneous action potentials two weeks after in vitro differentiation and were sensitive to TTX treatment. Scale bar: 20 μm. (B) Whole-cell currents were recorded in the voltage-clamp mode with a basal holding potential of −70 mV. Voltage steps ranging from −90 mV to +30 mV were delivered at 10 mV increments (lower panel). (C) At 2 weeks after differentiation, a few iNSC-derived neurons (3/12) showed mature TTX-sensitive action potentials evoked by step-current-injection in the current-clamp mode. (D) At 3 weeks after differentiation, a majority of neurons (8/14) showed mature evoked action potentials that were sensitive to TTX treatment. (E) Spontaneous EPSCs were recorded from an iNSC-derived neuron 3 weeks after differentiation. The membrane potential was held at −70 mV. The spontaneous postsynaptic currents could be blocked by bath application of CNQX and CPP (right panel).
Discussion
The adult central nervous system is limited in its regenerative capacity. Transplantation of lineage-committed neural stem/progenitor cells is a promising approach for treatment of neurological disorders. Autologous neural tissue/cells represent an ideal source for this purpose in that they pose fewer ethical issues and may avoid complications caused by mismatch-induced signaling22. Induced pluripotent stem cells1 and induced neurons (iNs)2 are two types of cells that are reprogrammed from somatic cells by introduction of a defined set of transcription factors, and thus retain very similar or identical genetic background as the donor cells. Although iPSCs represent an unlimited source for deriving neural progenitor cells, clinical applications still have a long way to go due to the limitation of gaining sufficiently tight control over the proliferative activity after transplantation23. In contrast, induced neurons are terminally differentiated cells that have completely exited the cell cycle and therefore would have difficulty surviving transplantation. In addition, iNs converted from mouse dermal fibroblasts have limited capacity to generate specific neuronal subtypes according to the reported protocol2. Therefore, a reliable strategy to induce a multipotent state may provide an alternative for a safer and more efficient clinical application.
Here we choose sertoli cells as the starting cells, which could also be isolated from adult human and non-human primate and cultured at high purity in vitro24,25,26,27,28, and show that mesoderm-derived terminally differentiated sertoli cells can be directly converted to a multipotent state of ectodermal neural lineage by a combination of nine transcription factors. The global gene expression of iNSCs has been broadly reprogrammed from the original sertoli cell patterns to a profile similar but not identical to that of normal neural stem cells. The iNSCs can self-renew and differentiate into astrocytes, oligodendrocytes, and different subtypes of neurons that are dopaminergic, GABAergic, or cholinergic. The iNSC-derived neurons possess normal electrophysiological properties in vitro, and may have formed functional synapses following transplantation into the DG of adult mouse brain.
The first iNSC colony normally emerged 3 days after infection, a much shorter period of time compared with the lengthy induction process required for iPSCs. The shape of the colony, as well as the nucleus to cytoplasm ratio, and the bipolar morphology of iNSCs were all distinct from iPSCs, suggesting that iNSCs had been established without going through a pluripotent state. Staining of the emerging colonies 6 days post infection with Oct4, Nanog, and Nestin (Supplementary information, Figure S3) further excluded the possibility of contamination from induced pluripotent cells and confirmed a direct conversion nature of iNSCs.
Neural stem cells possess a multipotent potential that requires a complex molecular network to restrict the cells from diversion into a non-neural lineage and yet maintain the self-renewal ability and the plasticity to differentiate into neurons, astrocytes, and oligodendrocytes. Along the developmental process, a set of critical transcription factors are activated, and coordinate a series of feed-back and feed-forward loops to stabilize the cells in a dynamic balance. A relatively stable homeostasis normally involves antagonistic pathways that reciprocally regulate each other so that cells can tolerate small intracellular and/or environmental perturbations without shifting away from their normal properties. The inducing factors used in this study are all expressed in normal neural stem cells and are involved in antagonistic regulatory pathways. For example, Hes1 and Ascl1 negatively regulate each other and the right ratio may help determine self-renewal or initiation of neurogenesis29. The factors are important for cell cycle regulation, inhibition of differentiation, or neurogenesis. We chose these factors to try to establish each important property of a neural stem cell. We reasoned that only if a sertoli cell has incorporated the right relative amount of each factor (another possible reason for the low efficiency of conversion), reciprocal regulatory networks may be established and progress to a balance that mimics a neural stem cell state. However, the real-time PCR results showed that most of the exogenous factors had not been silenced. Future studies applying inducible expression strategies are needed to illustrate whether silencing exogenous factors affects the neural stem cell property, although incomplete silencing of the exogenous factors seemed not to have interfered with the differentiation and other key properties of neural stem cells.
Although induced neural progenitor cells were also reported in a recent study6 by applying a very different strategy, where four “Yamanaka factors” were used to induce fibroblasts into a highly unstable intermediate state followed by directing those cells to neural lineages, the limited proliferative ability of those obtained cells needs to be much improved before the cells can be considered as a potential source for cell replacement therapy. In contrast, the iNSCs generated in the current study can readily establish cell lines and retain the self-renewal ability even after 25 passages. The iNSCs at passage 10-18 were used for electrophysiological examinations and the differentiated neurons were shown to possess proper membrane properties. In addition to forming synapses in culture dishes, iNSCs can survive transplantation and give rise to neurons that may have formed synapses in vivo. The great expandability and the multipotency to generate neurons and glia suggest a close similarity to normal NSCs and may have important implications for regenerative medicine.
The present study convincingly demonstrates that differentiated somatic cells can be directly converted into multipotent stem/progenitor cells across germ layers. However, it is possible that induction of iNSCs is context-dependent. It might require a different cocktail of factors to convert cells of a different type or species. Replacement of induction genes with small molecules, application of non-integrating strategies, and selection of originating cell sources that are more readily accessible may further refine the reprogramming pool to the minimal and most efficient combination, and provide a safer and more efficient source material for clinical applications.
Materials and Methods
Isolation of sertoli cells and neural stem cells
The isolation of sertoli cells with high purity was modified from a previous publication16. Briefly, decapsulated testes from 5-day-old C57BL/6 or B6D2F1 mice were digested sequentially with 0.1% collagenase IV (Invitrogen) at 37 °C for 20 min followed by 0.1% collagenase IV plus 0.1% hyaluronidase (Sigma) at 37°C for 10 min. Cells were washed and treated with 0.1% collagenase IV, 0.1% hyaluronidase, 0.25% trypsin (Amresco) and 0.04% DNase (Sigma) at 37 °C for 20 min. After termination of digestion with fetal bovine serum, the cells were filtered through a 70-μm mesh, washed twice with DMEM/F12, and cultured in sertoli cell medium containing DMEM/F12 supplemented with 10% fetal bovine serum, 100 U/ml penicillin and 100 μg/ml streptomycin sulfate (all from Invitrogen). Cells were grown at 37 °C, 5% CO2 for 2-3 days and passaged upon 80% confluence. Sertoli cells of passage number two were used for purity analysis and retrovirus infection.
Normal neural stem cells were isolated from brains of newborn pups (NSC1 line on B6D2F1 and NSC2 on C57BL/6 background) or E12.5 fetal brain (NSC3 line on C57BL/6 background) as previously described18, and cultured in N2B27 medium with EGF and bFGF (both at 20 ng/ml, R&D Systems)30.
All the animal experiments were performed according to the Guidelines for the Care and Use of Laboratory Animals established by the Beijing Association for Laboratory Animal Science.
Molecular cloning and retrovirus infection
cDNAs of candidate factors and EGFP were cloned into pQCXIN retrovirus vectors (Clontech). pMX-c-Myc and pMX-Klf4 were purchased from Addgene1. The retroviruses were packaged in gag/pol-293T cells and concentrated with Amicon ultra-15 centrifugal filter unit (6 000× g, 1 h), and titrated by infecting 293T cells with series of dilutions. The infection efficiency of sertoli cells was up to 30% monitored by using pQCXIN-EGFP retrovirus, probably due to the weak proliferative ability of sertoli cells. About 200 000 sertoli cells on C57BL/6 or B6D2F1 background were infected with ∼10 μl of each retrovirus (108 particles per ml) with 4 μg/ml polybrene for about 24 h and recovered for another 24 h. The infection was repeated to improve efficiency, followed by replating the cells into poly-D-lysine (PDL, Sigma) and Laminin (Invitrogen)-coated 24-well plates with N2B27 medium plus EGF and bFGF (both at 20 ng/ml). The medium was changed every 2 days during the induction period.
Neuronal differentiation
iNSCs were propagated in N2B27 medium with EGF and bFGF. For pan-neuronal differentiation, 5 000–10 000 iNSCs were plated onto PDL/Laminin-coated 24-well plates in N2B27 medium with withdrawal of EGF and bFGF, and addition of BDNF and NT-3 (both at 10 ng/ml, R&D Systems). For a longer differentiation paradigm, the concentration of BDNF and NT-3 was increased to 20 ng/ml. iNSCs differentiated for 2 weeks were examined for expression of neuronal markers (MAP2 and NeuN) and oligodendroglial marker (O4). At least 300 DAPI-positive cells from five randomly selected areas were counted to calculate the fraction of each lineage. Staining of TH, GABA, and ChAT was performed after a 3-week differentiation, and the percentage of each neuronal subtype was obtained by counting at least 300 MAP2-positive cells that were co-labeled with TH, GABA, or ChAT staining. To generate TH-positive neurons, iNSCs were exposed to bFGF (10 ng/ml), sonic hedgehog (400 ng/ml), and fibroblast growth factor-8 (100 ng/ml) according to a previously reported protocol30. For astrocyte differentiation, iNSCs were exposed to 1% fetal bovine serum and BMP4 (10 ng/ml) in N2B27 medium without EGF or bFGF for 5 days followed by GFAP staining18.
Immunofluorescence
For immunofluorescent staining, cells grown on PDL/Laminin-coated coverslips were washed and fixed with 4% paraformaldehyde for 10 min at room temperature. After washes with PBS, cells were blocked in PBS containing 3% donkey serum (Jackson Immunoresearch) plus 0.3% Triton X-100 for 1 hour at room temperature. The primary and secondary antibodies were diluted in PBS containing 1% donkey serum plus 0.3% Triton X-100 and incubated overnight at 4 °C and 1 h at room temperature, respectively. The following primary antibodies were used: mouse anti-Nestin (Millipore, 1:200), rabbit anti-Pax6 (Abcam, 1:200), rabbit anti-Sox2 (Millipore, 1:1 000), rabbit anti-Olig2 (Millipore, 1:500), goat anti-Doublecortin (Santa Cruz, 1:100), rabbit anti-Mki67 (Millipore, 1:300), rabbit anti-Tyrosine Hydroxylase (Millipore, 1:500), sheep anti-Tyrosine Hydroxylase (Millipore, 1:500), mouse anti-MAP2 (Millipore, 1:300), rabbit anti-GABA (Sigma, 1:2 000), goat anti-ChAT (Sigma, 1:500), mouse anti-NeuN (Millipore, 1:100), rabbit anti-Synapsin (Sigma, 1:100), rabbit anti-Pitx3 (Abcam, 1:100), mouse anti-GFP (Millipore, 1:500), rabbit anti-GFP (Invitrogen 1:500), rabbit anti-Wt1 (Santa Cruz, 1:100), goat anti-Gata4 (Santa Cruz, 1:100), chicken anti-GFAP (Millipore, 1:400), mouse anti-O4 (Millipore, 1:60), rabbit anti-Ddx4 (Abcam, 1:200), goat anti-3β-HSD (Santa Cruz, 1:200), goat anti-Oct4 (Santa Cruz, 1:100), and rabbit anti-Nanog (Millipore, 1:300). FITC-, Cy3-, or Cy5-conjugated secondary antibodies were obtained from Jackson Immunoresearch and Abcam. Streptavidin-FITC was obtained from Vector Labs. The images were collected by using a Zeiss LSM 710 microscope.
Electrophysiology
iNSCs were grown on coated coverslips and differentiated for 14 or 21 days before recording. The coverslip was placed in a recording chamber (0.5 ml in volume) on the fixed stage of an upright microscope (DMLFSA, Leica) equipped with epifluorescence and a 40× water-immersion objective lens configured for DIC. The cells were continuously superfused with oxygenated bicarbonate-buffered ACSF (in mM: 119 NaCl, 26.2 NaHCO3, 11 Glucose, 2.5 KCl, 1.0 K2HPO4, 2.5 CaCl2, and 1.3 MgCl2). Spontaneous and step-current-injection evoked action potentials were recorded in the current-clamp mode. Whole-cell currents were recorded in the voltage-clamp mode with a basal holding potential of −70 mV; voltage steps ranging from −90 to +30 mV were delivered at 10 mV increments. TTX was applied at a concentration of 100nM. Spontaneous EPSCs were recorded in the voltage-clamp mode 3 weeks after differentiation. The cells were held at −70 mV. Both CNQX and CPP (all from Sigma) were used at a concentration of 10 μM. Stimulus delivery and data acquisition were carried out via the Axopatch 200B amplifier. The data was digitized at 10 kHz with a 2 kHz low-pass filter, and analyzed using Clampfit 9 (Axon Instruments). The pipette solution for whole-cell recordings contained (in mM) 120 potassium gluconate, 5 NaCl, 10 KCl, 1 MgCl2, 1 EGTA, 10 Hepes, 2 ATP, and 0.5 GTP, adjusted to pH 7.2 with 1 M KOH. Neurobiotin (0.4%, Molecular Probes) was included to label the recorded cells.
Cell transplantation and tissue processing
iNSCs were labeled with GFP by lentivirus infection and about 105 cells were transplanted into each DG (A/P: −0.22 cm, M/L: ± 0.14 cm, D/V: −0.26 cm) of C57BL/6 mice (♂6-week-old, anesthetized with 80 mg/kg ketamine and 10 mg/kg xylazine). The mice were perfused with 0.9% saline followed by 4% paraformaldehyde 2, 3, or 4 weeks post transplantation. The brains were dissected out and fixed in 4% paraformaldehyde overnight followed by dehydration in 0.1 M PBS containing 30% sucrose for 2 days at 4 °C. Consecutive coronal sections (40 μm) were sliced using a Leica SM 2000R Sliding Microtome and stored in tissue collecting solution (25% glycerin (Sigma), 30% ethylene glycol (Sigma) in 0.1 M phosphate buffer (v/v)) at −20 °C until use.
Karyotype analysis
iNSCs of passage 15 were used for karyotype analysis as described previously31. Images were captured using the Genus System (Applied Imaging Corp.) with a CCD camera mounted on a Zeiss Axioplan2 microscope. The chromosomes were numbered according to a published G-banded karyotype ideogram (http://www.pathology.washington.edu/research/cytopages/ideograms/mouse).
Global gene expression analysis
Mouse genome-wide gene expression analysis was performed using Illumina MouseWG-6 v2.0 Expression BeadChips. Total RNA was extracted from NSC, iNSC and sertoli cells with three independent biological repeats. The raw data were processed using R/Bioconductor package. Linear models and empirical Bayes methods were used to pick the differentially expressed genes that show a fold change > 2 and an adjusted P-value < 0.05. Euclidean distance matrix and complete linkage clustering method were applied for hierarchical clustering analysis.
References
Takahashi K, Yamanaka S . Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006; 126:663–676.
Vierbuchen T, Ostermeier A, Pang ZP, et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 2010; 463:1035–1041.
Ieda M, Fu JD, Delgado-Olguin P, et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 2010; 142:375–386.
Szabo E, Rampalli S, Risueno RM, et al. Direct conversion of human fibroblasts to multilineage blood progenitors. Nature 2010; 468:521–526.
Efe JA, Hilcove S, Kim J, et al. Conversion of mouse fibroblasts into cardiomyocytes using a direct reprogramming strategy. Nat Cell Biol 2011; 13:215–222.
Kim J, Efe JA, Zhu S, et al. Direct reprogramming of mouse fibroblasts to neural progenitors. Proc Natl Acad Sci USA 2011; 108:7838–7843.
Toma JG, Akhavan M, Fernandes KJ, et al. Isolation of multipotent adult stem cells from the dermis of mammalian skin. Nat Cell Biol 2001; 3:778–784.
Mruk DD, Cheng CY . Sertoli-Sertoli and Sertoli-germ cell interactions and their significance in germ cell movement in the seminiferous epithelium during spermatogenesis. Endocr Rev 2004; 25:747–806.
Lyden D, Young AZ, Zagzag D, et al. Id1 and Id3 are required for neurogenesis, angiogenesis and vascularization of tumour xenografts. Nature 1999; 401:670–677.
Ross SE, Greenberg ME, Stiles CD . Basic helix-loop-helix factors in cortical development. Neuron 2003; 39:13–25.
Sansom SN, Griffiths DS, Faedo A, et al. The level of the transcription factor Pax6 is essential for controlling the balance between neural stem cell self-renewal and neurogenesis. PLoS Genet 2009; 5:e1000511.
Castro DS, Skowronska-Krawczyk D, Armant O, et al. Proneural bHLH and Brn proteins coregulate a neurogenic program through cooperative binding to a conserved DNA motif. Dev Cell 2006; 11:831–844.
Episkopou V . SOX2 functions in adult neural stem cells. Trends Neurosci 2005; 28:219–221.
Amati B, Frank SR, Donjerkovic D, Taubert S . Function of the c-Myc oncoprotein in chromatin remodeling and transcription. Biochim Biophys Acta 2001; 1471:M135–145.
Sridharan R, Tchieu J, Mason MJ, et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 2009; 136:364–377.
Richardson LL, Kleinman HK, Dym M . Basement membrane gene expression by Sertoli and peritubular myoid cells in vitro in the rat. Biol Reprod 1995; 52:320–330.
Guan K, Nayernia K, Maier LS, et al. Pluripotency of spermatogonial stem cells from adult mouse testis. Nature 2006; 440:1199–1203.
Conti L, Pollard SM, Gorba T, et al. Niche-independent symmetrical self-renewal of a mammalian tissue stem cell. PLoS Biol 2005; 3:e283.
Pollard SM, Conti L, Sun Y, Goffredo D, Smith A . Adherent neural stem (NS) cells from fetal and adult forebrain. Cereb Cortex 2006; 16(Suppl 1):i112–i120.
Monje ML, Toda H, Palmer TD . Inflammatory blockade restores adult hippocampal neurogenesis. Science 2003; 302:1760–1765.
Greenberg KP, Pham A, Werblin FS . Differential targeting of optical neuromodulators to ganglion cell soma and dendrites allows dynamic control of center-surround antagonism. Neuron 2011; 69:713–720.
Chen Z, Palmer TD . Cellular repair of CNS disorders: an immunological perspective. Hum Mol Genet 2008; 17:R84–R92.
Wernig M, Zhao JP, Pruszak J, et al. Neurons derived from reprogrammed fibroblasts functionally integrate into the fetal brain and improve symptoms of rats with Parkinson's disease. Proc Natl Acad Sci USA 2008; 105:5856–5861.
Lipshultz LI, Murthy L, Tindall DJ . Characterization of human Sertoli cells in vitro. J Clin Endocrinol Metab 1982; 55:228–237.
Lejeune H, Skalli M, Sanchez P, Avallet O, Saez JM . Enhancement of testosterone secretion by normal adult human Leydig cells by co-culture with enriched preparations of normal adult human Sertoli cells. Int J Androl 1993; 16:27–34.
Cudicini C, Lejeune H, Gomez E, et al. Human Leydig cells and Sertoli cells are producers of interleukins-1 and -6. J Clin Endocrinol Metab 1997; 82:1426–1433.
Lee BC, Pineda JL, Spiliotis BE, Brown TJ, Bercu BB . Male sexual development in the nonhuman primate. III. Sertoli cell culture and age-related differences. Biol Reprod 1983; 28:1207–1215.
Majumdar SS, Winters SJ, Plant TM . Procedures for the isolation and culture of Sertoli cells from the testes of infant, juvenile, and adult rhesus monkeys (Macaca mulatta). Biol Reprod 1998; 58:633–640.
Kageyama R, Ishibashi M, Takebayashi K, Tomita K . bHLH transcription factors and mammalian neuronal differentiation. Int J Biochem Cell Biol 1997; 29:1389–1399.
Ying QL, Stavridis M, Griffiths D, Li M, Smith A . Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nat Biotechnol 2003; 21:183–186.
Seabright M . A rapid banding technique for human chromosomes. Lancet 1971; 2:971–972.
Acknowledgements
This study was supported in part by grants from the National Science Foundation of China 90919060 (to Q Z) and 31070946 (to Z C), the “Strategic Priority Research Program” of the Chinese Academy of Sciences XDA01020100 (to Q Z), and the China National Basic Research Program 2007CB947702 (to L W), 2011CBA01101 (to Q Z), and 2011CB965103 (to Z C). We thank Yu Ren, Xiaona Wang, Yixun Liu, Wenhui Nai and Y Alex Zhang for their helpful discussion. There is no conflict of interest declared by any of the authors.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Characterization of isolated sertoli cells. (PDF 373 kb)
Supplementary information, Figure S2
Infection efficiency of sertoli cells is monitored by using pQCXIN-EGFP retrovirus. (PDF 98 kb)
Supplementary information, Figure S3
Characterization of emerging iNSC colonies. (PDF 98 kb)
Supplementary information, Figure S4
Detection of viral integration by PCR and Southern blot analysis. (PDF 64 kb)
Supplementary information, Figure S5
Expression of reprogramming factors detected by quantitative RT-PCR. (PDF 199 kb)
Supplementary information, Figure S6
Expression profiles of selected genes important to neural lineage (A) or sertoli cells (B) from microarray analysis. (PDF 50 kb)
Supplementary information, Figure S7
Differentiation of iNSCs in vitro and in vivo. (PDF 82 kb)
Supplementary information, Table S1
Differentially expressed genes between NSCs and sertoli cells. (XLSX 370 kb)
Supplementary information, Movie S1
A GFP+ cell co-labeled with synapsin. Ten consecutive scans were shown with an interval of 0.5 μm. (AVI 4283 kb)
Supplementary information, Data S1
Analysis of viral integration by PCR (PDF 22 kb)
Rights and permissions
About this article
Cite this article
Sheng, C., Zheng, Q., Wu, J. et al. Direct reprogramming of Sertoli cells into multipotent neural stem cells by defined factors. Cell Res 22, 208–218 (2012). https://doi.org/10.1038/cr.2011.175
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2011.175
Keywords
This article is cited by
-
Human iPSC-Derived Neural Models for Studying Alzheimer’s Disease: from Neural Stem Cells to Cerebral Organoids
Stem Cell Reviews and Reports (2022)
-
Biological importance of OCT transcription factors in reprogramming and development
Experimental & Molecular Medicine (2021)
-
Direct conversion of mouse astrocytes into neural progenitor cells and specific lineages of neurons
Translational Neurodegeneration (2018)
-
Direct reprogramming of fibroblasts into neural stem cells by single non-neural progenitor transcription factor Ptf1a
Nature Communications (2018)
-
Current progress in the derivation and therapeutic application of neural stem cells
Cell Death & Disease (2017)