Abstract
In their seminal publication describing the structure of the DNA double helix 1, Watson and Crick wrote what may be one of the greatest understatements in the scientific literature, namely that “It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material.” Half a century later, we more fully appreciate what a huge challenge it is to replicate six billion nucleotides with the accuracy needed to stably maintain the human genome over many generations. This challenge is perhaps greater than was realized 50 years ago, because subsequent studies have revealed that the genome can be destabilized not only by environmental stresses that generate a large number and variety of potentially cytotoxic and mutagenic lesions in DNA but also by various sequence motifs of normal DNA that present challenges to replication. Towards a better understanding of the many determinants of genome stability, this chapter reviews the fidelity with which undamaged and damaged DNA is copied, with a focus on the eukaryotic B- and Y-family DNA polymerases, and considers how this fidelity is achieved.
Similar content being viewed by others
Introduction
In order to pass genetic information from one generation to the next, all organisms must accurately replicate their genomes during each cell division. This includes the nuclear genome and mitochondrial and chloroplast genomes. These are normally replicated with high fidelity that is achieved through the combined action of accurate DNA polymerases and DNA mismatch repair (MMR). The major replicative DNA polymerases have evolved mechanisms to strongly favor correct over incorrect dNTP incorporation. In addition, several DNA polymerases contain an associated 3′→5′ exonuclease activity that can excise incorrect bases from the growing DNA chain, allowing another attempt at correct synthesis. In the event that the polymerase makes an error that escapes this proofreading activity, post-replication DNA MMR monitors the DNA for errors, excises the error in the newly synthesized strand and then re-synthesizes DNA. In total, these three discrimination steps result in an in vivo mutation rate estimated to be lower than 1 × 10−9, i.e., less than one error for every billion (or more) bases pairs copied (Figure 1A). Moreover, at each step of the process, there are competing forces (Figure 1B) that can affect the fidelity with which DNA is replicated. In this review, we focus on the contributions and mechanisms of DNA polymerase selectivity and proofreading. Readers interested in DNA MMR can consult recent comprehensive reviews of that subject (see refs. 2 and 3; also see the article by Li in this issue). The focus of this chapter will be on eukaryotic DNA polymerases in the B and Y families, with discussion of links to human disease where possible.
Determinants of replication fidelity. (A) The relative contribution levels of the three main components of replication fidelity are shown above the scale, estimated from the mutation rates of systems defective in one or more of the components. The overlapping ovals represent the fact that there is a range of possible increases in the level of fidelity that each mechanism provides dependent on many factors. The range of fidelity that a given mechanism is capable of providing is the critical factor (i.e. MMR can still provide up to four orders of magnitude increase in fidelity for polymerase errors that occur at a frequency of 10−2). The horizontal bars below the graph show the ranges of in vitro determined error rates for the different families of polymerases and the estimated mutation rate range of the in vivo complete replication complex. Within each family, the error rates can differ widely between polymerases and types of errors. The broken bars at the left and right ends indicate that the rates could be even higher and lower than indicated. (B) Graphic depicting the various means by which DNA replication can be modulated. DNA is shown as a stylized double helix (backbone is black and gray), with purine-pyrimidine base pairs indicated as red-green and blue-purple bars. The single-strand region is meant to depict the unwound DNA at a replication fork, with the kink in the DNA representing the bend in the template strand identified by crystallography 119. Red arrows and text indicate conditions that lead to lower fidelity. Green arrows and text indicate conditions that promote higher fidelity, and green bars indicate conditions that block mutations. M=Mutation; C=Correct.
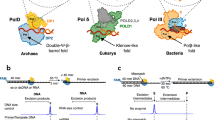
DNA replication requires the combined activity of dozens of proteins 4, a subset of which are shown in Figure 2. Three members of the B-family 5 of polymerases are involved in the bulk of DNA replication, pols α, δ and ε. After the DNA duplex is unwound, likely by the MCM2-7 helicase complex 6, synthesis is initiated on both the leading and lagging strands by the four subunit pol α-primase complex that synthesizes a short RNA-DNA hybrid primer. For leading strand synthesis, a polymerase then binds and extends the primer in a continuous fashion for as long as the polymerase is able to stay bound. For replication of the lagging strand, a discontinuous mode of synthesis occurs in patches of ∼250 base pairs called Okazaki fragments, each of which must be initiated by pol α-primase activity 4. The complexity of the system is illustrated by the fact that five decades after the discovery of the structure of DNA, uncertainty still remains as to the identity of major leading and lagging strand DNA polymerase(s) 4, 7. In mitochondria, pol γ is responsible for all DNA synthesis activities 8, 9. The importance of accurate replication of mitochondrial DNA will be discussed below.
Simplified cartoon model of a eukaryotic replication fork. Protein depictions are based on currently accepted subunit composition of S. cerevisiae proteins but are not meant to be accurate structure-based models. The assignment of pol ε to the leading strand is based on a recent report 120, but has not been definitively established for all replication. Pol δ is consequently assigned to the lagging strand, consistent with earlier reports 121, 122, 123. Helicase hexamer (magenta); replication protein A (RPA; light blue ovals); proliferating cellnuclear antigen (PCNA; purple torus); pol α-primase complex (blue); RNA-DNA hybrid primer (red zig-zag and arrow); pol δ (red); pol ε (green); template strand DNA (black lines); newly synthesized DNA (gray lines). Image inspired by and adapted from Figure 1 in ref. 7 and Figure 7 ref. 4.
In addition to the major replicative polymerases, there are a number of other DNA polymerases that have specialized roles in replicating the nuclear genome. Several of these, including the B-family member pol ζ, and multiple members of the Y-family (η, κ, ι, Rev1) are involved in bypassing DNA lesions that otherwise impede the major replicative polymerases. At least one of these, pol η, not only has the remarkable ability to copy damaged DNA more efficiently than the equivalent undamaged sequence, it can also “sense” that the lesion has been bypassed, triggering a lesion-dependent dissociation from the DNA 10. This results in the simple model for translesion synthesis (TLS) shown in Figure 3A. Other results using different combinations of polymerases and lesions give rise to the “multiple-polymerase” model shown in Figure 3B 11, wherein one polymerase may insert opposite a lesion and another extends from the resulting primer terminus. Given the range of possible lesions and number of TLS polymerases, it is possible that events depicted in both models occur, depending on both the lesion and polymerase involved. Adding another layer of complexity, it may be that TLS can in some instances occur at the replication fork, whereas for other lesions, bypass may occur during gap-filling after the fork has moved on (Figure 3C and 3D) 12. There is currently no data indicating that either the 1 or 2 polymerase model would be specific to a certain timing of TLS. In addition to their roles in TLS, many of the specialized polymerases have also been implicated in other DNA transactions, including somatic hypermutation (SHM; pols ζ, η, ι, Rev1) 13, 14, homologous recombination (pol η) 15, nucleotide excision repair (pol κ) 16 and base excision repair (pol ι) 17. The Y-family member Rev1 is a G-template specific deoxycytidyl transferase and it has a non-catalytic role in TLS as well 18, 19. Rev1p interacts with multiple polymerases 20 and is required for in vivo mutagenesis, although the transferase activity of the enzyme is dispensable 21. In addition, although not covered in detail here (except for the family A member pol γ), there exist several mammalian family A and X polymerases with widely varying fidelities and whose in vivo functions are the subject of intense investigation 5.
Models of translesion synthesis. (A) The 1-polymerase model of TLS, shown here for a thymine-thymine dimer, states that a single polymerase is responsible for the complete bypass of a lesion, including insertion opposite all lesion bases and extension from the primer terminus opposite a damaged template base. (B) The 2-polymerase model of TLS, shown here for a thymine-thymine 6-4 photoproduct, states that different polymerases are responsible for the insertion steps at the various lesion positions. In the example given, note that while pol ζ is responsible for extension from the template-3′ T primer terminus, it also carries out an insertion at the 5′ T position of the lesion. For a single base lesion, the insertion step would be opposite undamaged DNA. A more comprehensive listing of 2-polymerase/lesion combinations is given elsewhere 11. Note that for both examples given, the actual TLS reaction is flanked relatively closely both upstream (1-2 bases) and downstream (1-5 bases) of the lesion by replicative polymerase synthesis. (C) Model for TLS that occurs at a replication fork during the process of ongoing synthesis. (D) Model for TLS that takes place as a “gap-filling” reaction, away from the main replication machinery. Note that both of these models are consistent with either the 1- or 2-polymerase model of TLS given in panels A and B. In both cases, post-translational modification of PCNA and possible other proteins is critical for the polymerase switch. Note that panels A and B are models of the actual TLS process while panels C and D depict models for the timing of TLS. As such (and as noted in the text), there is overlap between the panels.
With this brief background, we first consider DNA replication fidelity and how defects in the steps required for high fidelity lead to genome instability (Figure 1B). After describing the fidelity of the major A- and B-family replicative polymerases, we then discuss how nucleotide selectivity and proofreading help maintain genome stability during DNA synthesis. We then consider the role that replication accessory proteins have in modulating DNA synthesis fidelity, followed by a discussion of the fidelity of the TLS polymerases and the lesion bypass process. We conclude with a discussion of how defects in several of these pathways are associated with human disease.
High fidelity of eukaryotic replicative DNA polymerases when copying undamaged DNA
The bulk of DNA synthesis in a eukaryotic cell occurs during replication of undamaged DNA templates. This synthesis is catalyzed by the B-family polymerases α, δ and ε for nuclear DNA and by the A-family polymerase γ for mitochondrial DNA. These four DNA polymerases are highly accurate, generating on average less than one single base substitution or single base insertion/deletion (indel) for every 10 000 correct incorporation events (Table 1). Such low error rates are consistent with other reports using homologous polymerases and different methods of analysis 22, 23, 24, 25. The rates listed in Table 1 are averages for all 12 possible single base-base mismatches and for a variety of single base indels in different sequence contexts. Rates for individual base substitutions and indels can vary by more than 100-fold, depending on the composition of the mismatch and the DNA sequence flanking the mismatch (error rate ranges are shown as purple boxes in Figure 1A). Each of the different polymerases listed in Table 1 also differs to some degree from the others regarding error specificity (see refs. listed in Table 1 for examples).
High nucleotide selectivity
Consistent with the need to accurately replicate billions of base pairs during every cell division cycle, the major replicative polymerases almost always initially insert the correct dNTP onto properly aligned primer templates. Among the three major steps, high selectivity against misincorporation provides the single greatest contribution to replication fidelity. This is illustrated by the low base substitution and indel error rates of pol α, which naturally lacks proofreading activity, and by the low error rates of derivatives of pols δ, ε and γ that have been engineered to inactivate their intrinsic proofreading activities (Table 1, top). The fidelity of all four enzymes is much higher than that of polymerases involved in translesion DNA synthesis, which are also naturally exonuclease-deficient (Table 1, bottom). The importance of high nucleotide selectivity to genome stability, and its relationship to disease outcomes, is illustrated by recent studies of the effects of conservative amino acid substitutions in the active site of replicative DNA polymerases. When a highly conserved residue in the active site of S. cerevisiae B-family polymerases that is predicted to interact with the incoming dNTP is replaced with a different amino acid, the mutant enzymes have decreased DNA synthesis fidelity in vitro and they generate mutator phenotypes in vivo 26, 27, 28, 29, 30. The amino acid changes introduced (L868F in pol α, L612M in pol δ and M644F in pol ε) do not greatly affect the overall activity of the polymerases, and error rates are elevated for only certain types of errors that differ among the polymerases. Mice with replacements for the homologous residue (L604G or L604K) in murine pol δ display homozygous lethality, and heterozygotes have a decreased lifespan, increased genomic instability and accelerated tumorigenesis 31. A second example of the importance of polymerase selectivity is the Y955C substitution in human pol γ. This change reduces pol γ fidelity in vitro when copying undamaged DNA and when bypassing template 7,8-dihydro-8-oxo-guanine (8-oxo-dG) 32, 33, and the Y955C substitution is associated with autosomal dominant progressive external ophthalmoplegia (PEO) 8.
How is high nucleotide selectivity achieved? Hydrogen bonding between template bases and incoming dNTPs is clearly important for replication fidelity 34. However, this alone is unlikely to explain high selectivity, since the free energy difference between correct and incorrect base pairs in solution 35, 36, 37 accounts for error rates of ∼1:100, while the selectivity of the exonuclease-deficient major replicative polymerases is much greater than this (Table 1). Several ideas have been put forth to account for high selectivity. One is enthalpy-entropy compensation 36, 38. In order for the incoming dNTP to hydrogen bond to a template base, water molecules that are hydrogen bonded to the base of the incoming dNTP must be stripped away, thereby decreasing the entropy of the system. This magnifies the contribution of enthalpy to the free energy difference (ΔΔG°=ΔΔH°-TΔΔS°), thereby increasing nucleotide selectivity. Consistent with this idea, the error rates of Y-family enzymes (Table 1, bottom), which have solvent accessible active sites (see below), are in the range predicted if enthalpy-entropy compensation was not contributing to selectivity. Kinetic analysis of insertion of non-polar base analogs by yeast pol η further supports that they do not use enthalpy-entropy compensation to increase selectivity 39.
A wealth of evidence supports the idea that the high nucleotide selectivity of accurate DNA polymerases results partly from the exquisite shape complementarity of their nascent base pair binding pockets 34, 36, 37, 40. The four canonical Watson-Crick base pairs are nearly identical in size and shape, and numerous structural studies reveal that these correct base pairs fit snugly with the binding pocket, without steric clashes 34, 41. Among many beautiful and informative structures now available (alas, none yet with pols α, δ, ε or γ), Figure 4B and 4D shows the structure of T7 DNA polymerase, a highly accurate family A homolog of Pol γ, with a correct base pair bound in its nascent base pair binding pocket. While the correct pair fits snugly in the active site, the presence of mismatches, which have different and variable geometries 40, 42, is predicted to create steric clashes that would (1) reduce incorrect dNTP-binding affinity, (2) affect subsequent conformational changes needed to set up the proper geometry for catalysis, and/or (3) reduce the rate of phosphodiester bond formation, i.e., chemistry. The relative contribution of these three parameters to the fidelity of DNA replication has and continues to be the subject of many structural and kinetic investigations that elegantly employ mutant and wild-type DNA polymerases and modified dNTPs. Of particular recent interest in the field is the extent to which incorrect incorporation is limited by chemistry or by a dNTP-induced conformational change that has been inferred from kinetic studies. The latter possibility also raises the important issue of the nature of the relevant conformational change among many that occur to assemble the active site, and whether the rate-limiting conformational change differs for different replication errors. These issues are not considered in further detail here because they have been discussed at length in recent articles that interested readers can consult 43, 44, 45.
Open and closed active sites of low and high fidelity polymerases. (A and B) Molecular surfaces of Dpo4 from S. solfataricus and T7 DNA polymerase with blue representing positively charged regions and red representing negatively charged regions are shown. Note the tighter fit of DNA (ball and stick model) to the higher fidelity T7 DNA polymerase, evidenced by closer contact with polymerase regions. In Dpo4, the DNA is located at further distance from the polymerase, and a “hole” in the polymerase structure is visible, indicating a much looser association of the DNA with the polymerase. (C and D) Closeup of the active site showing the templating base and nucleotide. Again, note the relatively open and solvent accessible region in Dpo4 compared to the snug fit in T7 DNA polymerase. Also note the increased amount of neutral region of protein in T7 DNA polymerase, indicating that it is the geometry of the replicative polymerase active site that plays an important role in fidelity, as noted in the text. This figure appears originally as Figure 7 in 124 and has been reproduced with the permission of the authors.
Polymerization errors due to substrate misalignments
In addition to direct misincorporation of an incorrect dNTP resulting in a base substitution error, all four major eukaryotic replicative polymerases (and TLS polymerases) also insert and delete one or more nucleotides during DNA synthesis. These errors result from strand misalignments that generate one or more unpaired bases, either in the primer strand, leading to additions, or in the template strand, leading to deletions. Several ideas have been proposed to account for how these misalignments initiate and are stabilized for continued synthesis to generate a mutation, including DNA strand slippage 46, misinsertion followed by primer relocation 47, melting-misalignment 48 and misalignment of a nucleotide at the active site 49, also referred to as dNTP stabilized misalignment 50. Extensive biochemical and initial structural support for several of these three models has recently been comprehensively reviewed 51.
The exonuclease-deficient major eukaryotic replicative DNA polymerases generate single base deletions at rates that are similar to those for single base substitutions (Table 1, top). Single base deletion error rates are typically substantially (10-fold or more) higher than for single base insertions, with possible explanations as discussed in ref. 51. It is also important to note that the “average” error rates in Table 1 are perhaps somewhat misleading, because the indel error rates of the replicative polymerases are highly dependent on sequence context, being higher in repetitive rather than in non-repetitive sequences and increasing as the length of a repetitive sequence increases 51, 52, just as predicted by Streisinger et al. 46 over 40 years ago. For this reason (and see proofreading section below), long repetitive sequences in eukaryotic genomes are “hot spots” for replication errors (reviewed in 51).
Contribution of proofreading by intrinsic 3′ exonuclease activity to genome stability
Once an incorrect dNTP is incorporated into DNA, the mismatched primer terminus is more difficult to extend than is a correctly paired and properly aligned primer terminus. Extension of such aberrant termini occurs with lower efficiency than does extension of a matched terminus. For mismatch extension, the geometric and kinetic considerations mentioned above are important. The delay in extension caused by a mismatch allows the primer terminus to fray and move to the 3′ exonuclease active site (if present) for excision of the incorrect nucleotide 37, 38. Interestingly, among many mammalian DNA polymerases, only those responsible for the bulk of chain elongation during replication (pols δ, ε and γ) contain intrinsic 3′ exonucleolytic proofreading activity. The contribution of proofreading to base substitution fidelity is illustrated by the lower error rates of the exonuclease-proficient pols δ, ε and γ as compared to their exonuclease-deficient derivatives (Table 1; see also ref. 25 and references therein that demonstrate similar high fidelity for pol δ and pol ε isolated from mammalian sources). Although not obvious from the “less than or equal to” error rates of the exonuclease-proficient wild-type polymerases listed in Table 1, a variety of in vitro studies indicate that proofreading improves replication fidelity by factors ranging from a few-fold to more than 100-fold, depending on the mismatch, the sequence context and the polymerase. The critical role of proofreading in maintaining eukaryotic genome stability is illustrated by genetic studies of yeast strains harboring genes for exonuclease-deficient pol δ, ε and γ, all of which have a mutator phenotype 53, 54, 55, 56, 57. The importance of proofreading to suppressing tumorigenesis is suggested by seminal studies showing that mice harboring exonuclease-deficient pol δ have a shortened life span and increased susceptibility to several types of cancer 58, 59. Also in mice, inactivating the 3′ exonuclease of pol γ elevates levels of mitochondrial DNA mutations and leads to loss of mitochondria and premature ageing 60, 61.
Proofreading also corrects misaligned intermediates containing extra bases in one strand or the other near the primer terminus, as illustrated by the higher indel error rates of exonuclease-deficient pols δ, ε and γ (Table 1, top) when compared with their proofreading proficient counterparts (Table 1, middle). However, the efficiency of proofreading of indels decreases as the length of a repetitive sequence increases, both in vitro 62 and in vivo 63. This is because the extra base in the misalignment substrate is protected from excision, since it can be located far upstream of the polymerase active site. Such diminished proofreading further contributes to the observation that long repetitive sequences are at risk for a high rate of replication errors, as evidenced by the now well-known “microsatellite instability” phenotype of eukaryotic cells defective in DNA MMR, especially tumors from humans and mice with mutations that inactivate MMR.
“Extrinsic proofreading” may also contribute to genome stability
Assuming that pol α, which lacks its own proofreading activity, synthesizes 10 nucleotides of each ∼250-nucleotide Okazaki fragment on the lagging strand, it synthesizes about 2% of the human genome. Given its base substitution error rate of ∼10−4 (Table 1, top), this amount of replication would generate 12 000 mismatches during each replication cycle. This leads to the issue of whether such a potentially heavy load of replication errors might be offset by proofreading of pol α errors by a separate exonuclease, a process referred to as extrinsic proofreading. This possibility was recently examined in a genetic study of yeast pol α with a Leu868Met substitution at the polymerase active site 64. L868M pol α copies DNA in vitro with normal activity and processivity but with reduced fidelity. In vivo, the pol1-L868M allele confers a mutator phenotype. This mutator phenotype is strongly increased upon inactivation of the 3′ exonuclease of pol δ but not that of pol ε. Among several non-exclusive explanations that were considered, the results support the hypothesis that the 3′ exonuclease of pol δ proofreads errors generated by pol α during initiation of Okazaki fragments. Given that eukaryotes encode many other specialized, naturally proofreading-deficient DNA polymerases with even lower fidelity than pol α 5, extrinsic proofreading could be relevant to several other DNA transactions that control genome stability 65, such as base excision repair and possibly TLS by pol η (see below).
Modest contribution of accessory proteins to eukaryotic replication fidelity
In addition to polymerases, DNA replication requires the coordinated action of a large number of other proteins 4 (Figure 2). Thus, it is reasonable to ask whether these proteins influence replication fidelity. Although relatively few investigations have been performed to investigate this subject with eukaryotic replication accessory proteins, experiments to date (see ref. 66 and references therein) suggest that replication accessory proteins like polymerase clamps and single-stranded DNA-binding proteins generally have relatively small effects on fidelity (i.e., typically a few-fold) in comparison to the large contribution of the polymerases themselves. In maintaining the focus of this review on the eukaryotic replication, one notable exception is the ability of RPA and PCNA to strongly suppress formation of large deletion errors occurring between direct repeat sequences during synthesis in vitro by pol δ 67. PCNA and RPA may suppress large deletions by preventing the primer terminus from fraying and/or by preventing primer relocation and annealing to the downstream repeat.
Fidelity of TLS polymerases when copying undamaged DNA
Complete replication of the nuclear genome occasionally requires TLS by specialized polymerases, including family B pol ζ and family Y pols η, κ and ι. These polymerases all lack proofreading activity and they also have lower nucleotide selectivity than the major replicative polymerases, as indicated by their higher error rates for base substitutions and indels (Table 1, bottom). The extreme case is for pol ι, which although reasonably accurate in preventing some mismatches (e.g., error rate of ∼10−4 for A•dCTP), preferentially inserts dGTP more often than the correct dATP opposite template T 68, 69. Structural studies suggest that the low fidelity of family Y enzymes is partly due to relaxed geometric selectivity in the nascent base pair binding pocket, which is more open and solvent accessible (e.g., see Sso Dpo4; Figure 4A and 4C) than those of more accurate DNA polymerases. This fact is likely relevant to the ability of family Y polymerases to more efficiently bypass lesions that distort helix geometry than can the major replicative polymerases (see below). Pols η and κ are not only error-prone for base substitution but also for indels (Table 1, bottom). Indeed, overexpression of pol κ in cultured cells increases indel mutations 70. While not the focus of this review, much of the work on family Y polymerases has been performed using bacterial enzymes, whose functions and properties are reviewed elsewhere 71, 72.
Another TLS polymerase is the B-family member pol ζ. When copying undamaged DNA, pol ζ has somewhat higher fidelity than the Y-family polymerases, but lower fidelity than the other B-family members (Table 1, bottom). The ability of pol ζ to generate both base substitutions and indels at relatively high rates is consistent with its known role in generating a large majority of spontaneous mutations, as well as mutations induced by a variety of DNA-damaging agents 21. The high base substitution error rate of pol ζ clearly demonstrates that it has low nucleotide selectivity, consistent with a possible direct role in mutagenic misinsertion of dNTPs in vivo. Also relevant are kinetic studies demonstrating that pol ζ efficiently extends terminal mismatches 21, 73, 74. This is true with undamaged DNA as well as for extending primer termini opposite damaged template bases, the latter being consistent with a role for pol ζ in the extension step of TLS in a 2-polymerase model (Figure 3B). A similar role has also been proposed for pol κ, which like pol ζ, is promiscuous for mismatch extension 75. During in vitro DNA synthesis, pol ζ also generates “complex” mutations that contain multiple substitutions and indels within a short tract of DNA 76. Consistent with this property, pol ζ also generates complex errors in vivo 77, 78. Most recently, two other eukaryotic DNA polymerases have been shown to have low fidelity and are implicated in TLS, pol ν 79, 80 and pol θ 81, 82. Interestingly, both are members of the family A group of polymerases, other members of which are high fidelity enzymes (e.g., T7 DNA polymerase, pol γ).
The fidelity of TLS
There are numerous reports describing the TLS ability and fidelity of various Y-family (and other) polymerases when encountering a wide range of structurally diverse lesions 5, 83, 84, 85. These studies show that for a given lesion, the bypass efficiency and fidelity is polymerase specific. Many of these reports focus on the specificity and kinetics with which polymerases insert individual correct or incorrect dNTPs opposite lesions, and/or their ability to extend matched and mismatched termini opposite damaged bases. These studies have been extremely valuable towards understanding two of the multiple steps needed to completely bypass lesions. Rather than exhaustively review the details of these studies, here we focus on what is known about error rates for complete bypass reactions that require insertion and multiple extensions and are performed in the presence of all four dNTPs. Since such studies are fewer in number, we discuss three common and biologically important lesions with different coding abilities: an abasic site (non-coding), a cis-syn cyclobutane thymine-thymine dimer (TTD, which retains base coding potential) and 8-oxo-dG (highly ambiguous coding potential).
Since an abasic site contains no base coding information for a polymerase to use to direct dNTP insertion, it is not surprising that the “fidelity”, or more appropriately the “specificity”, of abasic site bypass by multiple polymerases is low. Like a number of other polymerases 86, the archeabacterial DinB homolog Dpo4 of Sulfolobus solfataricus preferentially inserts dAMP opposite an abasic site, and like many other DinB homologs, also generates a significant level of −1 deletions 87. Structurally, this has been suggested to occur because the polymerase “loops out” the abasic site and instead copies the next available base 88. Human pol η displays very similar abasic site bypass characteristics 87. By comparison, a crystal structure of the B-family polymerase from bacteriophage RB69 with a template abasic site in the active site shows that while dAMP is inserted opposite the abasic site, the enzyme was unable to extend the resulting primer terminus, with no translocation of the DNA occurring and no “closing” of the polymerase 89. This helps explain why the efficiency of bypass of abasic sites by replicative polymerases is low.
One of the most well-studied lesions to date has been the TTD, one of the major DNA lesions resulting from exposure to UV radiation. TTDs contain two covalently linked thymines, both of which retain base coding potential. The early characterization of TTD bypass by pol η was termed “error-free” because dAMP was inserted more frequently than other nucleotides 90. Subsequent studies have determined that the fidelity for TTD bypass by yeast and human pol η is actually quite low, e.g., one dGMP incorporated opposite the 3′-T of the dimer for ∼30 dAMP incorporations. This high error rate is similar to that observed when pol η copies the corresponding undamaged thymine 91, 92. The fidelity of homologous Sso Dpo4 is even lower (approaching 1 error in every 10 bypass events), and in this case the error rate is far higher than for copying the equivalent undamaged base 91. Interestingly, both enzymes are more accurate when copying the 5′-T of the TTD. A possible explanation for higher fidelity at the 5′-T is provided by crystal structures of Dpo4 bound to TTD-containing templates 93. Incoming ddATP is paired with the 3′-T of the dimer in a normal Watson-Crick pair, while ddATP orients in a syn configuration to accommodate the template distortion resulting from covalent linkage to the 3′-T, such that the ddATP is paired with the 5′-T of the dimer in a Hoogsteen pair. It may be that Hoogsteen bonded mispairs are less stable, thereby disfavoring errors at this position.
Another well studied lesion is 8-oxo-dG, a common lesion generated by oxidative stress. 8-oxo-dG in an anti-configuration can pair correctly with dCMP, while in a syn configuration it pairs incorrectly with dAMP 94, 95. Kinetic studies of mammalian pol δ have shown that although dCMP is inserted more efficiently opposite the lesion, it is the 8-oxo-dG:dAMP pair that is preferentially extended 96. Moreover, the 8-oxo-dG:dAMP pair interacts with bacteriophage T7 DNA polymerase in a manner that allows this mispair to escape proofreading 97, thereby further increasing the mutagenic potential of this lesion. Readers interested in further information of mutagenesis resulting from 8-oxo-dG can consult excellent reviews on this subject 98, 99.
The biological relevance of TLS is best illustrated by the observation that loss of pol η function in humans and in mice results in sensitivity to sunlight and predisposition to skin cancer 100, 101. Knocking out mouse pol ι in a pol η-deficient background further increased susceptibility to UV light-induced skin cancer, suggesting that pol ι also has a role in bypassing UV photoproducts 102. Studies in cultured cells also implicate pol ι in UV-induced mutagenesis either when pol η is absent or when present 103, 104. Pol ι has also been linked to susceptibility of urethane-induced lung cancers 105. The importance of 8-oxo-dG bypass fidelity is illustrated by the pol γ mutant described above (Y955C), which has a higher propensity for inserting 8-oxo-dGTP and for bypassing template 8-oxo-dG in a mutagenic manner 33. Patients with this mutation have an autosomal dominant form of PEO that manifests with a large number of clinical phenotypes. Finally, while the loss of pol ζ results in embryonic lethality in mice 106, 107, 108, cells lacking pol ζ display severe chromosomal instability 109. These examples clearly demonstrate the importance of the TLS polymerases in maintaining genomic stability. While these examples predict an anti-mutagenic effect of the polymerases, the process of SHM during the generation of immunoglobulin diversity requires the mutagenic properties of these polymerases for normal function. In fact, a recent model of SHM invokes both pols η and ζ, possibly in the bypass of abasic sites 13, 14. Clearly, the mechanisms that control where, when and which TLS polymerases have access to primer templates are critical to genomic stability.
Controlling TLS
The low fidelity of family Y polymerases when copying undamaged DNA implies the need to strictly limit their activities to specific circumstances, or too much error-prone synthesis could lead to error catastrophe and cell death. One means of control is through the polymerase clamp PCNA. Studies have shown that all three eukaryotic Y-family polymerases interact with PCNA 110, 111, 112, and it is known that upon replication fork stalling induced by DNA damage, PCNA becomes mono-ubiquitylated 113. It is currently thought that this post-translational modification is important for delivering TLS polymerases to sites of damage. However, no studies to date have indicated that PCNA, whether unmodified or mono-ubiquitylated, affects the fidelity of TLS polymerases. The eukaryotic TLS polymerases also interact with Rev1 20, which as mentioned above plays a critical role in mutagenesis independent of its DNA synthetic capacity. It is possible that Rev1, which binds to single-stranded DNA and primer termini 114, could help deliver the TLS polymerases to gaps or sites of stalled replication forks, in effect becoming a polymerase accessory protein. Again, however, there is currently no evidence that Rev1/polymerase interactions alter the fidelity of TLS. Thus, the few studies that have been carried out so far come to the same conclusions as the results with the major replicative polymerases, namely that the polymerases themselves are the prime determinants of the fidelity of TLS 92, 111.
Concluding remarks
It has been hypothesized that multistage carcinogenesis requires a mutator phenotype 115. This idea is now supported by examples wherein defects in several pathways that determine DNA replication fidelity result in decreased genome stability and increased susceptibility to cancer. These defects include reduced dNTP selectivity 31, loss of the 3′ exonuclease activity of a replicative polymerase 58, 59, defects in other nucleases involved in DNA replication 116, defects in MMR 2, 3 and defects in TLS 101, 102. While cancer is the most cited example, a number of other diseases also have some connections to defects in replication fidelity and genomic stability. These include (but are not limited to) Alzheimers, Cockayne's Syndrome, Parkinson's disease, Friedreich's ataxia, Huntington's disease, tricothiodystrophy, a number of progressive neuropathies and multiple mitochondrial wasting diseases 117, 118. Clearly, much work remains to be done in this area, with the goal of not only better understanding but also ultimately preventing and/or treating these diseases.
Abbreviations
- (MMR):
-
mismatch repair
- (pol):
-
polymerase
- (TLS):
-
translesion synthesis
- (SHM):
-
somatic hypermutation
- (indel):
-
insertion/deletion
- (8-oxo-dG):
-
7,8-dihydro-8-oxo-guanine
- (PEO):
-
progressive external ophthalmoplegia
- (Dpo4):
-
DNA polymerase 4 from Sulfolobus solfataricus
- (TTD):
-
cis-syn thymine-thymine cyclobutane pyrimidine dimer
References
Watson JD, Crick FH . Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 1953; 171:737–738.
Jiricny J . The multifaceted mismatch-repair system. Nat Rev Mol Cell Biol 2006; 7:335–346.
Kunkel TA, Erie DA . DNA mismatch repair. Annu Rev Biochem 2005; 74:681–710.
Johnson A, O'Donnell M . Cellular DNA replicases: components and dynamics at the replication fork. Annu Rev Biochem 2005; 74:283–315.
Bebenek K, Kunkel TA . Functions of DNA polymerases. Adv Protein Chem 2004; 69:137–165.
Forsburg SL . Eukaryotic MCM proteins: beyond replication initiation. Microbiol Mol Biol Rev 2004; 68:109–131.
Garg P, Burgers PM . DNA polymerases that propagate the eukaryotic DNA replication fork. Crit Rev Biochem Mol Biol 2005; 40:115–128.
Graziewicz MA, Longley MJ, Copeland WC . DNA polymerase γ in mitochondrial DNA replication and repair. Chem Rev 2006; 106:383–405.
Falkenberg M, Larsson N-G, Gustafsson CM . DNA replication and transcription in mammalian mitochondria. Annu Rev Biochem 2007; 76:679–699.
McCulloch SD, Kokoska RJ, Kunkel TA . Efficiency, fidelity and enzymatic switching during translesion DNA synthesis. Cell Cycle 2004; 3:580–583.
Prakash S, Prakash L . Translesion DNA synthesis in eukaryotes: a one- or two-polymerase affair. Genes Dev 2002; 16:1872–1883.
Lehmann AR, Fuchs RP . Gaps and forks in DNA replication: rediscovering old models. DNA Repair (Amst) 2006; 5:1495–1498.
Diaz M, Lawrence C . An update on the role of translesion synthesis DNA polymerases in Ig hypermutation. Trends Immunol 2005; 26:215–220.
Casali P, Pal Z, Xu Z, Zan H . DNA repair in antibody somatic hypermutation. Trends Immunol 2006; 27:313–321.
Kawamoto T, Araki K, Sonoda E, et al. Dual roles for DNA polymerase η in homologous DNA recombination and translesion DNA synthesis. Mol Cell 2005; 20:793–799.
Ogi T, Lehmann AR . The Y-family DNA polymerase κ (pol kappa) functions in mammalian nucleotide-excision repair. Nat Cell Biol 2006; 8:640–642.
Prasad R, Bebenek K, Hou E, et al. Localization of the deoxyribose phosphate lyase active site in human DNA polymerase ι by controlled proteolysis. J Biol Chem 2003; 278:29649–29654.
Haracska L, Prakash S, Prakash L . Yeast Rev1 protein is a G template-specific DNA polymerase. J Biol Chem 2002; 277:15546–15551.
Nelson JR, Lawrence CW, Hinkle DC . Deoxycytidyl transferase activity of yeast REV1 protein. Nature 1996; 382:729–731.
Guo C, Fischhaber PL, Luk-Paszyc MJ, et al. Mouse Rev1 protein interacts with multiple DNA polymerases involved in translesion DNA synthesis. EMBO J 2003; 22:6621–6630.
Lawrence CW, Gibbs PE, Murante RS, et al. Roles of DNA polymerase ζ and Rev1 protein in eukaryotic mutagenesis and translesion replication. Cold Spring Harb Symp Quant Biol 2000; 65:61–69.
Einolf HJ, Guengerich FP . Kinetic analysis of nucleotide incorporation by mammalian DNA polymerase δ. J Biol Chem 2000; 275:16316–16322.
Hashimoto K, Shimizu K, Nakashima N, Sugino A . Fidelity of DNA polymerase δ holoenzyme from Saccharomyces cerevisiae: the sliding clamp proliferating cell nuclear antigen decreases its fidelity. Biochemistry 2003; 42:14207–14213.
Kunkel TA, Sabatino RD, Bambara RA . Exonucleolytic proofreading by calf thymus DNA polymerase δ. Proc Natl Acad Sci USA 1987; 84:4865–4869.
Thomas DC, Roberts JD, Sabatino RD, et al. Fidelity of mammalian DNA replication and replicative DNA polymerases. Biochemistry 1991; 30:11751–11759.
McElhinny SAN, Stith CM, Burgers PMJ, Kunkel TA . Inefficient proofreading and biased error rates during inaccurate DNA synthesis by a mutant derivative of Saccharomyces cerevisiae DNA polymerase δ. J Biol Chem 2007; 282:2324–2332.
Niimi A, Limsirichaikul S, Yoshida S, et al. Palm mutants in DNA polymerases α and η alter DNA replication fidelity and translesion activity. Mol Cell Biol 2004; 24:2734–2746.
Pursell ZF, Isoz I, Lundstrom EB, Johansson E, Kunkel TA . Regulation of B family DNA polymerase fidelity by a conserved active site residue: characterization of M644W, M644L and M644F mutants of yeast DNA polymerase ε. Nucl Acids Res 2007; 35:3076–3086.
Venkatesan RN, Hsu JJ, Lawrence NA, Preston BD, Loeb LA . Mutator phenotypes caused by substitution at a conserved motif A residue in eukaryotic DNA polymerase δ. J Biol Chem 2006; 281:4486–4494.
Li L, Murphy KM, Kanevets U, Reha-Krantz LJ . Sensitivity to phosphonoacetic acid: a new phenotype to probe DNA polymerase δ in Saccharomyces cerevisiae. Genetics 2005; 170:569–580.
Venkatesan RN, Treuting PM, Fuller ED, et al. Mutation at the polymerase active site of mouse polymerase δ increases genomic instability and accelerates tumorigenesis. Mol Cell Biol 2007; 27:7669–7682.
Ponamarev MV, Longley MJ, Nguyen D, Kunkel TA, Copeland WC . Active site mutation in DNA polymerase γ associated with progressive external ophthalmoplegia causes error-prone DNA synthesis. J Biol Chem 2002; 277:15225–15228.
Graziewicz MA, Bienstock RJ, Copeland WC . The DNA polymerase γ Y955C disease variant associated with PEO and parkinsonism mediates the incorporation and translesion synthesis opposite 7,8-dihydro-8-oxo-2′-deoxyguanosine. Hum Mol Genet 2007; 16:2729–2739.
Kool ET . Active site tightness and substrate fit in DNA replication. Annu Rev Biochem 2002; 71:191–219.
Petruska J, Goodman MF, Boosalis MS, et al. Comparison between DNA melting thermodynamics and DNA polymerase fidelity. Proc Natl Acad Sci USA 1988; 85:6252–6256.
Echols H, Goodman MF . Fidelity mechanisms in DNA replication. Annu Rev Biochem 1991; 60:477–511.
Kunkel TA, Bebenek K . DNA replication fidelity. Annu Rev Biochem 2000; 69:497–529.
Goodman MF, Creighton S, Bloom LB, Petruska J . Biochemical basis of DNA replication fidelity. Crit Rev Biochem Mol Biol 1993; 28:83–126.
Washington MT, Helquist SA, Kool ET, Prakash L, Prakash S . Requirement of Watson-Crick hydrogen bonding for DNA synthesis by yeast DNA polymerase η. Mol Cell Biol 2003; 23:5107–5112.
Goodman MF . Hydrogen bonding revisited: geometric selection as a principal determinant of DNA replication fidelity. Proc Natl Acad Sci USA 1997; 94:10493–10495.
Beard WA, Wilson SH . Structural insights into the origins of DNA polymerase fidelity. Structure 2003; 11:489–496.
Johnson SJ, Beese LS . Structures of mismatch replication errors observed in a DNA polymerase. Cell 2004; 116:803–816.
Rothwell PJ, Waksman G . Structure and mechanism of DNA polymerases. In: John MS, David ADP, eds. Fibrous proteins: Muscle and molecular motors. Advances in Protein Chemistry 2005; 71:401–440.
Tsai YC, Johnson KA . A new paradigm for DNA polymerase specificity. Biochemistry 2006; 45:9675–9687.
Rothwell PJ, Waksman G . A pre-equilibrium before nucleotide binding limits fingers subdomain closure by Klentaq1. J Biol Chem 2007; 282:28884–28892.
Streisinger G, Okada Y, Emrich J, et al. Frameshift mutations and the genetic code. Cold Spring Harb Symp Quant Biol 1966; 31:77–84.
Kunkel TA, Soni A . Mutagenesis by transient misalignment. J Biol Chem 1988; 263:14784–14789.
Fujii S, Akiyama M, Aoki K, et al. DNA replication errors produced by the replicative apparatus of Escherichia coli. J Mol Biol 1999; 289:835–850.
Kunkel TA . Frameshift mutagenesis by eucaryotic DNA polymerases in vitro. J Biol Chem 1986; 261:13581–13587.
Efrati E, Tocco G, Eritja R, Wilson SH, Goodman MF . Abasic translesion synthesis by DNA polymerase β violates the “A-rule”. Novel types of nucleotide incorporation by human DNA polymerase β at an abasic lesion in different sequence contexts. J Biol Chem 1997; 272:2559–2569.
Garcia-Diaz M, Kunkel TA . Mechanism of a genetic glissando: structural biology of indel mutations. Trends Biochem Sci 2006; 31:206–214.
Bebenek K, Garcia-Diaz M, Blanco L, Kunkel TA . The frameshift infidelity of human DNA polymerase λ. Implications for function. J Biol Chem 2003; 278:34685–34690.
Foury F, Vanderstraeten S . Yeast mitochondrial DNA mutators with deficient proofreading exonucleolytic activity. EMBO J 1992; 11:2717–2726.
Morrison A, Johnson AL, Johnston LH, Sugino A . Pathway correcting DNA replication errors in Saccharomyces cerevisiae. EMBO J 1993; 12:1467–1473.
Morrison A, Sugino A . The 3′→5′ exonucleases of both DNA polymerases δ and ε participate in correcting errors of DNA replication in Saccharomyces cerevisiae. Mol Gen Genet 1994; 242:289–296.
Pavlov YI, Maki S, Maki H, Kunkel TA . Evidence for interplay among yeast replicative DNA polymerases α, δ and ε from studies of exonuclease and polymerase active site mutations. BMC Biol 2004; 2:11.
Vanderstraeten S, Van den Brule S, Hu J, Foury F . The role of 3′-5′ exonucleolytic proofreading and mismatch repair in yeast mitochondrial DNA error avoidance. J Biol Chem 1998; 273:23690–23697.
Goldsby RE, Lawrence NA, Hays LE, et al. Defective DNA polymerase-δ proofreading causes cancer susceptibility in mice. Nat Med 2001; 7:638–639.
Goldsby RE, Hays LE, Chen X, et al. High incidence of epithelial cancers in mice deficient for DNA polymerase δ proofreading. Proc Natl Acad Sci USA 2002; 99:15560–15565.
Zhang D, Mott JL, Chang SW, et al. Construction of transgenic mice with tissue-specific acceleration of mitochondrial DNA mutagenesis. Genomics 2000; 69:151–161.
Trifunovic A, Wredenberg A, Falkenberg M, et al. Premature ageing in mice expressing defective mitochondrial DNA polymerase. Nature 2004; 429:417–423.
Kroutil LC, Register K, Bebenek K, Kunkel TA . Exonucleolytic proofreading during replication of repetitive DNA. Biochemistry 1996; 35:1046–1053.
Tran HT, Gordenin DA, Resnick MA . The 3′→5′ Exonucleases of DNA polymerases δ and ε and the 3′→5′ exonuclease Exo1 have major roles in postreplication mutation avoidance in Saccharomyces cerevisiae. Mol Cell Biol 1999; 19:2000–2007.
Pavlov YI, Frahm C, Nick McElhinny SA, et al. Evidence that errors made by DNA polymerase α are corrected by DNA polymerase δ. Curr Biol 2006; 16:202–207.
McElhinny SAN, Pavlov YI, Kunkel TA . Evidence for extrinsic exonucleolytic proofreading. Cell Cycle 2006; 5:958–962.
Bebenek A, Carver GT, Dressman HK, et al. Dissecting the fidelity of bacteriophage RB69 DNA polymerase: site-specific modulation of fidelity by polymerase accessory proteins. Genetics 2002; 162:1003–1018.
Fortune JM, Stith CM, Kissling GE, Burgers PM, Kunkel TA . RPA and PCNA suppress formation of large deletion errors by yeast DNA polymerase δ. Nucleic Acids Res 2006; 34:4335–4341.
Tissier A, McDonald JP, Frank EG, Woodgate R . pol ι, a remarkably error-prone human DNA polymerase. Genes Dev 2000; 14:1642–1650.
Bebenek K, Tissier A, Frank EG, et al. 5′-Deoxyribose phosphate lyase activity of human DNA polymerase ι in vitro. Science 2001; 291:2156–2159.
Ogi T, Kato T Jr, Kato T, Ohmori H . Mutation enhancement by DINB1, a mammalian homologue of the Escherichia coli mutagenesis protein dinB. Genes Cells 1999; 4:607–618.
Fuchs RP, Fujii S, Wagner J . Properties and functions of Escherichia coli: Pol IV and Pol V. Adv Protein Chem 2004; 69:229–264.
Jarosz DF, Beuning PJ, Cohen SE, Walker GC . Y-family DNA polymerases in Escherichia coli. Trends Microbiol 2007; 15:70–77.
Haracska L, Prakash S, Prakash L . Yeast DNA polymerase ζ is an efficient extender of primer ends opposite from 7,8-dihydro-8-oxoguanine and O6-methylguanine. Mol Cell Biol 2003; 23:1453–1459.
Lawrence CW, Hinkle DC . DNA polymerase ζ and the control of DNA damage induced mutagenesis in eukaryotes. Cancer Surv 1996; 28:21–31.
Haracska L, Prakash L, Prakash S . Role of human DNA polymerase κ as an extender in translesion synthesis. Proc Natl Acad Sci USA 2002; 99:16000–16005.
Zhong X, Garg P, Stith CM, et al. The fidelity of DNA synthesis by yeast DNA polymerase ζ alone and with accessory proteins. Nucleic Acids Res 2006; 34:4731–4742.
Minesinger BK, Abdulovic AL, Ou TM, Jinks-Robertson S . The effect of oxidative metabolism on spontaneous Pol zeta-dependent translesion synthesis in Saccharomyces cerevisiae. DNA Repair (Amst) 2006; 5:226–234.
Harfe BD, Jinks-Robertson S . DNA polymerase ζ introduces multiple mutations when bypassing spontaneous DNA damage in Saccharomyces cerevisiae. Mol Cell 2000; 6:1491–1499.
Arana ME, Takata K, Garcia-Diaz M, Wood RD, Kunkel TA . A unique error signature for human DNA polymerase ν. DNA Repair (Amst) 2007; 6:213–223.
Takata K, Shimizu T, Iwai S, Wood RD . Human DNA polymerase N (POLN) is a low fidelity enzyme capable of error-free bypass of 5S-thymine glycol. J Biol Chem 2006; 281:23445–23455.
Seki M, Marini F, Wood RD . POLQ (Pol θ), a DNA polymerase and DNA-dependent ATPase in human cells. Nucl Acids Res 2003; 31:6117–6126.
Seki M, Wood RD . DNA polymerase θ (POLQ) can extend from mismatches and from bases opposite a (6-4) photoproduct. DNA Repair (Amst) 2008; 7:119–127.
Goodman MF . Error-prone repair DNA polymerases in prokaryotes and eukaryotes. Annu Rev Biochem 2002; 71:17–50.
Shcherbakova PV, Bebenek K, Kunkel TA . Functions of eukaryotic DNA polymerases. Sci Aging Knowledge Environ 2003; 2003:RE3.
Prakash S, Johnson RE, Prakash L . Eukaryotic translesion synthesis DNA polymerases: specificity of structure and function. Annu Rev Biochem 2005; 74:317–353.
Strauss BS . The “A” rule revisited: polymerases as determinants of mutational specificity. DNA Repair (Amst) 2002; 1:125–135.
Kokoska RJ, McCulloch SD, Kunkel TA . The efficiency and specificity of apurinic/apyrimidinic site bypass by human DNA polymerase η and Sulfolobus solfataricus Dpo4. J Biol Chem 2003; 278:50537–50545.
Ling H, Boudsocq F, Woodgate R, Yang W . Snapshots of replication through an abasic lesion; structural basis for base substitutions and frameshifts. Mol Cell 2004; 13:751–762.
Hogg M, Wallace SS, Doublie S . Crystallographic snapshots of a replicative DNA polymerase encountering an abasic site. EMBO J 2004; 23:1483–1493.
Washington MT, Johnson RE, Prakash L, Prakash S . Accuracy of lesion bypass by yeast and human DNA polymerase η. Proc Natl Acad Sci USA 2001; 98:8355–8360.
McCulloch SD, Kokoska RJ, Masutani C, et al. Preferential cis-syn thymine dimer bypass by DNA polymerase η occurs with biased fidelity. Nature 2004; 428:97–100.
McCulloch SD, Wood A, Garg P, Burgers PM, Kunkel TA . Effects of accessory proteins on the bypass of a cis-syn thymine-thymine dimer by Saccharomyces cerevisiae DNA polymerase η. Biochemistry 2007; 46:8888–8896.
Ling H, Boudsocq F, Plosky BS, Woodgate R, Yang W . Replication of a cis-syn thymine dimer at atomic resolution. Nature 2003; 424:1083–1087.
Lipscomb LA, Peek ME, Morningstar ML, et al. X-ray structure of a DNA decamer containing 7,8-dihydro-8-oxoguanine. Proc Natl Acad Sci USA 1995; 92:719–723.
McAuley-Hecht KE, Leonard GA, Gibson NJ, et al. Crystal structure of a DNA duplex containing 8-hydroxydeoxyguanine-adenine base pairs. Biochemistry 1994; 33:10266–10270.
Einolf HJ, Guengerich FP . Fidelity of nucleotide insertion at 8-oxo-7,8-dihydroguanine by mammalian DNA polymerase δ. Steady-state and pre-steady-state kinetic analysis. J Biol Chem 2001; 276:3764–3771.
Brieba LG, Eichman BF, Kokoska RJ, et al. Structural basis for the dual coding potential of 8-oxoguanosine by a high-fidelity DNA polymerase. EMBO J 2004; 23:3452–3461.
Grollman AP, Moriya M . Mutagenesis by 8-oxoguanine: an enemy within. Trends Genet 1993; 9:246–249.
Boiteux S, Gellon L, Guibourt N . Repair of 8-oxoguanine in Saccharomyces cerevisiae: interplay of DNA repair and replication mechanisms. Free Radical Biol Med 2002; 32:1244–1253.
Masutani C, Kusumoto R, Yamada A, et al. The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature 1999; 399:700–704.
Lin Q, Clark AB, McCulloch SD, et al. Increased susceptibility to UV-induced skin carcinogenesis in polymerase η-deficient mice. Cancer Res 2006; 66:87–94.
Dumstorf CA, Clark AB, Lin Q, et al. Participation of mouse DNA polymerase ι in strand-biased mutagenic bypass of UV photoproducts and suppression of skin cancer. Proc Natl Acad Sci USA 2006; 103:18083–18088.
de Feraudy S, Limoli CL, Giedzinski E, et al. Pol η is required for DNA replication during nucleotide deprivation by hydroxyurea. Oncogene 2007; 26:5713–5721.
Wang Y, Woodgate R, McManus TP, et al. Evidence that in Xeroderma Pigmentosum variant cells, which lack DNA polymerase η, DNA polymerase ι causes the very high frequency and unique spectrum of UV-induced mutations. Cancer Res 2007; 67:3018–3026.
Wang M, Devereux TR, Vikis HG, et al. Pol ι is a candidate for the mouse pulmonary adenoma resistance 2 locus, a major modifier of chemically induced lung neoplasia. Cancer Res 2004; 64:1924–1931.
Bemark M, Khamlichi AA, Davies SL, Neuberger MS . Disruption of mouse polymerase zeta (Rev3) leads to embryonic lethality and impairs blastocyst development in vitro. Curr Biol 2000; 10:1213–1216.
Esposito G, Godindagger I, Klein U, et al. Disruption of the Rev3l-encoded catalytic subunit of polymerase zeta in mice results in early embryonic lethality. Curr Biol 2000; 10:1221–1224.
Wittschieben J, Shivji MK, Lalani E, et al. Disruption of the developmentally regulated Rev3l gene causes embryonic lethality. Curr Biol 2000; 10:1217–1220.
Wittschieben JP, Reshmi SC, Gollin SM, Wood RD . Loss of DNA polymerase ζ causes chromosomal instability in mammalian cells. Cancer Res 2006; 66:134–142.
Haracska L, Acharya N, Unk I, et al. A single domain in human DNA polymerase ι mediates interaction with PCNA: implications for translesion DNA synthesis. Mol Cell Biol 2005; 25:1183–1190.
Haracska L, Johnson RE, Unk I, et al. Physical and functional interactions of human DNA polymerase η with PCNA. Mol Cell Biol 2001; 21:7199–7206.
Haracska L, Unk I, Johnson RE, et al. Stimulation of DNA synthesis activity of human DNA polymerase κ by PCNA. Mol Cell Biol 2002; 22:784–791.
Kannouche PL, Lehmann AR . Ubiquitination of PCNA and the polymerase switch in human cells. Cell Cycle 2004; 3:1011–1013.
Masuda Y, Kamiya K . Role of single-stranded DNA in targeting REV1 to primer termini. J Biol Chem 2006; 281:24314–24321.
Loeb LA . Mutator phenotype may be required for multistage carcinogenesis. Cancer Res 1991; 51:3075–3079.
Zheng L, Dai H, Zhou M, et al. Fen1 mutations result in autoimmunity, chronic inflammation and cancers. Nat Med 2007; 13:812–819.
Bohr VA, Ottersen OP, Tonjum T . Genome instability and DNA repair in brain, ageing and neurological disease. Neuroscience 2007; 145:1183–1186.
DePamphilis ML . DNA replication and human disease. Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, 2006.
Doublie S, Tabor S, Long AM, Richardson CC, Ellenberger T . Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 1998; 391:251–258.
Pursell ZF, Isoz I, Lundstrom E-B, Johansson E, Kunkel TA . Yeast DNA polymerase ε participates in leading-strand DNA replication. Science 2007; 317:127–130.
Garg P, Stith CM, Sabouri N, Johansson E, Burgers PM . Idling by DNA polymerase δ maintains a ligatable nick during lagging-strand DNA replication. Genes Dev 2004; 18:2764–2773.
Jin YH, Obert R, Burgers PM, et al. The 3′→5′ exonuclease of DNA polymerase δ can substitute for the 5′ flap endonuclease Rad27/Fen1 in processing Okazaki fragments and preventing genome instability. Proc Natl Acad Sci USA 2001; 98:5122–5127.
Jin YH, Ayyagari R, Resnick MA, Gordenin DA, Burgers PM . Okazaki fragment maturation in yeast. II. Cooperation between the polymerase and 3′→5′-exonuclease activities of pol δ in the creation of a ligatable nick. J Biol Chem 2003; 278:1626–1633.
Ling H, Boudsocq F, Woodgate R, Yang W . Crystal structure of a Y-family DNA polymerase in action: a mechanism for error-prone and lesion-bypass replication. Cell 2001; 107:91–102.
Bebenek K, Kunkel TA . Analyzing fidelity of DNA polymerases. Methods Enzymol 1995; 262:217–232.
Longley MJ, Nguyen D, Kunkel TA, Copeland WC . The fidelity of human DNA polymerase γ with and without exonucleolytic proofreading and the p55 accessory subunit. J Biol Chem 2001; 276:38555–38562.
Kunkel TA, Hamatake RK, Motto-Fox J, Fitzgerald MP, Sugino A . Fidelity of DNA polymerase I and the DNA polymerase I-DNA primase complex from Saccharomyces cerevisiae. Mol Cell Biol 1989; 9:4447–4458.
Shcherbakova PV, Pavlov YI, Chilkova O, et al. Unique error signature of the four-subunit yeast DNA polymerase ε. J Biol Chem 2003; 278:43770–43780.
Fortune JM, Pavlov YI, Welch CM, et al. Saccharomyces cerevisiae DNA polymerase δ: high fidelity for base substitutions but lower fidelity for single- and multi-base deletions. J Biol Chem 2005; 280:29980–29987.
Matsuda T, Bebenek K, Masutani C, et al. Error rate and specificity of human and murine DNA polymerase η. J Mol Biol 2001; 312:335–346.
Ohashi E, Bebenek K, Matsuda T, et al. Fidelity and processivity of DNA synthesis by DNA polymerase κ, the product of the human DINB1 gene. J Biol Chem 2000; 275:39678–39684.
Acknowledgements
We thank Drs Stephanie Nick-McElhinny and Zachary Pursell of the NIEHS for their constructive comments on the manuscript. This work was supported in part by the Intramural Research Program of the NIH (USA), National Institute of Environmental Health Sciences (TAK) and by the College of Agriculture and Life Sciences at North Carolina State University (SDM), USA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
McCulloch, S., Kunkel, T. The fidelity of DNA synthesis by eukaryotic replicative and translesion synthesis polymerases. Cell Res 18, 148–161 (2008). https://doi.org/10.1038/cr.2008.4
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2008.4
Keywords
This article is cited by
-
Glycosylase-based base editors for efficient T-to-G and C-to-G editing in mammalian cells
Nature Biotechnology (2024)
-
DNA polymerase κ suppresses inflammation and inflammation-induced mutagenesis and carcinogenic potential in the colon of mice
Genes and Environment (2023)
-
Recommendations for the classification of germline variants in the exonuclease domain of POLE and POLD1
Genome Medicine (2023)
-
Leveraging the replication stress response to optimize cancer therapy
Nature Reviews Cancer (2023)
-
Revisiting mutagenesis at non-B DNA motifs in the human genome
Nature Structural & Molecular Biology (2023)