Abstract
A microRNA expression screen was performed analyzing 157 different microRNAs in laser-microdissected tissues from benign melanocytic nevi (n = 10) and primary malignant melanomas (n = 10), using quantitative real-time PCR. Differential expression was found for 72 microRNAs. Members of the let-7 family of microRNAs were significantly downregulated in primary melanomas as compared with benign nevi, suggestive for a possible role of these molecules as tumor suppressors in malignant melanoma. Interestingly, similar findings had been described for lung and colon cancer. Overexpression of let-7b in melanoma cells in vitro downregulated the expression of cyclins D1, D3, and A, and cyclin-dependent kinase (Cdk) 4, all of which had been described to play a role in melanoma development. The effect of let-7b on protein expression was due to targeting of 3′-untranslated regions (3′UTRs) of individual mRNAs, as exemplified by reporter gene analyses for cyclin D1. In line with its downmodulating effects on cell cycle regulators, let-7b inhibited cell cycle progression and anchorage-independent growth of melanoma cells. Taken together, these findings not only point to new regulatory mechanisms of early melanoma development, but also may open avenues for future targeted therapies of this tumor.
Similar content being viewed by others
Introduction
MicroRNAs are 19-23 nucleotide (nt) noncoding RNA molecules that regulate gene expression by targeting the 3′-untranslated region (3′UTR) of mRNAs with consequent inhibition of protein translation or degradation of target mRNA 1. Recent studies indicate that microRNAs may be involved in tumor development in humans, being either under-expressed in tumors with consequent upregulation of oncogenes or over-expressed with consequent downregulation of tumor suppressor genes 2. MicroRNA let-7, a founding member of the microRNA family, inversely correlated with the expression of the RAS oncogene in human lung cancer tissues 3. Furthermore, overexpression of let-7a in tumor cells in vitro downmodulated Ras expression due to binding of let-7a to the 3′UTR of RAS. This was shown by immunofluorescence analysis and reporter gene assays. These experiments were suggestive for an important role of let-7a as a tumor suppressor in lung cancer.
Up to now, a considerable number of microRNAs were found to be downregulated in human cancers compared with corresponding benign tissues. For example, miR-143 and miR-145 expression was significantly reduced in colorectal tumours, and miR-145 expression was reduced in breast cancer 4, 5. In a large-scale analysis of 540 tumor samples from lung, breast, stomach, prostate, colon, and pancreatic tumors, a so-called solid cancer microRNA signature was identified, which also included a series of overexpressed microRNAs such as miR-17-5p, miR-20a, miR-21, miR-92, miR-106a, and miR-155, some of which had already been implicated in cancer development in earlier studies 6.
Little is known about the possible role of microRNAs for melanoma development. One recent report showed that microRNA patterns were able to differentiate between different tumor types such as colon cancer, melanoma, hematologic, and central nervous system tumors, based on the analysis of cell lines derived from these tissues 7. These analyses were suggestive for a tumor-specific microRNA pattern. However, melanocytes or benign melanocytic nevi, which would allow a direct comparison of both stages of melanoma development, were not included in that study.
Results
Let-7 microRNA family members are downregulated in primary melanomas
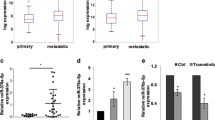
In the present study, the expression of 157 different microRNAs was analyzed in 10 benign melanocytic nevi and 10 primary malignant melanomas, respectively. Overall 72 differentially expressed microRNAs were identified (Table 1). Among these were a series of let-7 family members, as well as miR-15a, miR-17, miR-28, miR-106a, and miR-144, which had already been described to be deregulated in malignant tumors. We focused on the let-7 family, represented by let-7a, let-7b, let-7d, let-7e, and let-7g, all of which were downregulated in primary melanomas (Table 1). Let-7 family members might play a particular role in melanoma development, because the predicted targets of this family, according to public microRNA databases (http://pictar.bio.nyu.edu/; http://microrna.sanger.ac.uk), include a series of molecules involved in melanoma development and progression, such as N-Ras, Raf, c-Myc, cyclins D1 and D3, and cyclin-dependent kinase (Cdk) 4. Validation experiments performed for let-7a, let-7b, and let-7d using the 10 samples of benign nevi and primary melanomas, respectively, for individual measurements, confirmed the data from our screening analyses with pooled samples (Figure 1).
Validation of microRNA expression screening of benign melanocytic nevi and primary malignant melanomas. In validation experiments each sample (from 10 benign nevi and 10 primary melanomas) was tested individually for expression of microRNAs let-7a, let-7b, and let-7d, respectively, using commercially available TaqMan® MicroRNA Assays. Data are given as relative percent of average signals in benign nevi and primary melanomas. Asterisks indicate statistical significance of differences in microRNA expression between benign nevi and primary melanomas (P < 0.05; Mann-Whitney-U test).
Let-7b represses expression of cyclins D1, D3, Cdk4, and cyclin A
The predicted targets of let-7b comprise most of the mentioned melanoma-associated genes (Table 1), and thus it might be of particular importance for malignant melanoma development. In order to test the functional activity of let-7b on these targets, immunoblot and immunofluorescence analyses were performed after transient transfection of melanoma cells with let-7b precursor microRNA. As shown in Figure 2A, SK-Mel-147 melanoma cells showed downregulation of cyclins D1, D3, Cdk4, and cyclin A by let-7b after 24 h of let-7b transfection, and downregulation of cyclin A after 48 and 72 h, respectively, of let-7b transfection (left panel). G361 cells showed downregulation of Cdk4 and cyclin D3 after 24, 48, and 72 h, respectively, of let-7b transfection, and downregulation of cyclin A after 48 h of let-7b transfection (right panel). Downregulation of cyclin A in G361 cells was less pronounced than in SK-Mel-147 cells. These results were confirmed by parallelly performed immunofluorescence staining of SK-Mel-147 cells (Figure 2B-2D). Taken together, let-7b represses the expression of a series of melanoma-associated genes. Since cyclin A is not a predicted target of let-7b (http://pictar.bio.nyu.edu/; http://microrna.sanger.ac.uk), indirect mechanisms might contribute to cyclin A downregulation after let-7b overexpression.
Influence of microRNA let-7b on cyclins D1, D3, Cdk4, and cyclin A expression in melanoma cells. (A) SK-Mel-147 or G361 melanoma cells were transiently transfected with 30 nM of let-7b or control (co) precursor microRNA. Total protein extracts of cells were subjected to immunoblotting of cyclins D1, D3, Cdk4, and cyclin A. Immunoblots were re-probed with an anti-tubulin mouse monoclonal Ab to verify equal loading of proteins. (B-D) Immunofluorescence analyses of SK-Mel-147 melanoma cells transfected with 30 nM let-7b or control precursor microRNA. At 36 h after transfection melanoma cells were fixed with paraformaldehyde and stained with anti-cyclin D1 (B), anti-Cdk4 (C), or anti-cyclin D3 (D) antibodies. An Alexa Fluor 488 goat anti-mouse IgG antibody or goat anti-rabbit IgG antibody was used as secondary antibody. Cells were counterstained with DAPI. Upper panel shows microscopic pictures of immunofluorescence staining of one representative experiment (× 100 and × 400 magnifications, respectively). Lower panel shows graphic presentation of mean fluorescence intensities of three independent experiments. Asterisks in (B-D) indicate statistical significance of differences in cyclin D1, Cdk4, or cyclin D3 expression in let-7b-transfected cells compared with control precursor microRNA-transfected cells (P < 0.05; Mann-Whitney-U test).
Let-7b targets 3′UTR of CCND1
To determine whether the negative regulatory effects of let-7b on cyclin expression were indeed mediated through binding to 3′UTR of target mRNAs, in vitro luciferase assays were performed. For this purpose, corresponding 3′UTRs of cyclin D1 (CCND1) and cyclin A (CCNA) were cloned into the pGL3-Promoter Vector, immediately downstream of the luciferase gene. SK-Mel-147 melanoma cells, which express endogenous let-7b, were transfected with these reporter gene constructs.
As shown in Figure 3, luciferase activity in cells transfected with pGL3-CCND1-3′UTR was significantly reduced as compared with cells transfected with the control pGL3 vector. Furthermore, inhibition of let-7b activity by co-transfection of a let-7b inhibitor (antagomir) relieved the repression of the reporter gene. In experiments using a pGL3-CCNA-3′UTR construct, significant downmodulation of luciferase activity was also observed. However, in this case, repression of the reporter gene was not relieved after co-transfection of the let-7b inhibitor. These latter findings argue for the presence of cyclin A targeting microRNAs in SK-Mel-147 melanoma cells, and also indicate that cyclin A is not a direct target of let-7b.
Reporter gene analyses using CCND1 and CCNA 3′UTR constructs. SK-Mel-147 melanoma cells were transfected with the pGL3 Promotor Vector (pGL3), constructs containing the 3′UTR of CCND1 (pGL3-CCND1-3′UTR), or 3′UTR of CCNA (pGL3-CCNA-3′UTR), and co-transfected with either anti-let-7b antagomir or left untransfected (Ø). Single asterisks indicate statistical significance of differences in pGL3-CCND1-3′UTR or pGL3-CCNA-3′UTR luciferase activity compared with pGL3 luciferase activity; double asterisks indicate statistical significance of difference in pGL3-CCND1-3′UTR luciferase activity of anti-let-7b-transfected compared with untransfected cells (P < 0.05; Mann-Whitney-U test).
Let-7b reduces cell cycle progression and anchorage-independent growth
Based on the findings of let-7b-mediated downregulation of different cyclins and Cdk4, we next examined the effects of let-7b on cell cycle regulation and cell proliferation of melanoma cells. For this purpose, Sk-Mel-147 and G361 melanoma cells were transfected with let-7b precursor or control microRNA, stained with BrdU and analyzed by flow cytometry. As shown in Figure 4, in SK-Mel-147 cells let-7b overexpression induced a significant reduction of the number of cells in S-phase, paralleled by an increase of the number of cells in G0-G1 phase. In contrast, after transfection of let-7b antagomir, the number of cells in S-phase increased together with a slight decrease of the number of cells in G2-M phase. Similar results were obtained for G361 cells (data not shown). Finally, soft agar assays were performed to analyze the influence of let-7b on anchorage-independent growth (colony formation) of melanoma cells. As shown in Figure 5A/B, overexpression of let-7b in G361 melanoma cells resulted in a significant reduction of colony numbers after 20 days of culture. Taken together, let-7b might be an important regulator of melanoma cell growth via inhibition of cell cycle progression.
Influence of let-7b on cell cycle progression of melanoma cells. SK-Mel-147 melanoma cells were transfected for 48 h with 30 nM let-7b precursor microRNA (Pre-let-7b) or anti-let-7b antagomir. Cell cycle analysis was performed by flow cytometry. Data are given as mean values +/− SD of three independent experiments. Asterisks indicate statistical significance of differences in the number of cells in G0-G1, S, or G2-M between Pre-let-7b- or anti-let7b-transfected versus control transfected cells (P < 0.05; Mann-Whitney-U test).
Influence of let-7b on colony formation of melanoma cell in vitro. G361 melanoma cells were transfected for 48 h with 30 nM let-7b or control precursor microRNA and grown in soft agar for 20 days at 37 °C. (A) High-power views (× 100 magnification) of colonies from one representative experiment. (B) Graphic presentation of numbers of colony-forming units from three independent experiments. Data in (B) are given as mean values +/− SD. Asterisk indicates statistical significance of the difference in number of colonies of let-7b-transfected compared with control miRNA-transfected cells (P < 0.05; Mann-Whitney-U test).
Discussion
In the present report, a series of 72 microRNAs were found to be differentially expressed in benign melanocytic nevi and primary malignant melanomas. Interestingly, the majority of differentially expressed microRNAs were downregulated in melanoma tissues. Similar findings regarding a predominance of downregulated microRNAs in malignant tumors compared with normal tissues were reported recently 8. The authors stated that these findings are consistent with the hypothesis that miRNAs function to prevent cell division and induce terminal differentiation. Thus, downregulation of microRNAs in malignant tumors might support uncontrolled cell proliferation and de-differentiation.
In our study, members of the let-7 family were all downregulated in melanomas compared with benign nevi. In line with this observation, copy number losses were demonstrated for both let-7a and let-7b genes in a recent study on a series of 40 melanoma cell lines tested for alterations in genomic microRNA loci 9. In the mentioned study, microRNA copy number losses correlated with reduced microRNA expression levels. Since the predicted targets of let-7 family microRNAs include important molecules for melanoma development, such as N-Ras, Raf, c-Myc, cyclins D1 and D3, and Cdk4 10, our current study further analyzed the let-7 family microRNAs in more detail.
Although microRNA patterns in tumors have been a focus of recent research, target validation has only been performed for a limited number of genes. Here we focused on let-7b and its predicted targets, because the latter comprise most of the mentioned melanoma-associated genes, suggesting that let-7b might play a central role in melanoma development. Immunoblot and immunofluorescence analyses showed that overexpression of let-7b in melanoma cells resulted in significant downregulation of cyclins D1, D3, and Cdk4 protein expression. Downregulation of cyclin A protein expression was less prominent. Moreover, the negative regulatory effect of let-7b on cyclin D1 expression was indeed mediated through binding to 3′UTR of the target, as shown by in vitro luciferase assays. Downmodulation of these target molecules is likely due to inhibition of translation, since a recent study performing large-scale mRNA expression profiling of let-7b-transfected HepG2 liver cancer cells failed to show that the above-mentioned cyclins and Cdk4 are directly affected by let-7b-mediated mRNA degradation 11.
The findings of let-7b downregulation in primary melanomas combined with its influence on cell cycle molecules might be of relevance for the in vivo situation, since cyclins D1 and D3 have been described to be upregulated in primary melanomas compared with benign melanocytic nevi, in a series of immunohistochemical studies 12, 13, 14. The role of cyclin D1 for melanoma development was also emphasized in a study using comparative genomic hybridization 15. In that study, a considerable number of melanomas showed enhanced cyclin D1 expression independent of gene copy number changes, suggestive of a further level of regulation. Moreover, the functional relevance of cyclin D1 for melanoma growth was demonstrated in xenotransplant experiments in nude mice using CCND1 antisense RNA 15.
As shown earlier, a mutation in the CDK4 gene, leading to amino-acid substitution at residue 24 (R24C), inhibited the binding of tumor suppressor p16INK4A in melanoma cells 16. This mechanism had been proposed to play a role in melanoma development, since this CDK4 mutation had been detected in three melanoma families. Moreover, mice carrying this Cdk4 activating mutation developed malignant melanomas after skin treatment with different carcinogenic substances 17. In line with this, we observed downregulation of Cdk4 protein in let-7b-transfected melanoma cells, which paralleled the findings for cyclins D1 and D3. Taken together, cell cycle regulation via cyclin D1 and Cdk4 appears to play an important role in malignant melanoma and may be disturbed on different levels. In one of the mentioned immunohistochemical studies, enhanced cyclin D3 staining correlated with decreased overall survival of patients with superficial spreading melanomas 12. Furthermore, a concordant staining pattern was observed for cyclins D3 and D1, and for cyclin D3 and proliferation marker Ki-67 in superficial spreading melanomas, suggestive of a particular role of both cyclin D1 and D3 in tumor cell proliferation in this melanoma subtype.
Downregulation of cyclin A was observed in both cell lines 48 h after let-7b transfection. However, as mentioned above, it was less prominent, in particular in G361 cells. This gene (CCNA) has not yet been described as a target for let-7b. However, HMGA2, which is a known target of let-7b 18, is a major activator of cyclin A expression 19. Cyclin A downregulation might thus occur indirectly via let-7b/HMGA2 interaction. In line with this, in experiments using a CCNA 3′UTR reporter construct, a significant downmodulation of luciferase activity was observed. However, repression of the reporter gene could not be relieved after co-transfection of a let-7b inhibitor. These latter findings argue for the presence of cyclin A targeting microRNAs in melanoma cells, and also suggest that the effect of let-7b overexpression on cyclin A levels is indirect.
Based on three immunohistochemical studies showing significant upregulation of cyclin A in melanomas compared with benign nevi 12, 13, 14, cyclin A appears to play a role in melanoma development. Interestingly, cyclin A was described as an independent prognostic factor in malignant melanoma, since its expression correlated with disease-free survival of melanoma patients, at least for patients with superficial spreading malignant melanomas 13. More recently, it was demonstrated that treatment with deacetylase inhibitor trichostatin A induced apoptosis in melanoma cells in vitro, which was associated with reduced cyclin A expression 20. These findings are suggestive of a protective role of cyclin A against apoptosis in melanoma.
To examine the effects of let-7b on melanoma cell growth in vitro, cell cycle analyses and colony-forming assays were performed. In cell cycle experiments, overexpression of let-7b in melanoma cells resulted in a significant reduction of the number of proliferating cells (cells in S-phase), accompanied by an increased number of cells in G1. These findings are in line with the observed repression of cyclin D1 and D3 expression by let-7b, both of which play a central role in G1-phase of the cell cycle 21. Similar findings regarding cell cycle inhibition by let-7b and other let-7 family members were reported very recently for HepG2 liver cancer cells 11. Finally, let-7b led to a significant reduction of melanoma cell growth and proliferation in soft agar assays. Taken together, these findings support the notion that let-7b might be an important negative regulator of melanoma cell growth and proliferation via inhibition of cell cycle progression.
In conclusion, a microRNA pattern was identified associated with primary melanoma development. Here let-7 family members, and in particular let-7b, might play an important role. Our results might help explain many of the findings in malignant melanoma such as overexpressed cell cycle molecules and high proliferative activity, and might open avenues for innovative treatment approaches.
Materials and Methods
Patient tissues
Biopsy material was obtained from 10 benign melanocytic nevi and 10 primary malignant melanomas, respectively (Supplementary information, Table S1). Tumor thickness of primary melanomas was measured according to Breslow 22 and ranged from 0.40 to 8.00 mm. Melanocytes or melanoma cells were laser-microdissected from tissues as described 23. The presented study was conducted according to the Principles of the Declaration of Helsinki and was approved by the local Ethics Committee of the University of Rostock. Informed consent was obtained from all patients included in the study.
MicroRNA extraction and quantification by real-time PCR
MicroRNA was extracted using the mirVanaTM RNA Isolation Kit (Applied Biosystems, Darmstadt, Germany). RNA concentrations were determined with a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). RNAs from 10 samples in each group (benign nevi or primary melanomas) were pooled for expression profiling of 157 different mature microRNAs, using a commercially available quantitative real-time PCR microRNA Assay (Human Panel-Early Access Kit; Applied Biosystems, Darmstadt, Germany), according to the manufacturer's specifications. PCR amplification and signal detection of amplified microRNAs was performed using an ABI PRISMTM 7700 system (Applied Biosystems). Ct (threshold cycle) values of real-time PCR were normalized to U43 small nucleolar RNA (RNU43, #4373375; Applied Biosystems), which was measured in parallel. Fold changes in microRNA copy numbers between benign nevi and primary melanomas are given as log10 of 2−ΔCt values. In validation experiments each sample (from 10 benign nevi or 10 primary melanomas) was tested individually for expression of microRNAs let-7a, let-7b, and let-7d, respectively, using commercially available TaqMan® MicroRNA Assays (#4373169, #4373168, and #4373166; Applied Biosystems). Ct values of real-time PCR analyses were normalized to RNU43 expression, which was measured in parallel. Data are given as relative percent of average signals in benign nevi and primary melanomas.
Immunoblotting and immunofluorescence analyses
SK-Mel-147 or G361 melanoma cells were transiently transfected with 30 nM of let-7b (PM11050, Ambion, Austin, Texas, USA) or control precursor microRNA #1 (AM17110, Ambion), using Lipofectamine 2000 reagent (Invitrogen, Karlsruhe, Germany). Cells were harvested in radioimmunoprecipitation buffer [50 mmol/l Tris-HCl (pH 7.4), 150 mmol/l NaCl, 1% NP-40, 0.5% deoxycholic acid, 0.1% SDS], supplemented with protease inhibitor mix (Roche Molecular Biochemicals, Mannheim, Germany). In all, 40 μg of total protein extract were denatured in electrophoresis sample buffer for 5 min at 95 °C, and subjected to SDS-polyacrylamide gel electrophoresis (PAGE). Gels were electroblotted onto nitrocellulose membranes (Highbond ECLTM, Amersham, Braunschweig, Germany). Immunodetection was performed, using the following primary antibodies (all purchased from Santa Cruz Biotechnologies, Santa Cruz, CA, USA): anti-cyclin D1 mouse monoclonal Ab (sc-8396), anti-cyclin D3 rabbit polyclonal Ab (sc-182), anti-cyclin A mouse monoclonal Ab (sc-239), and anti-Cdk4 rabbit polyclonal Ab (sc-601). Goat anti-mouse or donkey anti-rabbit HRP-conjugated antibodies were used as secondary antibodies (W4021 or V7951, Promega, Mannheim, Germany). Immunoblots were re-probed with an anti-tubulin mouse monoclonal Ab (sc-5274, Santa Cruz Biotechnologies) to verify equal loading of proteins. For immunofluorescence analyses of SK-Mel-147, cells were plated on glass coverslips in six-well plates and transfected with 30 nM let-7b or control precursor microRNA. At 36 h after transfection, coverslips were stained with the mentioned anti-cyclin D1, anti-Cdk4, or anti-cyclin D3 antibodies. Alexa Fluor 488 goat anti-mouse IgG antibody or goat anti-rabbit IgG antibody was used as secondary antibody (A11001 or A11008, Invitrogen). Coverslips were counterstained with DAPI (Invitrogen-Molecular Probes, Eugene, Oregon, USA) for visualization of nuclei. Microscopic analysis was performed with a confocal laser-scanning microscope (Leica Microsystems, Bensheim, Germany). Fluorescence intensities were measured in a few viewing areas for 200-300 cells per coverslip and analyzed using ImageJ 1.37v software (http://rsb.info.nih.gov/ij/index.html).
Luciferase reporter assay
Luciferase reporter gene constructs carrying the 3′UTR of CCND1 or CCNA gene were generated. For this purpose, human genomic DNA was PCR amplified using the following primers: 3′UTR of CCND1 (gene bank accession number NM 053056):
5′-GCT CTA GAG CTC CTC TCC GGA GCA TT-3′,
5′-GCT CTA GAG CTT CTG CTG GAA ACA TGCC-3′;
3′UTR of CCNA (gene bank accession number NM 001237):
5′-GCT CTA GAG CTC TGT AAC AAT GAA AG-3′,
5′-GCT CTA GAG CGT TTT CCA AAT CAA T-3′.
Primers were purchased from Sigma-Genosys (Steinheim, Germany). Primers generated XbaI restriction sites, which were used for cloning of PCR products into the XbaI site of pGL3-Promotor Vector (Promega, Mannheim, Germany), located immediately downstream of the luciferase gene. SK-Mel-147 cells were transfected with the pGL3-Promotor Vector (pGL3), constructs containing the 3′UTR of CCND1 (pGL3-CCND1-3′UTR) or 3′UTR of CCNA (pGL3-CCNA-3′UTR), and co-transfected with either anti-let-7b antagomir or left untransfected (Ø).
Cell cycle analysis and colony formation
For cell cycle analysis, SK-Mel-147 and G361 cells were stained with BrdU for 6 h, using a FITC BrdU Flow Kit (BD Biosciences, Heidelberg, Germany). Populations in G0-G1, S, and G2-M phase were measured by flow cytometry with a FACScalibur (BD Biosciences). Data were processed by CellQuestTM Software (BD Biosciences). For analysis of anchorage-independent cell growth in vitro, soft agar assays were performed. For this purpose, G361 melanoma cells were transfected for 48 h with 30 nM let-7b or control precursor microRNA and grown in soft agar for 20 days at 37 °C. Influence of let-7b on colony formation of melanoma cells was estimated by counting the colony forming units by microscopic inspection.
Statistical analysis
Data are presented as mean +/− SD. Statistical significance of differences in microRNA expression between benign nevi and primary melanomas, differences in promoter activity, and differences in cell cycle populations and the number of colony-forming units in soft agar assays were analyzed by Mann-Whitney-U test (P < 0.05 was regarded as significant).
(Supplementary information is available at Cell Research's website.)
References
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116:281–297.
Esquela-Kerscher A, Slack FJ . Oncomirs – microRNAs with a role in cancer. Nat Rev Cancer 2006; 6:259–269.
Johnson SM, Grosshans H, Shingara J, et al. RAS is regulated by the let-7 microRNA family. Cell 2005; 120:635–647.
Michael MZ, O'Connor SM, van Holst Pellekaan NG, Young GP, James RJ . Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 2003; 1:882–891.
Iorio MV, Ferracin M, Liu CG, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005; 65:7065–7070.
Volinia S, Calin GA, Liu CG, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 2006; 103:2257–2261.
Gaur A, Jewell DA, Liang Y, et al. Characterization of microRNA expression levels and their biological correlates in human cancer cell lines. Cancer Res 2007; 67:2456–2468.
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435:834–838.
Zhang L, Huang J, Yang N, et al. microRNAs exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci USA 2006; 103:9136–9141.
Miller AJ, Mihm MC Jr. Melanoma. N Engl J Med 2006; 355:51–65.
Johnson CD, Esquela-Kerscher A, Stefani G, et al. The let-7 MicroRNA represses cell proliferation pathways in human cells. Cancer Res 2007; 67:7713–7722.
Florenes VA, Faye RS, Maelandsmo GM, Nesland JM, Holm R . Levels of cyclin D1 and D3 in malignant melanoma: deregulated cyclin D3 expression is associated with poor clinical outcome in superficial melanoma. Clin Cancer Res 2000; 6:3614–3620.
Florenes VA, Maelandsmo GM, Faye R, Nesland JM, Holm R . Cyclin A expression in superficial spreading malignant melanomas correlates with clinical outcome. J Pathol 2001; 195:530–536.
Alonso SR, Ortiz P, Pollan M, et al. Progression in cutaneous malignant melanoma is associated with distinct expression profiles: a tissue microarray-based study. Am J Pathol 2004; 164:193–203.
Sauter ER, Yeo UC, von Stemm A, et al. Cyclin D1 is a candidate oncogene in cutaneous melanoma. Cancer Res 2002; 62:3200–3206.
Wolfel T, Hauer M, Schneider J, et al. A p16INK4a-insensitive CDK4 mutant targeted by cytolytic T lymphocytes in a human melanoma. Science 1995; 269:1281–1284.
Sotillo R, Garcia JF, Ortega S, et al. Invasive melanoma in Cdk4-targeted mice. Proc Natl Acad Sci USA 2001; 98:13312–13317.
Lee YS, Dutta A . The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev 2007; 21:1025–1030.
Tessari MA, Gostissa M, Altamura S, et al. Transcriptional activation of the cyclin A gene by the architectural transcription factor HMGA2. Mol Cell Biol 2003; 23:9104–9116.
Florenes VA, Skrede M, Jorgensen K, Nesland JM . Deacetylase inhibition in malignant melanomas: impact on cell cycle regulation and survival. Melanoma Res 2004; 14:173–181.
Malumbres M, Barbacid M . Mammalian cyclin-dependent kinases. Trends Biochem Sci 2005; 30:630–641.
Breslow A . Thickness, cross-sectional areas and depth of invasion in the prognosis of cutaneous melanoma. Ann Surg 1970; 172:902–908.
Jaeger J, Koczan D, Thiesen HJ, et al. Gene expression signatures for tumor progression, tumor subtype, and tumor thickness in laser-microdissected melanoma tissues. Clin Cancer Res 2007; 13:806–815.
Acknowledgements
We thank R Waterstradt for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary information, Table S1
List of melanoma specimens analysed by TaqMan® real-time PCR for microRNA expression (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Schultz, J., Lorenz, P., Gross, G. et al. MicroRNA let-7b targets important cell cycle molecules in malignant melanoma cells and interferes with anchorage-independent growth. Cell Res 18, 549–557 (2008). https://doi.org/10.1038/cr.2008.45
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2008.45
Keywords
This article is cited by
-
Tumor suppressor let-7 acts as a key regulator for pluripotency gene expression in Muse cells
Cellular and Molecular Life Sciences (2024)
-
CADM1 impairs the effect of miR-1246 on promoting cell cycle progression in chemo-resistant leukemia cells
BMC Cancer (2023)
-
Current understanding of epigenetics role in melanoma treatment and resistance
Cancer Cell International (2022)
-
MicroRNA let-7b downregulates AML1-ETO oncogene expression in t(8;21) AML by targeting its 3′UTR
Experimental Hematology & Oncology (2021)
-
Signal pathways of melanoma and targeted therapy
Signal Transduction and Targeted Therapy (2021)