Abstract
The ubiquitin-proteasome system (UPS) in plants, like in other eukaryotes, targets numerous intracellular regulators and thus modulates almost every aspect of growth and development. The well-known and best-characterized outcome of ubiquitination is mediating target protein degradation via the 26S proteasome, which represents the major selective protein degradation pathway conserved among eukaryotes. In this review, we will discuss the molecular composition, regulation and function of plant UPS, with a major focus on how DELLA protein degradation acts as a key in gibberellin signal transduction and its implication in the regulation of plant growth.
Similar content being viewed by others
The ubiquitin and 26S proteasome pathway
Ubiquitin is a conserved 76-amino acid protein that is conjugated to lysine residues within target proteins and itself via the ubiquitination pathway 1, 2. The ubiquitination pathway is complex and the entire process is under tight regulations from other cellular signaling events. The early step of ubiquitination is carried out through the actions of three enzymes: E1 (the ubiquitin activating enzyme), E2 (the ubiquitin conjugating enzyme) and E3 (the ubiquitin ligase). The E1 hydrolyzes ATP to form a thioester bond with the C-terminal glycine of ubiquitin and transfers the activated ubiquitin to a cysteinyl residue of the E2 enzyme. The E2-ubiquitin can either bind with E3 to directly transfer ubiquitin to substrate protein, or in the case of HECT (homology to E6-AP C terminus) E3s, conjugate the ubiquitin to E3 to form an E3-ubiquitin intermediate, and then transfer the ubiquitin to substrate proteins. In both cases, it is the E3 that dictates the substrate specificity of the ubiquitination process and makes the ubiquitin system a major selective degradation pathway conserved in eukaryotes 3, 4, 5. The ubiquitination process can repeat several times to attach new ubiquitin molecule to the lysine residue of a former ubiquitin, which has already been conjugated to the substrate protein. These reiterated processes lead to the modification of the substrate protein by a ubiquitin chain (referred to as polyubiquitination), which is essential for the 26S proteasome recognition, and leads to the subsequent degradation of the polyubiquitinated substrate 2. The polyubiquitin chain can be disassembled by the activity of DUB (deubiquitinating enzyme) to release ubiquitin moieties that are reused in the next ubiquitination cycle (Figure 1) 6.
The ubiquitin-proteasome pathway for protein degradation. A polyubiquitin chain is synthesized via an enzyme cascade including E1, E2 and E3 enzymes, and removed by DUBs. The polyubiquitin chain serves as a tag to be recognized by the 26S proteasome, which mediates the subsequent protein degradation. Both the ubiquitination and degradation are ATP-dependent processes.
The 26S proteasome is a 2.5-MDa ATP-dependent protease complex that consists of a cylindrical 20S core particle (CP), capped on each end by a 19S regulatory particle (RP) (Figure 2) 7. The 20S CP consists of a stack of two outer α-subunit rings and two proteolytic β-subunit rings to hold the protease activity within the internal chamber. The opening to the CP chamber is sufficiently narrow to make sure only unfolded proteins can enter the chamber and access the active proteolytic sites 8. The 19S RP can be further divided into two components, lid and base (Figure 2), and protein components of RP regulate many activities related with proteasome-dependent degradation, including recognition of ubiquitinated substrates 9, 10, removing and recycling the ubiquitin moieties 11, 12, unfolding and transporting the target protein into the central chamber of CP 13, 14. Previous studies show that some of the 19S RP components have substrate-specific functions in plants. For example, the deficiency of RPN10, a base subunit serving as a ubiquitin receptor, impairs ABA singling by stabilization of the transcription factor ABI5 15.
The structure (left) and a simplified model (right) of the yeast 26S proteasome. The structure of 26S proteasome was reprinted from Proceedings of the National Academy of Sciences, USA 92. The proteolytic active subunits (β1, β2 and β5) are highlighted in red color. Rpt, regulatory particle triple-A ATPase, Rpn, regulatory particle non-ATPase.
Genomic analysis revealed that more than 6% of the Arabidopsis genome (over 1 600 loci) encodes core components of the (UPS) 8. For example, Arabidopsis has two E1s, at least 37 E2s and more than 1 400 potential E3s. Since a large number of E3s exist in plant proteome, it is not surprising to find that most of them are plant-specific enzymes without obvious counterparts in yeast and mammalian cells. The diversity of E3s also suggests that protein degradation control in plants is a vital process to regulate growth and development 16.
Plant E3 ubiquitin ligases
The E3 ubiquitin ligases are encoded by diverse gene families in plants. E3s can carry out the ubiquitination function either as single subunit proteins or multi-subunit protein complexes 4, 16. According to the type of E2-binding domain, the single subunit E3s can be further divided into HECT domain and really interesting new gene (RING)/U-box domain E3s, with different ubiquitin transferring mechanisms 3. The HECT domain is a 350-amino acid protein domain that consists of both a ubiquitin-binding motif and an E2-binding motif. The HECT domain E3 protein family is the smallest E3 subfamily in Arabidopsis genome, with only seven members 8. The RING domain is the most abundant E2 interaction domain in Arabidopsis, which contains approximately 477 single subunit protein members, although it is not known whether all the RING domain proteins can function as E3 ubiquitin ligases 17. The RING domains are characterized by the ∼70-amino acid zinc-binding motif (referred to as RING finger) 18, 19. The U-box domain is a modified form of RING-finger domain with approximately 64 members 20. Unlike the RING-finger domain, the U-box domain does not use zinc ions to maintain its secondary structure, whereas the overall structure of both domains is quite similar and both of them contain a conserved surface for E2 interaction 21. In contrast with HECT domain, E3s that accept activated ubiquitin to form an E3-ubiquitin intermediate and then transfer ubiquitin to the target proteins, the RING/U-box E3s directly catalyze the ubiquitin transfer from E2s to substrate proteins (Figure 3) 3.
HECT and RING/U-box single subunit E3 ubiquitin ligases. During the process of protein ubiquitination, ubiquitin forms an intermediate thioester linkage with HECT E3s before transfer to the lysine residue in the substrate protein. RING E3s do not form an intermediate with ubiquitin, rather, RING E3s provide a scaffold to support the direct transfer of ubiquitin from E2 to the substrate protein.
Cullin-based multi-subunit E3 families
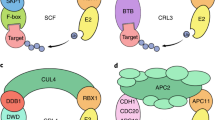
The SCF complexes, a large group of multi-subunit RING domain E3 ligases, are the most abundant and best characterized E3 family 22. The SCF complex is named after three of its four subunits: Skp1 (in plant referred to as ASK), Cullin 1 (CUL1) and the F-box protein. The fourth subunit is the RING-finger-containing protein RBX1 4. In SCF complex, the cullin protein serves as a scaffold that binds RBX1 to its C-terminus and the linker subunit SKP1 to its N-terminal domain. The SKP1 protein in turn interacts with the F-box motif at the N-terminus of F-box proteins to form a complete SCF complex 23. The C-termini of F-box proteins contain protein-protein interaction domains including, but not limited to, WD-40, KELCH and Leu-rich repeat domains. These domains can recruit specific substrates to the SCF complex (Figure 4A) 24. In Arabidopsis, the F-box protein superfamily is encoded by the largest gene family containing more than 700 members, and can be divided into 42 families according to the distinct domain organizations 25. By comparison, only 68 and 74 genes encode F-box proteins in human and mouse genomes, respectively 26.
Besides the SCF E3 complex, there are also other cullin-based RING domain E3 ligase complexes. First, the CUL3-BTB (broad complex/tramtrack/bric-a-brac) E3 ligase complexes. In this group of complex E3 ligases, BTB proteins are used both to recognize substates and to interact with the CUL3 scaffold protein (Figure 4B). Second, the CUL4-DDB (DNA damage-binding) E3 ligase complexes. The CUL4-DDB E3s use WD40 domain-containing DWD proteins for substrate recognition, and use the DDB1 protein to tether DWD proteins to the CUL4 scaffold protein (Figure 4C). Third, the APC (anaphase-promoting complex) E3 contains at least 11 subunits, including the cullin-like protein APC2, the RBX1-like protein APC11, and several substrate-recruting subunits, e.g. APC10, CDC20 (cell division cycle protein 20) and CDH1 (CDC20-homology 1; Figure 4D). Together, all the above mentioned four subtypes of multi-subunit E3 complexes were named as cullin–RING ligases (CRLs).
The assembly and regulation of cullin-based E3 ligases
The assembly and activity of CRL complexes are regulated by a small ubiquitin-related protein RUB (related to ubiquitin) 27. The RUB (also named Nedd8 in animals) protein is also highly conserved among eukaryotes. The RUB protein is attached to the cullin subunit of the CRL complex via an enzymatic cascade similar to that of the ubiquitin pathway. In Arabidopsis, AXR1 (auxin resistant 1) and ECR1 (E1 C-terminal related 1) form a dimer to function as the E1 28. RCE1 (RUB conjugating enzyme 1) serves as the E2 29. RING-finger protein RBX1, a subunit of the SCF complex, is the E3 of RUB conjugation 30, 31. The RUB conjugation (Rubylation) pathway was first identified in Arabidopsis through genetic screen for auxin resistant mutants, and AXR1 is the first protein shown to be required for auxin response 32, 33. Genetic studies in Arabidopsis suggest that RUB conjugation to the cullin subunit is required for the activity of SCF complexes 29, 33, 34. Subsequent studies in mammalian cells also demonstrated that RUB/Nedd8 is essential for the function of SCF complexes 22. At least for the SCF, CUL3-BTB and CUL4-DDB CRLs, RUB modification is highly dynamic and plays an important role in the assembly and disassembly of these CRL E3 complexes 8.
Given the importance of CRL complexes to cellular regulation and the highly dynamic feature of cullin rubylation, it is not surprising that the activity of CRL complexes is tightly regulated by other complexes that antagonize the RUB conjugation pathway. The COP9 signalsome (CSN) was first identified as an essential complex that represses photomorphogenesis, but is now known to have a broad role in plant growth and development 35, 36. The CSN is a conserved multi-protein complex consisting of eight subunits (CSN1-CSN8); it shares structural and sequence similarities to the 19S RP of the proteasome and the eukaryotic translation initiation factor 3 (eIF3) 37. The best-characterized biochemical function of the CSN complex is RUB isopeptidase activity that removes RUB modification from cullin proteins 38. The RUB deconjugation (derubylation) reaction is mediated by CSN5, a zinc metalloprotease, but loss of other CSN subunits also leads to destabilization of the entire CSN complex, causing severe development defects in plants 35. Impaired function of the CSN complex results in loss of cullin derubylation 39, 40. The derubylation activity of CSN directly links this complex to the regulation of SCF E3 ligases 36. Interestingly, the CSN5 partially deficient mutant has increased level of rubylated cullin proteins, but the phenotype of csn5 is quite similar to the axr1 mutant 39. Thus, the fact that both increased and decreased levels of rubylated cullin cause a similar effect on the function of the CRL complex suggests that the dynamic cycling of rubylation and derubylation is required for CRL activity.
CAND1 (cullin-associated and neddylation-dissociated 1) is a protein first identified in animals that can bind unmodified cullin proteins to regulate the activity of SCF complexes 41, 42. In Arabidopsis, the cand1 mutant was discovered by genetic screen and has a pleiotropic phenotype with altered responses to several phytohormones, including gibberellic acid (GA) and auxin 43. CAND1 preferentially binds to derubylated cullin and disrupts the formation of SCF complexes 43, 44, whereas a recent study indicated that, similar to the manner of CSN regulation, both decreased and increased CAND1-CUL1 interactions impaired the function of SCF complexes in vivo 45. These findings suggest that binding dynamics of CAND1 and the cycle of rubylation and derubylation intersect each other to fine tune the activity of CRL complexes.
Physiological function of plant ubiquitin-26S proteasome system (UPS)
The ubiquitin-26S proteasome pathway regulates almost all the aspects of plant growth and development, including, but not limited to, hormone perception and signaling 16, 46, light response 5, flower development 4, 5, self-incompatibility 4, 8, epigenetic regulation 8 and plant pathogenesis and disease control 8.
Light is one of the most important environmental cues for plants; thus, it is reasonable to find that protein degradation through the UPS is widely involved in regulating plant light responses. Both the red and far-red light absorbing photoreceptor PHYA (phytochrome A), the blue light absorbing photoreceptor CRY2 (cryptochrome 2) and the phytochrome interacting factors (PIFs) are proteolysis targets of plant UPS, and their degradation is conventionally regulated by phosphorylation 47, 48. The single-subunit RING-finger E3, COP1, is of critical importance to plant photomorphogenesis. The dark-dependent nucleus translocation of COP1 is responsible for the turnover of a number of transcription factors, including a key photomorphogenic effector protein HY5 (long hypocotyl 5) 49.
Phytohormones (plant hormones) are a structurally unrelated collection of small molecules that control and integrate a wide variety of processes in plant growth and development. Among the photohormones, auxin was the first to be discovered, followed by gibberellins, cytokinins, abscisic acid, ethylene, jasmonates, brassinosteroids and strigolactone. A number of UPS components have been implicated in the regulation of phytohormone responses. In the rest of this review, we will discuss in detail the relationship between ubiquitin-proteasome-mediated proteolysis and gibberellin signal transduction.
Plant hormone GA and development regulation
GA plays an important role in diverse growth and developmental processes throughout the whole life cycle of plants, including seed germination, stem elongation, leaf expansion and flower development 46, 50. GA perception is mediated by GID1 (gibberellin insensitive dwarf 1), a receptor that has similarity to hormone-sensitive lipases (HSLs), but lacks the conserved residues required for enzymatic activity 51. GID1 was originally discovered by genetic screen for GA signaling mutants in rice 52. Later, studies in Arabidopsis identified three orthologs, GID1a, GID1b and GID1c, as GA receptors (Figure 5) 53, 54. Genetic studies demonstrate that both in rice and Arabidopsis, GID1 proteins are essential to perceive GA and trigger all the GA-related responses 52, 53, 54.
GA responses are negatively regulated by DELLA proteins that belong to the GRAS family of putative transcription factors and are named from their N terminal-conserved DELLA motif 46. The DELLA proteins are key plant growth repressors first identified in Arabidopsis and widely distributed in other crop plants, including rice (Oryza sativa), maize (Zea mays), wheat (Triticum aestivum), barley (Hordeum vulgare) and grape (Vitis vinfera) 55, 56, 57, 58, 59, 60. A single gene encodes the DELLA protein SLR1 (slender rice 1) in rice, whereas a family of five genes in the Arabidopsis genome encodes DELLA proteins, including GA insensitive (GAI), repressor of ga1-3 (RGA) and other three repressor of ga1-3-LIKE proteins (RGL1, RGL2 and RGL3; Figure 5). The five DELLA proteins in Arabidopsis have both redundant and partially specialized functions. Genetic studies suggest that RGA and GAI synergistically suppress GA-regulated internode elongation, abaxial trichome initiation and leaf expansion 61, 62, whereas RGL1 and RGL2 are involved in controlling seed germination 63, 64, 65. Moreover, RGA, RGL1 and RGL2 can work together to regulate floral development 65, 66, 67. Recent studies highlighted that DELLA proteins serve as integrators to regulate plant growth and development by integrating the effects of multiple environmental cues, including light 68, 69, 70, 71 and salt 72, 73, 74, and cold 75 and biotic stresses 76.
The UPS in GA signaling
The growth restraint in plant is relieved by GA-induced degradation of DELLA proteins 77, 78, although the kinetics of degradation varies among different homologues of DELLA proteins 79. Based on the inhibition of DELLA degradation by proteasome-specific inhibitors and the existence of polyubiquitinated DELLA proteins, it was generally assumed that GA-induced degradation of DELLA proteins is via the ubiquitin-26S proteasome pathway 71, 80. This proteolysis-based GA signal transduction pathway is highly conserved among higher plants: the GA-induced degradation of DELLA proteins has not only been characterized in Arabidopsis, but also examined in other plant species, such as rice 55 and barley 56, 80.
The DELLA proteins accumulate at high levels in Arabidopsis and rice mutants, sly1-10 and gid2, which have defects in the F-box genes SLY1 (sleepy 1) and GID2 (gibberellin-insensitive dwarf 2), respectively 81, 82, 83. In contrast, a sly1 gain-of-function allele, sly1-d, causing a much stronger interaction with DELLA than the wild-type SLY1 protein, can promote DELLA protein turnover and reduce protein levels of RGA in vivo 77, 79, 84. Both sly1-10 and gid2 mutants exhibit GA-insensitive dwarf phenotypes that can be suppressed by additional loss-of-function mutations of DELLA proteins 81, 82, 85. Furthermore, physical interaction between the F-box protein, SLY1 and the SKP adaptor proteins can be detected through immunoprecipitation assays, supporting that SLY1/GID2 is a functional component of the SCF complex that recruits DELLA proteins for ubiquitination and subsequent degradation by the proteasome 79. Moreover, a recent study demonstrated that the F-box protein SLY1 is directly involved in DELLA protein degradation by using a cell-free assay system 86. Taken together, these results indicate that the SCFSLY1/GID2 E3 ligase complex is responsible for controlling the stability of DELLA proteins.
A cascade of protein-protein interactions triggered by GA perception control the degradation of DELLA proteins to mediate GA responses
DELLA proteins can interact with GID1 in a GA-dependent manner 87. Crystal structure data show that the GA binding site locates in a deep pocket of the GID1 protein, and binding to GA induces the N-terminal lid of GID1 to fold back over the GA-binding pocket to provide a rigid platform for interaction with the conserved N-terminal DELLA/VHYNP/LExLE motifs of DELLA protein (Figure 6) 88, 89. Yeast three-hybrid results showed that the interaction of DELLA protein and GID1 enhances the binding affinity between DELLA proteins and the F-box protein SLY1/GID2 53. A recent study suggests that the C-terminal GRAS domain of DELLA protein can further stabilize the DELLA-GID1 interaction by reducing the dissociation rate, and this stable interaction mediated through the GRAS domain is essential for the recruitment of DELLA protein by the SCFSLY1/GID2 complex 90. All the above data support that the formation of a GA-GID1-DELLA ternary complex promotes the interaction between DELLA and the SCFSLY1/GID2 complex, which results in ubiquitination and subsequent degradation of the DELLA proteins (Figure 6).
Model of GA-induced DELLA protein degradation and the regulation of PIF protein function. The formation of GA-GID1-DELLA complex in the presence of GA promotes the recruitment of DELLA by SCFSLY1/GID2 E3 complex. DELLA protein degradation releases growth-promoting transcription factors from sequestration, enabling previously inactive GA response genes to become activated.
Recent studies shed important light on how DELLA proteins function as a key repressor of plant growth. The data showed that the leucine-heptad-repeat (LZ) domain of DELLA proteins can interact directly with the basic helix-loop-helix (bHLH) DNA-binding domain of PIF3 and PIF4 to sequester these transcription factors in inactive complexes 70, 71. Thus, it is likely that DELLA proteins suppress plant growth, at least partially, through interfering with the functions of other transcription factors that act as positive regulators of plant growth 91.
Conclusion and future perspective
The 2004 Nobel Prize in Chemistry was awarded to Aaron Ciechanover, Avram Hershko and Irwin Rose for their pioneering biochemical studies that led to the discovery of the ubiquitin-mediated protein degradation. The pivotal role of the ubiquitin-26S proteasome pathway in eukaryotes has already become clear. Our understanding of the plant ubiquitin system and GA signaling pathway has expanded exponentially over the past several years, and the link between GA signal transduction and protein degradation has been firmly established. The molecular mechanism of DELLA protein ubiquitination and subsequently degradation by the UPS is a fertile area for future research. The role of the ubiquitin system controlling numerous key regulators in almost every aspect of the life cycle of a plant is certainly not limited to the GA pathway. As more plant E3s and their substrates are characterized, we can expect a better understanding of plant growth and development that will be invaluable for agriculture application, and may even have important implications for biomedical research.
References
Hershko A, Ciechanover A . The ubiquitin system. Annu Rev Biochem 1998; 67:425–479.
Smalle J, Vierstra RD . The ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol 2004; 55:555–590.
Pickart CM . Mechanisms underlying ubiquitination. Annu Rev Biochem 2001; 70:503–533.
Moon J, Parry G, Estelle M . The ubiquitin-proteasome pathway and plant development. Plant Cell 2004; 16:3181–3195.
Sullivan JA, Shirasu K, Deng XW . The diverse roles of ubiquitin and the 26S proteasome in the life of plants. Nat Rev Genet 2003; 4:948–958.
Reyes-Turcu FE, Ventii KH, Wilkinson KD . Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem 2009; 78:363–397.
Finley D . Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu Rev Biochem 2009; 78:477–513.
Vierstra RD . The ubiquitin-26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol 2009; 10:385–397.
Elsasser S, Chandler-Militello D, Muller B, Hanna J, Finley D . Rad23 and Rpn10 serve as alternative ubiquitin receptors for the proteasome. J Biol Chem 2004; 279:26817–26822.
Husnjak K, Elsasser S, Zhang N, et al. Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature 2008; 453:481–488.
Verma R, Aravind L, Oania R, et al. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science 2002; 298:611–615.
Abadi AH, Abouel-Ella DA, Lehmann J, et al. Discovery of colon tumor cell growth inhibitory agents through a combinatorial approach. Eur J Med Chem 2010; 45:90–97.
Navon A, Goldberg AL . Proteins are unfolded on the surface of the ATPase ring before transport into the proteasome. Mol Cell 2001; 8:1339–1349.
Rabl J, Smith DM, Yu Y, Chang SC, Goldberg AL, Cheng Y . Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases. Mol Cell 2008; 30:360–368.
Smalle J, Kurepa J, Yang P, et al. The pleiotropic role of the 26S proteasome subunit RPN10 in Arabidopsis growth and development supports a substrate-specific function in abscisic acid signaling. Plant Cell 2003; 15:965–980.
Santner A, Estelle M . The UPS regulates plant hormone signaling. Plant J 2010; 61:1029–1040.
Stone SL, Hauksdottir H, Troy A, Herschleb J, Kraft E, Callis J . Functional analysis of the RING-type ubiquitin ligase family of Arabidopsis. Plant Physiol 2005; 137:13–30.
Seol JH, Feldman RM, Zachariae W, et al. Cdc53/cullin and the essential Hrt1 RING-H2 subunit of SCF define a ubiquitin ligase module that activates the E2 enzyme Cdc34. Genes Dev 1999; 13:1614–1626.
Freemont PS . RING for destruction? Curr Biol 2000; 10:R84–R87.
Yee D, Goring DR . The diversity of plant U-box E3 ubiquitin ligases: from upstream activators to downstream target substrates. J Exp Bot 2009; 60:1109–1121.
Ohi MD, Vander Kooi CW, Rosenberg JA, Chazin WJ, Gould KL . Structural insights into the U-box, a domain associated with multi-ubiquitination. Nat Struct Biol 2003; 10:250–255.
Petroski MD, Deshaies RJ . Function and regulation of cullin-RING ubiquitin ligases. Nat Rev Mol Cell Biol 2005; 6:9–20.
Zheng N, Schulman BA, Song L, et al. Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 2002; 416:703–709.
Gagne JM, Downes BP, Shiu SH, Durski AM, Vierstra RD . The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc Natl Acad Sci USA 2002; 99:11519–11524.
Xu G, Ma H, Nei M, Kong H . Evolution of F-box genes in plants: different modes of sequence divergence and their relationships with functional diversification. Proc Natl Acad Sci USA 2009; 106:835–840.
Jin J, Cardozo T, Lovering RC, Elledge SJ, Pagano M, Harper JW . Systematic analysis and nomenclature of mammalian F-box proteins. Genes Dev 2004; 18:2573–2580.
Hotton SK, Callis J . Regulation of cullin RING ligases. Annu Rev Plant Biol 2008; 59:467–489.
del Pozo JC, Dharmasiri S, Hellmann H, Walker L, Gray WM, Estelle M . AXR1-ECR1-dependent conjugation of RUB1 to the Arabidopsis Cullin AtCUL1 is required for auxin response. Plant Cell 2002; 14:421–433.
Dharmasiri S, Dharmasiri N, Hellmann H, Estelle M . The RUB/Nedd8 conjugation pathway is required for early development in Arabidopsis. EMBO J 2003; 22:1762–1770.
Gray WM, Hellmann H, Dharmasiri S, Estelle M . Role of the Arabidopsis RING-H2 protein RBX1 in RUB modification and SCF function. Plant Cell 2002; 14:2137–2144.
Kamura T, Conrad MN, Yan Q, Conaway RC, Conaway JW . The Rbx1 subunit of SCF and VHL E3 ubiquitin ligase activates Rub1 modification of cullins Cdc53 and Cul2. Genes Dev 1999; 13:2928–2933.
Lincoln C, Britton JH, Estelle M . Growth and development of the axr1 mutants of Arabidopsis. Plant Cell 1990; 2:1071–1080.
Leyser HM, Lincoln CA, Timpte C, Lammer D, Turner J, Estelle M . Arabidopsis auxin-resistance gene AXR1 encodes a protein related to ubiquitin-activating enzyme E1. Nature 1993; 364:161–164.
Dharmasiri N, Dharmasiri S, Weijers D, Karunarathna N, Jurgens G, Estelle M . AXL and AXR1 have redundant functions in RUB conjugation and growth and development in Arabidopsis. Plant J 2007; 52:114–123.
Serino G, Deng XW . The COP9 signalosome: regulating plant development through the control of proteolysis. Annu Rev Plant Biol 2003; 54:165–182.
Wei N, Serino G, Deng XW . The COP9 signalosome: more than a protease. Trends Biochem Sci 2008; 33:592–600.
von Arnim AG, Schwechheimer C . Life is degrading – thanks to some zomes. Mol Cell 2006; 23:621–629.
Cope GA, Suh GS, Aravind L, et al. Role of predicted metalloprotease motif of Jab1/Csn5 in cleavage of Nedd8 from Cul1. Science 2002; 298:608–611.
Schwechheimer C, Serino G, Deng XW . Multiple ubiquitin ligase-mediated processes require COP9 signalosome and AXR1 function. Plant Cell 2002; 14:2553–2563.
Gusmaroli G, Figueroa P, Serino G, Deng XW . Role of the MPN subunits in COP9 signalosome assembly and activity, and their regulatory interaction with Arabidopsis Cullin3-based E3 ligases. Plant Cell 2007; 19:564–581.
Oshikawa K, Matsumoto M, Yada M, Kamura T, Hatakeyama S, Nakayama KI . Preferential interaction of TIP120A with Cul1 that is not modified by NEDD8 and not associated with Skp1. Biochem Biophys Res Commun 2003; 303:1209–1216.
Zheng J, Yang X, Harrell JM, et al. CAND1 binds to unneddylated CUL1 and regulates the formation of SCF ubiquitin E3 ligase complex. Mol Cell 2002; 10:1519–1526.
Chuang HW, Zhang W, Gray WM . Arabidopsis ETA2, an apparent ortholog of the human cullin-interacting protein CAND1, is required for auxin responses mediated by the SCF (TIR1) ubiquitin ligase. Plant Cell 2004; 16:1883–1897.
Feng S, Shen Y, Sullivan JA, et al. Arabidopsis CAND1, an unmodified CUL1-interacting protein, is involved in multiple developmental pathways controlled by ubiquitin/proteasome-mediated protein Degradation. Plant Cell 2004; 16:1870–1882.
Zhang W, Ito H, Quint M, Huang H, Noel LD, Gray WM . Genetic analysis of CAND1-CUL1 interactions in Arabidopsis supports a role for CAND1-mediated cycling of the SCFTIR1 complex. Proc Natl Acad Sci USA 2008; 105:8470–8475.
Santner A, Estelle M . Recent advances and emerging trends in plant hormone signalling. Nature 2009; 459:1071–1078.
Henriques R, Jang IC, Chua NH . Regulated proteolysis in light-related signaling pathways. Curr Opin Plant Biol 2009; 12:49–56.
Hoecker U . Regulated proteolysis in light signaling. Curr Opin Plant Biol 2005; 8:469–476.
Osterlund MT, Hardtke CS, Wei N, Deng XW . Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 2000; 405:462–466.
Fleet CM, Sun TP . A DELLAcate balance: the role of gibberellin in plant morphogenesis. Curr Opin Plant Biol 2005; 8:77–85.
Hirano K, Ueguchi-Tanaka M, Matsuoka M . GID1-mediated gibberellin signaling in plants. Trends Plant Sci 2008; 13:192–199.
Ueguchi-Tanaka M, Ashikari M, Nakajima M, et al. GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 2005; 437:693–698.
Griffiths J, Murase K, Rieu I, et al. Genetic characterization and functional analysis of the GID1 gibberellin receptors in Arabidopsis. Plant Cell 2006; 18:3399–3414.
Nakajima M, Shimada A, Takashi Y, et al. Identification and characterization of Arabidopsis gibberellin receptors. Plant J 2006; 46:880–889.
Itoh H, Ueguchi-Tanaka M, Sato Y, Ashikari M, Matsuoka M . The gibberellin signaling pathway is regulated by the appearance and disappearance of SLENDER RICE1 in nuclei. Plant Cell 2002; 14:57–70.
Gubler F, Chandler PM, White RG, Llewellyn DJ, Jacobsen JV . Gibberellin signaling in barley aleurone cells. Control of SLN1 and GAMYB expression. Plant Physiol 2002; 129:191–200.
Chandler PM, Marion-Poll A, Ellis M, Gubler F . Mutants at the Slender1 locus of barley cv Himalaya. Molecular and physiological characterization. Plant Physiol 2002; 129:181–190.
Boss PK, Thomas MR . Association of dwarfism and floral induction with a grape 'green revolution' mutation. Nature 2002; 416:847–850.
Peng J, Richards DE, Hartley NM, et al. 'Green revolution' genes encode mutant gibberellin response modulators. Nature 1999; 400:256–261.
Peng J, Carol P, Richards DE, et al. The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 1997; 11:3194–3205.
Dill A, Sun T . Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 2001; 159:777–785.
King KE, Moritz T, Harberd NP . Gibberellins are not required for normal stem growth in Arabidopsis thaliana in the absence of GAI and RGA. Genetics 2001; 159:767–776.
Wen CK, Chang C . Arabidopsis RGL1 encodes a negative regulator of gibberellin responses. Plant Cell 2002; 14:87–100.
Lee S, Cheng H, King KE, et al. Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev 2002; 16:646–658.
Tyler L, Thomas SG, Hu J, et al. Della proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 2004; 135:1008–1019.
Cheng H, Qin L, Lee S, et al. Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 2004; 131:1055–1064.
Yu H, Ito T, Zhao Y, Peng J, Kumar P, Meyerowitz EM . Floral homeotic genes are targets of gibberellin signaling in flower development. Proc Natl Acad Sci USA 2004; 101:7827–7832.
Oh E, Yamaguchi S, Hu J, et al. PIL5, a phytochrome-interacting bHLH protein, regulates gibberellin responsiveness by binding directly to the GAI and RGA promoters in Arabidopsis seeds. Plant Cell 2007; 19:1192–1208.
Achard P, Liao L, Jiang C, et al. DELLAs contribute to plant photomorphogenesis. Plant Physiol 2007; 143:1163–1172.
de Lucas M, Daviere JM, Rodriguez-Falcon M, et al. A molecular framework for light and gibberellin control of cell elongation. Nature 2008; 451:480–484.
Feng S, Martinez C, Gusmaroli G, et al. Coordinated regulation of Arabidopsis thaliana development by light and gibberellins. Nature 2008; 451:475–479.
Magome H, Yamaguchi S, Hanada A, Kamiya Y, Oda K . The DDF1 transcriptional activator upregulates expression of a gibberellin-deactivating gene, GA2ox7, under high-salinity stress in Arabidopsis. Plant J 2008; 56:613–626.
Achard P, Renou JP, Berthome R, Harberd NP, Genschik P . Plant DELLAs restrain growth and promote survival of adversity by reducing the levels of reactive oxygen species. Curr Biol 2008; 18:656–660.
Achard P, Cheng H, De Grauwe L, et al. Integration of plant responses to environmentally activated phytohormonal signals. Science 2006; 311:91–94.
Achard P, Gong F, Cheminant S, Alioua M, Hedden P, Genschik P . The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism. Plant Cell 2008; 20:2117–2129.
Navarro L, Bari R, Achard P, et al. DELLAs control plant immune responses by modulating the balance of jasmonic acid and salicylic acid signaling. Curr Biol 2008; 18:650–655.
Dill A, Jung HS, Sun TP . The DELLA motif is essential for gibberellin-induced degradation of RGA. Proc Natl Acad Sci USA 2001; 98:14162–14167.
Silverstone AL, Jung HS, Dill A, Kawaide H, Kamiya Y, Sun TP . Repressing a repressor: gibberellin-induced rapid reduction of the RGA protein in Arabidopsis. Plant Cell 2001; 13:1555–1566.
Fu X, Richards DE, Fleck B, Xie D, Burton N, Harberd NP . The Arabidopsis mutant sleepy1gar2-1 protein promotes plant growth by increasing the affinity of the SCFSLY1 E3 ubiquitin ligase for DELLA protein substrates. Plant Cell 2004; 16:1406–1418.
Fu X, Richards DE, Ait-Ali T, et al. Gibberellin-mediated proteasome-dependent degradation of the barley DELLA protein SLN1 repressor. Plant Cell 2002; 14:3191–3200.
McGinnis KM, Thomas SG, Soule JD, et al. The Arabidopsis SLEEPY1 gene encodes a putative F-box subunit of an SCF E3 ubiquitin ligase. Plant Cell 2003; 15:1120–1130.
Sasaki A, Itoh H, Gomi K, et al. Accumulation of phosphorylated repressor for gibberellin signaling in an F-box mutant. Science 2003; 299:1896–1898.
Strader LC, Ritchie S, Soule JD, McGinnis KM, Steber CM . Recessive-interfering mutations in the gibberellin signaling gene SLEEPY1 are rescued by overexpression of its homologue, SNEEZY. Proc Natl Acad Sci USA 2004; 101:12771–12776.
Dill A, Thomas SG, Hu J, Steber CM, Sun TP . The Arabidopsis F-box protein SLEEPY1 targets gibberellin signaling repressors for gibberellin-induced degradation. Plant Cell 2004; 16:1392–1405.
Steber CM, Cooney SE, McCourt P . Isolation of the GA-response mutant sly1 as a suppressor of ABI1-1 in Arabidopsis thaliana. Genetics 1998; 149:509–521.
Wang F, Zhu D, Huang X, et al. Biochemical insights on degradation of Arabidopsis DELLA proteins gained from a cell-free assay system. Plant Cell 2009; 21:2378–2390.
Willige BC, Ghosh S, Nill C, et al. The DELLA domain of GA INSENSITIVE mediates the interaction with the GA INSENSITIVE DWARF1A gibberellin receptor of Arabidopsis. Plant Cell 2007; 19:1209–1220.
Murase K, Hirano Y, Sun TP, Hakoshima T . Gibberellin-induced DELLA recognition by the gibberellin receptor GID1. Nature 2008; 456:459–463.
Shimada A, Ueguchi-Tanaka M, Nakatsu T, et al. Structural basis for gibberellin recognition by its receptor GID1. Nature 2008; 456:520–523.
Hirano K, Asano K, Tsuji H, et al. Characterization of the molecular mechanism underlying gibberellin perception complex formation in rice. Plant Cell 2010.
Lau OS, Deng XW . Plant hormone signaling lightens up: integrators of light and hormones. Curr Opin Plant Biol 2010.
Bohn S, Beck F, Sakata E, et al. Structure of the 26S proteasome from Schizosaccharomyces pombe at subnanometer resolution. Proc Natl Acad Sci USA 2010; 107:20992–20997.
Acknowledgements
We apologize to all colleagues whose relevant work could not be included because of space constraints. Work in the authors' laboratory (Beijing) is supported by grants from the Ministry of Science and Technology of China (2006AA10A101; 2006AA10A103; 2009DFB30030) and the Ministry of Agriculture of China (2009ZX08012-021B).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, F., Deng, X. Plant ubiquitin-proteasome pathway and its role in gibberellin signaling. Cell Res 21, 1286–1294 (2011). https://doi.org/10.1038/cr.2011.118
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2011.118
Keywords
This article is cited by
-
Transcriptome alterations of radish shoots exposed to cadmium can be interpreted in the context of leaf senescence
Protoplasma (2023)
-
Stalk architecture, cell wall composition, and QTL underlying high stalk flexibility for improved lodging resistance in maize
BMC Plant Biology (2020)
-
Genome-wide identification and transcriptome profiling reveal that E3 ubiquitin ligase genes relevant to ethylene, auxin and abscisic acid are differentially expressed in the fruits of melting flesh and stony hard peach varieties
BMC Genomics (2019)
-
A transcriptional analysis reveals an extensive range of genes responsible for increasing the tolerance of Carrizo citrange to oxygen deficiency
Tree Genetics & Genomes (2019)
-
Expression of candidate genes related to white mold resistance in common beans
Tropical Plant Pathology (2019)