Abstract
Human neurocristopathies include a number of syndromes, tumors, and dysmorphologies of neural crest (NC) stem cell derivatives. In recent years, many white spotting genes have been associated with hypopigmentary disorders and deafness in neurocristopathies resulting from NC stem cell-derived melanocyte deficiency during development. These include PAX3, SOX10, MITF, SNAI2, EDNRB, EDN3, KIT, and KITL. Recent studies have revealed surprising new insights into a central role of MITF in the complex network of interacting genes in melanocyte development. In this perspective, we provide an overview of some of the current findings and explore complex functional roles of these genes during NC stem cell-derived melanocyte development.
Similar content being viewed by others
Neural crest stem cell development
The neural crest (NC) is a unique embryonic structure and contains a remarkable multipotent stem cell population that arises during vertebrate embryogenesis 1, 2. NC has been referred to as the fourth germ layer because of its great importance during development 3. NC stem cells arise from the dorsal neural tube during neurolation in early development, then migrate out from the neural tube and along defined pathways throughout the body, where they contribute to numerous cell types and tissues, including melanocytes, ocular and periocular structures, bone and cartilage cells of the cranial skeleton, odontoblasts, autonomic neurons, sensory neurons, enteric neurons, smooth muscle, endocrine cells, chromaffin cells, and glial cells 1. Although it has long been thought that the fates of NC-derived lineages are controlled by transcription and growth factors, the physiological functions of these factors are not fully known.
Understanding NC development is medically important because defective derivatives of aberrant NC cell development give rise to numerous human diseases known as neurocristopathies 4. These diseases include ocular diseases (such as iris hypoplasia and optic nerve head melanocytoma), cardiocutaneous syndromes, craniofacial malformations of mesoectodermal origin, DiGeorge syndrome, Ewing's tumors, Hirschsprung disease, lentigo, medullary carcinoma of the thyroid, melanotic nevi, melanoma,multiple endocrine neoplasia (types 2A and 2B), neuroblastoma, neurocutaneous syndromes, neurofibromatosis type 1, PCWH (Peripheral demyelinating neuropathy, Central dysmyelinating leukodystrophy, Waardenburg syndrome (WS), and Hirschsprung disease), PHACES syndrome (Posterior fossa abnormalities and other structural brain abnormalities, Hemangioma(s) of the cervical facial region, Arterial cerebrovascular anomalies, Cardiac defects, aortic coarctation and other aortic abnormalities, Eye Anomalies, Sternal defects, and/or Supraumbilical raphe), pheochromocytoma, piebaldism, WS, Tietz syndrome, and more 4. Among these WS is an autosomal-dominant subtype of complex NC diseases and is named after the Dutch ophthalmologist who, in 1947, first described a patient with heterochromia iridis (different eye colors), congenital deafness, and dystopia canthorum (lateral displacement of the inner canthi of the eyes leading to a wide nasal bridge). WS patients also show additional defects, including white forelock, pigmentary disturbance of the skin, upper limb abnormalities, and megacolon 5. To date there are at least four types of WS that are due to mutations in separate transcription factors, including SOX10, MITF, PAX3, and SNAI2, and in signaling molecules, including EDNRB and EDN3. The four WS types are categorized based on presentation of various subsets of the phenotypic characteristics of the syndrome. For example, WS type 1 patients have craniofacial defects, WS type 3 patients have craniofacial and limb defects, and WS type 4 patients have megacolon. Intriguingly, distinct subtypes of WS and piebaldism, which is associated with mutation of KIT, often have the common phenotype of hypopigmentation, which is due to melanocyte defects in the skin. The comparable hypopigmentation defect in these diseases reflects a possible functional relationship among the disease-associated genes in melanocyte development. In this review, we discuss the known functional roles of these genes during NC stem cell-derived melanocyte development and propose alternative models of functional roles of these genes, with a focus on the central role of MITF.
White spotting mouse disease models and melanocyte development
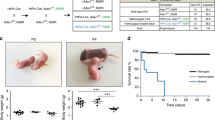
Gene expression programs that direct the development of distinct cell lineages from unspecified precursor cells are the result of complex interactions between cell-extrinsic signals and transcription factors. An excellent system to study such interactions is provided by the development of melanocytes. Their precursor cells, the melanoblasts, originate from multipotent NC stem cells and migrate along characteristic pathways to various destinations such as the iris and the choroid of the eye, the inner ear, the dermis, and the epidermis. In the skin, these precursors differentiate into melanin-producing cells that determine skin color and protect the organism from UV radiation, one of the risk factors for skin cancers such as melanoma 6. In addition, the precursors distribute into the bulged region of developing hair follicles, where they persist as self-renewing stem cells in the niche 7. For their development, melanoblasts depend on numerous transcription factors and signaling systems. These include the transcription factors PAX3 8, 9, SOX10 9, 10, 11, and MITF (Microphthalmia-associated Transcription Factor) 12, the WNT signaling pathway 13, 14, G protein-coupled endothelin receptor B (EDNRB) and its ligand, endothelin 3 (EDN3) 15, 16, and receptor tyrosine kinase KIT and KIT-ligand (KITL) 17, 18. Among the genes encoding these factors, Mitf, Sox10, Pax3, Kit, and Kitl comprise a particularly intriguing set, since heterozygosity for certain mutations in each of these genes leads to the strikingly similar phenotype of belly spotting in mice (Figure 1). Since many in vivo and in vitro observations suggest that there are mutual interactions between these genes 9, 19, a possible functional regulatory relationship may exist among these genes in melanocyte development.
Mutations in the genes encoding key transcriptional factors or signaling molecules result in characteristic white coat color phenotypes, reflecting the ability of these genes to regulate NC stem cell-derived melanocyte generation. Among these genes Mitf affects development of both melanocytes and RPE, whereas others affect only melanocyte development. (A) A Mitfmi-vga-9 homozygote (a null allele). Mitfmi-vga-9 homozygotes are white, with small eyes, whereas heterozygotes have normal pigmentation. (B) A Mitfmi-rw (microphthalmia-red eyed white) homozygote, which contains a genomic deletion starting downstream of exon 1E and ending upstream of exon 1M and encompassing the exons 1H, 1D, and 1B1a/1B1b and their flanking sequences 147. Mitfmi-rw homozygotes show abnormal RPE development and primarily white fur, but always display a black head spot, reflecting normal melanocyte generation in that region. (C) A Mitfmi-bws (microphthalmia-black and white spotting) homozygote. These mice harbor a point mutation that results in altered splicing so that the Mitf produces not only a wild-type transcript that contains exon 2b, but also a transcript that lacks exon 2b, which contains a KIT signaling-dependent phosphorylation site, serine-73. This causes deficiency of many skin melanocytes without affecting RPE development, resulting in widespread white spotting and black eyes. (D) A 10- month- old Mitfmi-vit (microphthalmia-vitiligo) homozygote. This allele contains a G to A transition that leads to an aspartate to asparagine substitution at amino acid 222 in the helix 1 region of MITF protein. Mitfmi-vit homozygous mice are born normally pigmented, but gradually lose their melanocyte stem cells with aging, resulting in a gray coat phenotype 146. (E) Ednrbtm1Myks homozygote (left) and heterozygote littermate (right). This homozygote contains a transgenic insertion of a LacZ reporter gene at the Ednrb locus, resulting in absence of Ednrb expression. Homozygotes are almost completely white, with pigmented regions remaining in the head and rump, and die from megacolon as juveniles 129. (F) Kittm1Alf (also known as KitW-lacZ) homozygote (top) and normal littermate (bottom), age P3. This mouse is homozygous for the transgenic insertion of a LacZ reporter gene at the Kit locus 148. Absence of normal Kit causes complete lack of skin melanocytes, resulting in white fur. (G) A Sox10Hry homozygote (left) and normal littermate (right). This mouse contains a 15.9- kb deletion of non-coding sequence located 47.3 kb upstream of the transcription start site in the gene Sox10, leading to loss of skin melanocytes and megacolon 30. (H) A Pax3Sp heterozygote, which displays a characteristic white belly patch. Mice harboring heterozygous mutations of certain alleles of Sox10, Mitf, Kit, and Kitl show a similar belly spotting phenotype.
Mouse mutations have long served as human disease models for many aspects of developmental studies 20, 21. In addition, mouse coat color mutants serve as an excellent model for the study of melanocyte development and pigmentation 22. Owing to the shared embryonic origin of various tissues or pleiotropic effects, these mutants also serve as models of disorders in vision, hearing, craniofacial development, enteric nervous system development, and neural tube closure. White spotting mutations produce white hair and skin in regions where melanocytes normally appear. This phenotype can result from a defect of survival, migration, proliferation, or differentiation at a particular time of melanocyte development when the specific gene product is required. The best-characterized models used in the studies of melanocyte development are Microphthalmia(Mi), Dominant megacolon(Dom) , Splotch(Sp) , Dominant white spotting (W), Steel (Sl), Piebald-lethal(sl) , and Lethal spotting (ls) (Figure 1). All of the mutated genes associated with these models are cloned, and they belong to two categories: transcription factors and receptor/ligand systems. These mutations provide a rich genetic resource for investigation of the mechanisms of melanocyte development at the molecular, cellular, and physiological levels.
Microphthalmia
This locus encodes the basic-helix-loop-helix-leucine-zipper transcription factor Mitf 12. At least 25 different murine mutant alleles of Mitf have been identified, providing a useful genetic resource for studies of development and disease 23, 24. Mitf homozygous mutant mice, such as MitfMi (microphthalmia), Mitfmi-ew (mi-eyeless-white), MitfMi-wh (Mi-white), or Mitfmi-vga-9 (a transgenic insertional allele), typically survive but are microphthalmic, deaf, and completely white, reflecting the complete abolishment of melanocytes (Figure 1) 12, 25. Additionally, a few mutations result in osteoclast defects.
Mutations of human MITF are associated with 10-15% of WS type 2, and patients show skin hypopigmentation, ocular pigmentation defects, and deafness caused by defects of melanocytes of the inner ear 26. Tietz syndrome shows more obvious hypopigmentation and deafness that is also associated with mutations in MITF27.
Dominant megacolon (Dom)
This mouse mutant exhibits white spotting and megacolon in heterozygotes. Homozygous Dom mutations are embryonic lethal (at E13.5) and exhibit absence of melanocytes and enteric neurons, size reductions in the dorsal trigeminal and facial ganglia, and defects in dorsal root ganglia, sympathetic ganglia, and terminal oligodendrocyte differentiation in spinal cord. This locus encodes Sox10 (Sry-like HMB box 10), a member of the high mobility group (HMG) family of transcription factors, showing HMG domain homology to the testis determining factor SRY 28, 29. The Sox10Dom allele results from a point mutation that introduces a frameshift and early truncation that generates a truncated SOX10 protein lacking the transcription activation domain 11. A transgene-insertion mutant mouse line (Hry) was shown to be the result of a 15.9- kb deletion of a non-coding sequence located 47.3 kb upstream of the transcription start site in Sox1030.
Mutations in human SOX10 are associated with WS type 4, also known as Waardenburg-Shah syndrome. The patients show WS characteristics of the white forelock and eyelashes, abnormal iris pigmentation, and deafness, along with enteric aganglionosis, which is seen in patients with Hirschsprung disease 10, 11. Recently, a complex neurocristopathy, PCWH, which shows WS phenotypes along with additional neurological defects, has been shown to result from mutated SOX10 mRNA escaping the nonsense-mediated decay pathway 31. Recent evidence suggests that some WS2 patients harbor SOX10 deletions, and some of these patients also show the neurological phenotypes of PCWH 32.
Splotch
This locus encodes a paired-box homeodomain transcription factor, Pax3, and the Splotch mouse mutant was due to a Pax3 loss- of- function mutation 33. Mice harboring heterozygous Pax3 mutations show ventral spotting, whereas homozygous mutations are embryonic lethal. Pax3 belongs to the Pax gene family, which is highly conserved across species and whose members contain a paired DNA- binding domain 34.
Mutations in human PAX3 are associated with WS type 1 and type 3 or Klein- WS 35. WS type 1 patients show dystopia canthorum, hypopigmentation most often manifested as a white blaze of hair at the forehead or leukoderma, heterochromia iridis, and deafness. WS type 3 patients show additional skeletal abnormalities and cardiopulmonary defects.
Piebald (s) and lethal spotting (ls)
Mutations in the recessive mutants s and ls also disrupt normal melanocyte development. The s locus encodes G protein-coupled Ednrb 16 and the ls locus encodes Edn3, a 21-residue peptide ligand with high affinity for EDNRB 15. The related ligands, EDN1 and EDN2, can also bind EDNRB. Activating mutations in the Ga subunits Gnaq and Gna11 can promote expansion of the early melanoblast population, suggesting that the G protein-coupled receptor plays an important role in regulating melanocyte development 36.
Mutations at the human EDNRB and EDN3 loci are also associated with WS type 4 or Waardenburg-Shah syndrome, which is inherited as an autosomal recessive trait. As described above, the patients show pigmentary defects and enteric aganglionosis 37, 38. Additionally, ABCD syndrome, named for the patients' phenotypic presentation of albinism, black lock, cell migration disorder of the neurocytes of the gut, and deafness, has been identified as a homozygous nonsense mutation in the EDNRB gene 39. EDNRB is also associated with melanoma risk and is required for the expansion of malignant melanoma 40, 41.
Dominant white spotting (W) and Steel (Sl)
Similar to mice with defects in EDN3/EDNRB signaling, mutations in W and Sl also disrupt normal melanocyte development. W encodes the receptor tyrosine kinase Kit (also known as c-Kit) 17. Sl encodes Kitl, also known as stem cell factor (SCF) and mast cell growth factor (MGF) 18. Most alleles of W and Sl in heterozygotes show head and belly spots and the homozygotes are often embryonic lethal; those homozygotes that survive are black-eyed white, sterile, and anemic 42. Kitl produces two KITL proteins, a transmembrane form and a soluble form. The membrane-bound form is required for melanocyte precursor survival in the dermis, whereas the soluble form is needed for melanocyte precursor dispersal on the lateral pathway and/or for their initial survival in the migration staging area 43. In addition, Kit and Kitl mutants have defects in the intestinal pacemaker system, T-cell precursors, and hippocampal learning and hearing 44, 45, 46.
Mutations in human KIT are associated with piebaldism, a rare autosomal- dominant disorder in which patients show patches of white skin and white hair 47. KIT mutations are also associated with human gastrointestinal stromal tumors, urticaria pigmentosa, and aggressive mastocytosis in which KIT proteins are constitutively activated 48, 49. To date KITL mutations have not been found in human patients.
Snai2 knockout mice
Recent evidence suggests that mutations in human SNAI2 are associated with WS2 and piebaldism 50, 51. The initial description of Snai2 knockout mice reported no NC defects and described normal melanocyte generation, migration, and development 52. However, another report showed a strain-dependent phenotype of a small amount of white spotting in the homozygous Snai2 knockout mice 50. The functional role of Snai2 in melanocyte development is not known and requires further investigation.
Microphthalmia-associated transcription factor
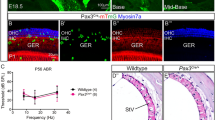
The first identification of the microphthalmia gene, now termed Mitf, was provided by cloning the gene from a microphthalmic and hypopigmented transgene-insertion mutant mouse line 12. Mitf encodes a transcription factor of the basic-helix-loop-helix-leucine zipper (bHLH-Zip) class, which, together with TFE3, TFEB, and TFEC, belongs to the MITF-TFE subfamily of bHLH proteins. The four mammalian members of this subfamily share very similar bHLH and leucine zipper domains and in vitro form all possible combinations of homo- and heterodimers with each other, but do not interact with other bHLH and bHLH-Zip proteins 53. Intriguingly, it has been shown that knockouts of Tfe3, Tfeb, and Tfec did not affect melanocyte development, suggesting that heterodimeric interactions are not essential for MITF-TFE function in melanocyte development 54, L Hou and H Arnheiter, unpublished results). The Mitf gene is quite complex, with at least nine promoters producing multiple isoforms, here termed A-MITF, J-MITF, C-MITF, MC-MITF, E-MITF, H-MITF, D-MITF, B-MITF, and M-MITF (Figure 2). These isoforms differ in their amino termini but share exons 2-9, which include all bHLH-Zip domains. MITF is broadly expressed though the protein levels and isoforms differ among cell types (for detailed structures of MITF, see Steingrimsson et al. 24 and Arnheiter et al. 23). M-MITF is a major isoform in NC stem cell-derived melanocytes. All of the isoforms also produce alternative splice forms modifying exon 6 that lead to inclusion (+) or exclusion (−) of the sequence ACIFPT upstream of the basic domain. The function of MITF (+) or MITF (−) forms are not fully understood in melanocyte development, but they may be related to cell proliferation and different transcriptional activities 55, 56. Distinct extracellular signaling pathways, such as those of WNT, KIT, EDNRB, and α-melanocyte-stimulating hormone (MSH), also regulate Mitf 57, 58, 59, 60. MITF proteins are modified by phosphorylation, ubiquitination, sumoylation, and acetylation 58, 61, 62, 63, and the protein inhibitor of activated STAT3 (PIAS3) inhibits MITF transcriptional activity 64.
Schematic diagram of the mouse Mitf gene and its mutations. The upper part of the figure shows the genomic organization of the gene. The boxes represent exons, with the numbers written on top indicating the corresponding nine distinct exons: 1A, 1J, 1C, 1MC, 1E, 1H, 1D, 1B, and 1M, each associated with a distinct mRNA isoform, and the common exons 2–9. The bHLH-Zip domain (colored pink) is contributed by part of exon 6B, all of exon 7 and 8, and part of exon 9. The lower part of the figure shows 20 of the currently known alleles that have been described in the literature 24, 25, 125, 147. Filled circles represent point mutations, filled triangles represent insertion mutations, and lines represent deletions (Courtesy: Heinz Arnheiter).
Mitf is expressed in developing NC-derived melanocyte precursors before the initial expression of Dopachrome tautomerase (Dct) and in the neuroepithelium-derived retinal pigmented epithelium (RPE) of the eye beginning at E10. On the basis of the coexpression of markers such as Kit and Dct, these NC-derived Mitf-positive cells are defined as melanocyte precursors 19, 65. Mitf is one of the key transcription factors regulating many aspects of melanocyte development and has been referred to as the melanocyte master regulator 66, 67. MITF is required for melanocyte cell survival by directly regulating Bcl2 and MET, the receptor for hepatocyte growth factor 67, 68, and is involved in melanocyte proliferation and cell cycle progression by its regulation of Tbx2, INK4A/p16, p21, and CDK2 69, 70, 71, 72, 73. MITF can also control melanocyte differentiation by directly activating transcription through E-box (CATGTG) binding sites in the melanocyte-specific genes, Dct, Tyrosinase (Tyr), Tyrosinase related protein 1 (Tyrp1), and Silver/Pmel17, Aim-1, Mart1, and MC1R62, 74, 75, 76. Interestingly, recent work suggests that MITF is not the sole regulator of Dct and Tyr in melanocyte development. SOX10 also regulates Dct expression by directly binding to the promoter of Dct 77 and melanocyte-specific expression of Dct is dependent on its synergistic activation by SOX10 and MITF 78, 79. In addition, MITF is not sufficient to induce Tyr expression and full melanocyte differentiation in the absence of functional SOX10, suggesting that Sox10 also may control expression of other melanocyte-specific gene(s) 80.
SOX10 and PAX3 are both broadly expressed in NC stem cells 81, 82. Supporting this similarity in expression patterns, ectopic expression of a Sox10 transgene under the control of regulatory regions from the Pax3 gene in Sox10-deficient NC cells rescues melanocyte differentiation 83. Intriguingly, Sox10 and Pax3 have more expansive expression patterns than M-Mitf and are required for several lineages of NC cells; yet, M-Mitf is activated in a small subset of NC cells and is only required for melanocyte development. These observations suggest that additional extrinsic signaling control must be involved in Mitf regulation. In support of this idea, we have found that MITF is not sufficient to induce the expression of Tyr without functional KIT signaling 19, suggesting that KIT signaling modulates the activity of MITF either directly or indirectly in melanocyte development. Further studies are needed to understand the roles of MITF in melanoblast survival, proliferation, differentiation, and disease.
Transcriptional regulation of Mitf
MITF plays an essential role in survival, migration, proliferation, and differentiation of melanocytes during development. Therefore, understanding the transcriptional regulation of Mitf will help us identify the transcriptional hierarchy that directs the development of melanocytes from NC stem cells. Here we discuss two transcription factors, SOX10 and PAX3.
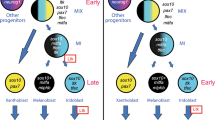
SOX10 is expressed in NC stem cells 81 and in NC-derived structures during embryonic development 11, and is required for proper development and survival of NC-derived melanocyte, glial, and enteric neuron lineages 84, 85, 86. Sox10 function is regulated by sumoylation in Xenopus NC development 87. SOX10 has been shown to strongly activate Mitf expression in cultured cell lines 9, 88 and to regulate Dct expression 77. In addition, SOX10 is required not only for inducing Mitf expression in NC cells, but also for Mitf-dependent Tyr expression 80. These results suggest that SOX10 regulates the expression of other melanocyte-specific gene(s) in addition to Mitf in melanocyte development. In contrast, in zebrafish melanocyte development Sox10 is only required for directly activating mitf, which, independent of the further actions of Sox10, rapidly stimulates downstream target genes and hence pigmentation 89. These results clearly show that distinct species differ in usage of homologous regulators and their targets for melanocyte development. In zebrafish, sox10, mitf, and downstream pigment genes are linked in a linear, seemingly simple, regulatory chain in which sox10 controls the expression of mitf, which in turn is sufficient to regulate melanocyte-specific gene expression and pigmentation. In mice, the situation is apparently more complex in that the generation of melanocytes requires both Sox10 and Mitf, and neither gene alone can overcome the lack of the other to generate tyrosinase-expressing, mature melanocytes (schematically illustrated in Figure 3). This regulatory model was confirmed in mouse melanocytes, in which it was shown that SOX10 cooperates with MITF to regulate Tyr gene expression by direct activation of the Tyr distal regulatory element 90. SOX10's interacting factors and its downstream targets are yet to be fully elucidated in NC stem cell and melanocyte development.
The transcriptional regulatory hierarchy of Sox10 and Mitf in melanocyte development and differentiation is distinct in zebrafish and mice. Examples of the transcriptional regulatory network models are based on reference 149. In zebrafish, melanocyte development exhibits a simple regulatory chain model. Here, Sox10 directly activates mitf and Mitf, independent of the further actions by Sox10, and rapidly stimulates downstream target genes and hence pigmentation. Mouse melanocyte development exhibits a more complex feed-forward loop network model. Here, SOX10 directly regulates Mitf and subsequently cooperates with MITF and/or additional SOX10-dependent regulators to activate downstream target genes, including Tyr. This model allows for temporal control of melanogenic gene expression in mouse.
PAX3 is expressed in NC cells and is required for early NC and melanocyte development 33, 82, 91. PAX3 controls neural tube closure through inhibition of p53-mediated apoptosis 92. PAX3 also up-regulates Tyrp1 promoter activity 93, and overexpression of Pax3 induces tyrosinase activity in ascidian embryos 94. It has been shown that PAX3 weakly transactivates the Mitf promoter 8 and that PAX3 synergistically transactivates the promoter of Mitf with SOX10 8, 9, 88. However, contradictory data showing that PAX3 does not synergistically act with SOX10 to activate Mitf transcription have also been reported 95. In vitro studies have shown that the phorbol ester, 12-tetradecanoylphorbol 13-acetate (TPA), induces melanocyte differentiation from NC cells through Mitf up-regulation, but that Pax3 expression level is not altered by the treatment 96. It is currently unknown whether PAX3 directly regulates Mitf in melanocyte development in vivo. Recently it has been shown that PAX3 represses Dct expression in the absence of activated β-catenin, and such repression is relieved by activated β-catenin in melanocyte stem cells 97. The precise function of PAX3 in NC stem cell and melanocyte development, however, is poorly understood and requires further investigation.
Signaling regulation of Mitf
During melanocyte development, the three major signaling pathways involving WNT, KIT, and EDNRB play essential roles, whereas the roles of other signaling pathways are not readily apparent from analysis of mouse models. For example, despite expression of Met and Erbb3 in melanoblasts, there were no melanocyte defects in Met and Erbb3 knockout mice 98, K Buac and WJ Pavan, unpublished results]. Although α-MSH utilizes cAMP to trigger melanin synthesis and pigmentation of melanocytes through activation of Mitf and Sox10 59, 99, it does not affect melanocyte differentiation in mouse. However, it does function in this capacity in other vertebrates, such as reptiles 100, 101. Below we discuss the roles of WNT, KIT, and EDNRB signaling in melanocyte development (Figure 4).
A simplified schematic showing the features of key signaling pathways in melanocyte development. Green lines represent three major signaling pathways, WNT, KIT, and EDNRB, which are all connected to Mitf. WNT/β-catenin signaling promotes melanoblast development by regulating MITF transcription. KIT and EDNRB signal pathways are not required for the initial expression of Mitf in melanocyte development, but both pathways induce the phosphorylation of MITF in mature melanocytes. However, the KIT signaling-dependent phosphorylation site at serine-73 is not essential for melanocyte development. It is unknown whether melanocytic KIT and EDNRB signaling pathways act through regulation of MITF and whether EDNRB signaling-dependent MITF phosphorylation plays any role in melanocyte development in vivo. MITF is involved in melanocyte survival, proliferation, and differentiation by regulating downstream genes. Blue lines represent MITF target genes, which include genes involved in cell survival (Bcl2 and Met), cell proliferation (p21, p16, CDK2, and Tbx2), and differentiation (Tyr, Tyrp1, Dct, Silver/Pmel17, Mart1, Aim-1, and MC1R). During melanocyte lineage development, the transcription factors SOX10, PAX3, and LEF1 regulate expression of the melanocyte-specific Mitf isoform. The relative positions of the binding sites for these factors within the proximal Mitf promoter are shown. MITF is insufficient to induce Tyr expression and full melanocyte differentiation in the absence of SOX10. SOX10 is required for Dct and Tyrosinase expression in addition to the control of Mitf. Although PAX3 is known to repress Dct expression, the precise function of PAX3 in melanocyte development is poorly understood. In addition, MSH can elevate cAMP levels that subsequently activate both the cAMP and the MAP kinase pathways, resulting in elevated Mitf promoter activity in melanogenesis. However, how CREB activation is involved in melanocyte development is unknown to date. KIT signaling-induced activation of PI3 K is not essential for melanocyte development 111. Question marks indicate insufficient data to describe the involvement of these signaling pathways in melanocyte development.
WNT/β-catenin signaling
WNT signaling is essential for NC induction and melanocyte development. Activation of Frizzled receptors by WNT leads to activation of downstream signal transduction molecules, such as β-catenin, PKC, CAMKII, PKA, and Rho GTPase, resulting in WNT-mediated complex cellular actions. In the best understood canonical WNT/β-catenin signaling pathway, when extracellular WNT ligand binds to its receptor (Frizzled), β-catenin accumulates, enters the nucleus, and subsequently interacts with members of the lymphoid enhancer binding factor 1/T-cell specific factor (Lef1/Tcf) family of transcription factors, which then modulate transcription of target genes 102. Wnt1 and Wnt3 are expressed in the dorsal part of the neural tube in spatiotemporal patterns consistent with the timing of NC induction 57, 103, and Dct-positive cells are markedly reduced in Wnt1/Wnt3 double knockout mouse E11.5 embryos 13.
In vivo and in vitro studies also indicate that the WNT/β-catenin signaling pathway is required for induction of melanocyte and other cell fates. Overexpression of β-catenin in zebrafish promotes melanoblast formation and reduces formation of neurons and glia 57. Similarly, WNT3a or WNT1 promotes the differentiation and expansion of melanocytes in cultured chick NC cells and in cultured mouse NC cells 104, 105. Furthermore, both melanoblasts and sensory neurons are absent in β-catenin conditional knockout mice during embryonic development 106. Interestingly, there is a highly conserved binding site for LEF-1 in the Mitf promoter 14, 107, and the interaction between MITF and LEF-1, but not TCF-1, results in synergistic transactivation of the Dct gene promoter 108. In addition, MITF can interact directly with β-catenin and can redirect its transcriptional activity away from canonical WNT signaling-regulated genes toward MITF-specific target promoters to activate transcription 109. Together these studies suggest that WNT/β-catenin signaling promotes melanoblast development by regulating MITF.
KIT signaling
KIT signaling is required for normal development of three migratory cell populations: blood cells, melanocytes, and primordial germ cells 17, 18, 110. Activation of KIT by KITL leads to receptor dimerization and autophosphorylation of specific tyrosine residues in the kinase domain. This activates downstream signal transduction molecules, such as MAPK, phosphatidylinositol 3′-kinase (PI3K), JAK/STAT, and Src family members. Although KIT signaling-induced activation of PI3K is required for male fertility, this activation is not essential for melanocyte development 111.
KIT is expressed in developing NC cells, hematopoietic stem cells, and primordial germ cells, whereas KITL is expressed in tissues associated with KIT-expressing cells and in the neural tube 43, 112, 113, 114, 115. KIT signaling is necessary for the survival and/or migration of melanoblasts 43, 116, 117. Injection of KIT antibody into early mouse embryos blocks proper melanocyte development 118, 119, and ectopic expression of KITL promotes migration, proliferation, and differentiation of melanocyte precursors 120. Tyrosine residues 567 and 569 of KIT are crucial for its function in melanocyte development, as specific mutation of both residues results in complete loss of melanocytes 121.
Understanding of the complex relationship between KIT signaling and MITF function also concerns their temporal expression patterns in melanocyte development. One possible model is that MITF can activate Kit transcription 122 and thus upregulate KIT expression 65. This model is supported by observations that zebrafish mitf mutants fail to express kit, suggesting that the initial expression of KIT is dependent on MITF 123. Another possible model is that KIT signaling induces the transcriptional activity of MITF. This model is supported by studies on cultured melanocytes and melanoma cells in which KIT signaling leads to an increase in MITF phosphorylation, which is associated with an enhanced recruitment of the transcriptional coactivator p300/CBP and a concomitant stimulation of MITF transcriptional activity 58, 124. This increase is transient and followed by rapid ubiquitination and proteasome-mediated degradation of MITF 61. However, we have shown that the initial expression of Kit is not dependent on MITF and that the initial expression of Mitf is not dependent on KIT. In addition, we have shown that the presence of MITF alone is not sufficient for Tyr expression in melanoblasts in the absence of functional KIT signaling, and that KIT signaling influences gene expression through MITF in a gene-selective manner during melanocyte development 19. Taken together, these results suggest that MITF and KIT are not related in a simple linear regulatory chain, and that both cooperatively regulate the expansion of melanocyte precursors in development. To date, it is unknown when and where KIT signaling-dependent MITF phosphorylation occurs and whether this post-translational modification plays any role in melanocyte development in vivo. Interestingly, one mutant Mitf allele that results in reduced skin melanocytes, Mitfmi-bws (mi-black and white spotting), produces not only a wild-type transcript that contains exon 2b, but also a transcript that lacks exon 2b, which contains a KIT signaling-dependent phosphorylation site, serine-73. This suggests that exon 2b may play a role in melanocyte development, potentially through KIT signaling-dependent phosphorylation. However, targeted mutation of serine-73 to alanine leads to normally pigmented mice 125. This indicates that mutation of this phosphorylation site is not deleterious to melanocyte development. The developmental mechanism of Mitfmi-bws mutant mice and the precise function of Kit signaling in melanocyte development require further investigation.
EDNRB signaling
EDNRB exerts pleiotropic effects on mouse development, and its function is required for the normal development of NC-derived melanocytes and enteric ganglia 15, 16. Binding of EDNRB by EDN3 leads to the activation of downstream signal transduction pathways, including PKC, CamKII, and MAPK 126. EDNRB is expressed in developing NC stem cells, melanoblasts, and enteric ganglia in mouse embryos 16, 127, 128, 129, 130, whereas EDN3 is expressed in tissues associated with Ednrb-expressing cells 127. Studies on mice harboring mutant Ednrb alleles showed that EDNRB signaling functions are necessary for the development of melanoblasts and enteric neural precursors 15, 16, 128, 131. Likewise, transgenic expression of Edn3 prevents aganglionosis and piebaldism in lethal spotted mice 132. Experiments in Ednrb mutant mice have shown that EDNRB is not needed for melanoblast formation, but is needed for migration of melanoblasts and enteric neuroblast precursors prior to cell differentiation, between E10.5 and E12.5 133. In addition, it is also required for melanoblast development in the epidermis beyond E12.5 119. Avian NC cells express an additional Ednrb gene, Ednrb2, which is involved in melanoblast differentiation and migration 134, 135.
Although EDNRB is expressed in unspecified NC cells and melanocyte precursors, it is not clear whether it acts solely in a cell-autonomous manner 16, 130, 136. By cross-explantation of embryonic tissues and NC cells, it has been found that the melanoblasts of the hypomorphic Ednrbs (piebald) allele show increased survival on in vitro cultured wild-type skin compared with mutant skin 137. To address the question of whether melanocyte development depends entirely on the cell-autonomous action of EDNRB, we have performed a series of tissue recombination experiments in vitro using NC cell cultures from EdnrblacZembryos, which contain a functionally null allele of Ednrb. These studies showed that EDNRB plays a significant role during melanocyte differentiation by sequential cell-autonomous and non-autonomous actions 138. Recently it has been shown that endothelin signaling leads to an increase in MITF phosphorylation and CREB phosphorylation in cultured human melanocytes 60. However, it is unknown whether melanocytic EDNRB signaling acts through regulation of MITF, when and where the MITF post-translational modifications occur, and whether EDNRB signaling-dependent MITF phosphorylation plays any role in melanocyte development in vivo.
In addition, how EDNRB itself is regulated in melanocyte development is unknown. Interestingly, it has been shown that SOX10 directly activates Ednrb transcription in NC stem cell-derived enteric neurons 139, but genetic evidence suggests that SOX10 does not directly activate Ednrb transcription in the melanocyte lineage 140. However, contradictory data showed that SOX10 transactivates the Ednrb promoter in human cultured melanocytes 141. These results suggest that SOX10 may regulate differentiation-related downstream target gene(s) based on the cellular context in development.
Notch signaling
Recent work showed that Notch signaling is involved in the maintenance of melanoblasts and melanocyte stem cells 142, 143. The precise function of Notch signaling in the maintenance of melanocyte stem cells, however, requires further investigation. For detailed information on the Notch signaling pathway and its general role in melanocytes, we refer the readers to other recent reviews 144, 145.
Conclusions
White spotting genes play essential roles in NC stem cell-derived melanocyte development and related diseases. Nevertheless, much work is required to complete our functional understanding of these genes in melanocyte development. One of the important questions is when and how this complex network of genes interacts with other genes to regulate proper melanocyte development. Current evidence suggests that MITF is extensively involved in melanocyte development, providing a central link between transcription factors and signaling pathways (Figure 4), and is also involved in melanocyte stem cell maintenance 146. Do all transcription factors and signaling pathways use MITF to regulate melanocyte development? More research is needed to answer this question. For example, it is unknown if and/or how PAX3 or SNAI2 regulates melanocyte development via Mitf. In addition, it will be very interesting to determine how WNT, KIT, and EDNRB signaling pathways regulate MITF and whether Notch signaling influences the maintenance of melanocyte stem cells through regulation of MITF. It is unknown whether KIT- and EDNRB-dependent MITF phosphorylation plays any role in melanocyte development in vivo. Increasing evidence suggests that signaling proteins tend to form networks of interactions rather than simple linear pathways. Therefore, most importantly, we need to further understand how distinct signaling pathways form interacting networks to regulate specification, survival, migration, proliferation, and differentiation of melanocyte precursors during development and how these pathways influence the dynamic balance between stem cell maintenance and differentiation in tissues of mature melanocytes. The field of melanocyte research has grown to include developmental cell biology and cancer biology, and will continue to provide a fruitful ground for basic and translational research in the future.
References
Le Douarin NM, Kalcheim C, eds. The Neural Crest. Cambridge: Cambridge University Press, 1999.
Hou L, Takeuchi T . Neural crest development in reptilian embryos, studied with monoclonal antibody, HNK-1. Zool Sci 1994; 11:423–431.
Hall BK, ed. The Neural Crest in Development and Evolution. New York: Springer, 1999.
Bolande RP . Neurocristopathy: its growth and development in 20 years. Pediatr Pathol Lab Med 1997; 17:1–25.
Read AP, Newton VE . Waardenburg syndrome. J Med Genet 1997; 34:656–665.
Perlis C, Herlyn M . Recent advances in melanoma biology. Oncologist 2004; 9:182–187.
Nishimura EK, Jordan SA, Oshima H, et al. Dominant role of the niche in melanocyte stem-cell fate determination. Nature 2002; 416:854–860.
Watanabe A, Takeda K, Ploplis B, Tachibana M . Epistatic relationship between Waardenburg syndrome genes MITF and PAX3. Nat Genet 1998; 18:283–286.
Potterf SB, Furumura M, Dunn KJ, Arnheiter H, Pavan WJ . Transcription factor hierarchy in Waardenburg syndrome: regulation of MITF expression by SOX10 and PAX3. Hum Genet 2000; 107:1–6.
Pingault V, Bondurand N, Kuhlbrodt K, et al. SOX10 mutations in patients with Waardenburg-Hirschsprung disease. Nat Genet 1998; 18:171–173.
Southard-Smith EM, Kos L, Pavan WJ . Sox10 mutation disrupts neural crest development in Dom Hirschsprung mouse model. Nat Genet 1998; 18:60–64.
Hodgkinson CA, Moore KJ, Nakayama A, et al. Mutations at the mouse microphthalmia locus are associated with defects in a gene encoding a novel basic-helix-loop-helix-zipper protein. Cell 1993; 74:395–404.
Ikeya M, Lee SM, Johnson JE, McMahon AP, Takada S . Wnt signalling required for expansion of neural crest and CNS progenitors. Nature 1997; 389:966–970.
Dorsky RI, Raible DW, Moon RT . Direct regulation of nacre, a zebrafish MITF homolog required for pigment cell formation, by the Wnt pathway. Genes Dev 2000; 14:158–162.
Baynash AG, Hosoda K, Giaid A, et al. Interaction of endothelin-3 with endothelin-B receptor is essential for development of epidermal melanocytes and enteric neurons. Cell 1994; 79:1277–1285.
Hosoda K, Hammer RE, Richardson JA, et al. Targeted and natural (piebald-lethal) mutations of endothelin-B receptor gene produce megacolon associated with spotted coat color in mice. Cell 1994; 79:1267–1276.
Chabot B, Stephenson DA, Chapman VM, Besmer P, Bernstein A . The proto-oncogene c-kit encoding a transmembrane tyrosine kinase receptor maps to the mouse W locus. Nature 1988; 335:88–89.
Copeland NG, Gilbert DJ, Cho BC, et al. Mast cell growth factor maps near the steel locus on mouse chromosome 10 and is deleted in a number of steel alleles. Cell 1990; 63:175–183.
Hou L, Panthier JJ, Arnheiter H . Signaling and transcriptional regulation in the neural crest-derived melanocyte lineage: interactions between KIT and MITF. Development 2000; 127:5379–5389.
Silver L, ed. Mouse Genetics Concepts and Applications. New York: Oxford University Press, 1995.
Barsh GS . The genetics of pigmentation: from fancy genes to complex traits. Trends Genet 1996; 12:299–305.
Baxter LL, Hou L, Loftus SK, Pavan WJ . Spotlight on spotted mice: a review of white spotting mouse mutants and associated human pigmentation disorders. Pigment Cell Res 2004; 17:215–224.
Arnheiter H, Hou L, Nguyen MT, et al. Mitf – A matter of life and death for the developing melanocyte. In: Hearing VJ, Leong SPL, eds. From Melanocytes to Malignant Melanoma. Totowa, NJ: Humana Press, 2006:27–49.
Steingrimsson E, Copeland NG, Jenkins NA . Melanocytes and the microphthalmia transcription factor network. Annu Rev Genet 2004; 38:365–411.
Steingrimsson E, Moore KJ, Lamoreux ML, et al. Molecular basis of mouse microphthalmia (mi) mutations helps explain their developmental and phenotypic consequences. Nat Genet 1994; 8:256–263.
Tassabehji M, Newton VE, Read AP . Waardenburg syndrome type 2 caused by mutations in the human microphthalmia (MITF) gene. Nat Genet 1994; 8:251–255.
Amiel J, Watkin PM, Tassabehji M, Read AP, Winter RM . Mutation of the MITF gene in albinism-deafness syndrome (Tietz syndrome). Clin Dysmorphol 1998; 7:17–20.
Mollaaghababa R, Pavan WJ . The importance of having your SOX on: role of SOX10 in the development of neural crest-derived melanocytes and glia. Oncogene 2003; 22:3024–3034.
Wegner M . Secrets to a healthy Sox life: lessons for melanocytes. Pigment Cell Res 2005; 18:74–85.
Antonellis A, Bennett WR, Menheniott TR, et al. Deletion of long-range sequences at Sox10 compromises developmental expression in a mouse model of Waardenburg-Shah (WS4) syndrome. Hum Mol Genet 2006; 15:259–271.
Inoue K, Khajavi M, Ohyama T, et al. Molecular mechanism for distinct neurological phenotypes conveyed by allelic truncating mutations. Nat Genet 2004; 36:361–369.
Bondurand N, Dastot-Le Moal F, Stanchina L, et al. Deletions at the SOX10 gene locus cause Waardenburg syndrome types 2 and 4. Am J Hum Genet 2007; 81:1169–1185.
Epstein DJ, Vekemans M, Gros P . Splotch (Sp2H), a mutation affecting development of the mouse neural tube, shows a deletion within the paired homeodomain of Pax-3. Cell 1991; 67:767–774.
Epstein JA . Pax3 and vertebrate development. Methods Mol Biol 2000; 137:459–470.
Tassabehji M, Read AP, Newton VE, et al. Mutations in the PAX3 gene causing Waardenburg syndrome type 1 and type 2. Nat Genet 1993; 3:26–30.
Van Raamsdonk CD, Fitch KR, Fuchs H, de Angelis MH, Barsh GS . Effects of G-protein mutations on skin color. Nat Genet 2004; 36:961–968.
Puffenberger EG, Hosoda K, Washington SS, et al. A missense mutation of the endothelin-B receptor gene in multigenic Hirschsprung's disease. Cell 1994; 79:1257–1266.
Edery P, Attie T, Amiel J, et al. Mutation of the endothelin-3 gene in the Waardenburg-Hirschsprung disease (Shah-Waardenburg syndrome). Nat Genet 1996; 12:442–444.
Verheij JB, Kunze J, Osinga J, van Essen AJ, Hofstra RM . ABCD syndrome is caused by a homozygous mutation in the EDNRB gene. Am J Med Genet 2002; 108:223–225.
Lahav R . Endothelin receptor B is required for the expansion of melanocyte precursors and malignant melanoma. Int J Dev Biol 2005; 49:173–180.
Soufir N, Meziani R, Lacapere JJ, et al. Association between endothelin receptor B nonsynonymous variants and melanoma risk. J Natl Cancer Inst 2005; 97:1297–1301.
Ruan HB, Zhang N, Gao X . Identification of a novel point mutation of mouse proto-oncogene c-kit through N-ethyl-N-nitrosourea mutagenesis. Genetics 2005; 169:819–831.
Wehrle-Haller B, Weston JA . Soluble and cell-bound forms of steel factor activity play distinct roles in melanocyte precursor dispersal and survival on the lateral neural crest migration pathway. Development 1995; 121:731–742.
Huizinga JD, Thuneberg L, Kluppel M, et al. W/kit gene required for interstitial cells of Cajal and for intestinal pacemaker activity. Nature 1995; 373:347–349.
Motro B, Wojtowicz JM, Bernstein A, van der Kooy D . Steel mutant mice are deficient in hippocampal learning but not long-term potentiation. Proc Natl Acad Sci USA 1996; 93:1808–1813.
Rodewald HR, Kretzschmar K, Swat W, Takeda S . Intrathymically expressed c-kit ligand (stem cell factor) is a major factor driving expansion of very immature thymocytes in vivo. Immunity 1995; 3:313–319.
Giebel LB, Spritz RA . Mutation of the KIT (mast/stem cell growth factor receptor) protooncogene in human piebaldism. Proc Natl Acad Sci USA 1991; 88:8696–8699.
Longley BJ, Tyrrell L, Lu SZ, et al. Somatic c-KIT activating mutation in urticaria pigmentosa and aggressive mastocytosis: establishment of clonality in a human mast cell neoplasm. Nat Genet 1996; 12:312–314.
Hirota S, Isozaki K, Moriyama Y, et al. Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 1998; 279:577–580.
Sanchez-Martin M, Rodriguez-Garcia A, Perez-Losada J, et al. SLUG (SNAI2) deletions in patients with Waardenburg disease. Hum Mol Genet 2002; 11:3231–3236.
Sanchez-Martin M, Perez-Losada J, Rodriguez-Garcia A, et al Deletion of the SLUG (SNAI2) gene results in human piebaldism. Am J Med Genet A 2003; 122:125–132.
Jiang R, Lan Y, Norton CR, Sundberg JP, Gridley T . The Slug gene is not essential for mesoderm or neural crest development in mice. Dev Biol 1998; 198:277–285.
Hemesath TJ, Steingrimsson E, McGill G, et al. Microphthalmia, a critical factor in melanocyte development, defines a discrete transcription factor family. Genes Dev 1994; 8:2770–2780.
Steingrimsson E, Tessarollo L, Pathak B, et al. Mitf and Tfe3, two members of the Mitf-Tfe family of bHLH-Zip transcription factors, have important but functionally redundant roles in osteoclast development. Proc Natl Acad Sci USA 2002; 99:4477–4482.
Bismuth K, Maric D, Arnheiter H . MITF and cell proliferation: the role of alternative splice forms. Pigment Cell Res 2005; 18:349–359.
Murakami M, Iwata Y, Funaba M . Expression and transcriptional activity of alternative splice variants of Mitf exon 6. Mol Cell Biochem 2007; 303:251–257.
Dorsky RI, Moon RT, Raible DW . Control of neural crest cell fate by the Wnt signalling pathway. Nature 1998; 396:370–373.
Hemesath TJ, Price ER, Takemoto C, Badalian T, Fisher DE . MAP kinase links the transcription factor Microphthalmia to c-Kit signalling in melanocytes. Nature 1998; 391:298–301.
Price ER, Horstmann MA, Wells AG, et al alpha-Melanocyte-stimulating hormone signaling regulates expression of microphthalmia, a gene deficient in Waardenburg syndrome. J Biol Chem 1998; 273:33042–33047.
Sato-Jin K, Nishimura EK, Akasaka E, et al. Epistatic connections between microphthalmia-associated transcription factor and endothelin signaling in Waardenburg syndrome and other pigmentary disorders. The FASEB Journal 2008; 22:1155–1168.
Xu W, Gong L, Haddad MM, et al. Regulation of microphthalmia-associated transcription factor MITF protein levels by association with the ubiquitin-conjugating enzyme hUBC9. Exp Cell Res 2000; 255:135–143.
Du J, Fisher DE . Identification of Aim-1 as the underwhite mouse mutant and its transcriptional regulation by MITF. J Biol Chem 2002; 277:402–406.
Murakami H, Arnheiter H . Sumoylation modulates transcriptional activity of MITF in a promoter-specific manner. Pigment Cell Res 2005; 18:265–277.
Levy C, Nechushtan H, Razin E . A new role for the STAT3 inhibitor, PIAS3: a repressor of microphthalmia transcription factor. J Biol Chem 2002; 277:1962–1966.
Opdecamp K, Nakayama A, Nguyen MT, et al Melanocyte development in vivo and in neural crest cell cultures: crucial dependence on the Mitf basic-helix-loop-helix-zipper transcription factor. Development 1997; 124:2377–2386.
Bejar J, Hong Y, Schartl M . Mitf expression is sufficient to direct differentiation of medaka blastula derived stem cells to melanocytes. Development 2003; 130:6545–6553.
McGill GG, Horstmann M, Widlund HR, et al. Bcl2 regulation by the melanocyte master regulator Mitf modulates lineage survival and melanoma cell viability. Cell 2002; 109:707–718.
McGill GG, Haq R, Nishimura EK, Fisher DE . c-Met expression is regulated by Mitf in the melanocyte lineage. J Biol Chem 2006; 281:10365–10373.
Carreira S, Liu B, Goding CR . The gene encoding the T-box factor Tbx2 is a target for the microphthalmia-associated transcription factor in melanocytes. J Biol Chem 2000; 275:21920–21927.
Du J, Widlund HR, Horstmann MA, et al. Critical role of CDK2 for melanoma growth linked to its melanocyte-specific transcriptional regulation by MITF. Cancer Cell 2004; 6:565–576.
Carreira S, Goodall J, Aksan I, et al. Mitf cooperates with Rb1 and activates p21Cip1 expression to regulate cell cycle progression. Nature 2005; 433:764–769.
Loercher AE, Tank EM, Delston RB, Harbour JW . MITF links differentiation with cell cycle arrest in melanocytes by transcriptional activation of INK4A. J Cell Biol 2005; 168:35–40.
Carreira S, Goodall J, Denat L, et al. Mitf regulation of Dia1 controls melanoma proliferation and invasiveness. Genes Dev 2006; 20:3426–3439.
Goding CR . Mitf from neural crest to melanoma: signal transduction and transcription in the melanocyte lineage. Genes Dev 2000; 14:1712–1728.
Du J, Miller AJ, Widlund HR, et al. MLANA/MART1 and SILV/PMEL17/GP100 are transcriptionally regulated by MITF in melanocytes and melanoma. Am J Pathol 2003; 163:333–343.
Adachi S, Morii E, Kim D, et al. Involvement of mi-transcription factor in expression of alpha-melanocyte-stimulating hormone receptor in cultured mast cells of mice. J Immunol 2000; 164:855–860.
Potterf SB, Mollaaghababa R, Hou L, et al. Analysis of SOX10 function in neural crest-derived melanocyte development: SOX10-dependent transcriptional control of dopachrome tautomerase. Dev Biol 2001; 237:245–257.
Jiao Z, Mollaaghababa R, Pavan WJ, et al. Direct interaction of Sox10 with the promoter of murine Dopachrome Tautomerase (Dct) and synergistic activation of Dct expression with Mitf. Pigment Cell Res 2004; 17:352–362.
Ludwig A, Rehberg S, Wegner M . Melanocyte-specific expression of dopachrome tautomerase is dependent on synergistic gene activation by the Sox10 and Mitf transcription factors. FEBS Lett 2004; 556:236–244.
Hou L, Arnheiter H, Pavan WJ . Interspecies difference in the regulation of melanocyte development by SOX10 and MITF. Proc Natl Acad Sci USA 2006; 103:9081–9085.
Kim J, Lo L, Dormand E, Anderson DJ . SOX10 maintains multipotency and inhibits neuronal differentiation of neural crest stem cells. Neuron 2003; 38:17–31.
Magnaghi P, Roberts C, Lorain S, Lipinski M, Scambler PJ . HIRA, a mammalian homologue of Saccharomyces cerevisiae transcriptional co-repressors, interacts with Pax3. Nat Genet 1998; 20:74–77.
Hou L, Loftus SK, Incao A, Chen A, Pavan WJ . Complementation of melanocyte development in SOX10 mutant neural crest using lineage-directed gene transfer. Dev Dyn 2004; 229:54–62.
Britsch S, Goerich DE, Riethmacher D, et al. The transcription factor Sox10 is a key regulator of peripheral glial development. Genes Dev 2001; 15:66–78.
Aoki Y, Saint-Germain N, Gyda M, et al. Sox10 regulates the development of neural crest-derived melanocytes in Xenopus. Dev Biol 2003; 259:19–33.
Honore SM, Aybar MJ, Mayor R . Sox10 is required for the early development of the prospective neural crest in Xenopus embryos. Dev Biol 2003; 260:79–96.
Taylor KM, Labonne C . SoxE factors function equivalently during neural crest and inner ear development and their activity is regulated by SUMOylation. Dev Cell 2005; 9 : 593–603.
Bondurand N, Pingault V, Goerich DE, et al. Interaction among SOX10, PAX3 and MITF, three genes altered in Waardenburg syndrome. Hum Mol Genet 2000; 9:1907–1917.
Elworthy S, Lister JA, Carney TJ, Raible DW, Kelsh RN . Transcriptional regulation of mitfa accounts for the sox10 requirement in zebrafish melanophore development. Development 2003; 130:2809–2818.
Murisier F, Guichard S, Beermann F . The tyrosinase enhancer is activated by Sox10 and Mitf in mouse melanocytes. Pigment Cell Res 2007; 20:173–184.
Hornyak TJ, Hayes DJ, Chiu LY, Ziff EB . Transcription factors in melanocyte development: distinct roles for Pax-3 and Mitf. Mech Dev 2001; 101:47–59.
Pani L, Horal M, Loeken MR . Rescue of neural tube defects in Pax-3-deficient embryos by p53 loss of function: implications for Pax-3- dependent development and tumorigenesis. Genes Dev 2002; 16:676–680.
Galibert MD, Yavuzer U, Dexter TJ, Goding CR . Pax3 and regulation of the melanocyte-specific tyrosinase-related protein-1 promoter. J Biol Chem 1999; 274:26894–26900.
Yajima I, Endo K, Sato S, et al. Cloning and functional analysis of ascidian Mitf in vivo: insights into the origin of vertebrate pigment cells. Mech Dev 2003; 120:1489–1504.
Lee M, Goodall J, Verastegui C, Ballotti R, Goding CR . Direct regulation of the Microphthalmia promoter by Sox10 links Waardenburg-Shah syndrome (WS4)-associated hypopigmentation and deafness to WS2. J Biol Chem 2000; 275:37978–37983.
Prince S, Wiggins T, Hulley PA, Kidson SH . Stimulation of melanogenesis by tetradecanoylphorbol 13-acetate (TPA) in mouse melanocytes and neural crest cells. Pigment Cell Res 2003; 16:26–34.
Lang D, Lu MM, Huang L, et al. Pax3 functions at a nodal point in melanocyte stem cell differentiation. Nature 2005; 433:884–887.
Kos L, Aronzon A, Takayama H, et al. Hepatocyte growth factor/scatter factor-MET signaling in neural crest-derived melanocyte development. Pigment Cell Res 1999; 12:13–21.
Huber WE, Price ER, Widlund HR, et al. A tissue-restricted cAMP transcriptional response: SOX10 modulates alpha-melanocyte-stimulating hormone-triggered expression of microphthalmia-associated transcription factor in melanocytes. J Biol Chem 2003; 278:45224–45230.
Hou L, Takeuchi T . Differentiation of reptilian neural crest cells in vitro. In Vitro Cell Dev Biol 1992; 28A:348–354.
Hou L, Kwon BS . Turtle lung cells produce a melanization-stimulating activity that promotes melanocytic differentiation of avian neural crest cells. Pigment Cell Res 1995; 8:113–119.
Nelson WJ, Nusse R . Convergence of Wnt, beta-catenin, and cadherin pathways. Science 2004; 303:1483–1487.
Roelink H, Nusse R . Expression of two members of the Wnt family during mouse development – restricted temporal and spatial patterns in the developing neural tube. Genes Dev 1991; 5:381–388.
Jin EJ, Erickson CA, Takada S, Burrus LW . Wnt and BMP signaling govern lineage segregation of melanocytes in the avian embryo. Dev Biol 2001; 233:22–37.
Dunn KJ, Brady M, Ochsenbauer-Jambor C, et al. WNT1 and WNT3a promote expansion of melanocytes through distinct modes of action. Pigment Cell Res 2005; 18:167–180.
Hari L, Brault V, Kleber M, et al. Lineage-specific requirements of beta-catenin in neural crest development. J Cell Biol 2002; 159:867–880.
Takeda K, Yasumoto K, Takada R, et al. Induction of melanocyte-specific microphthalmia-associated transcription factor by Wnt-3a. J Biol Chem 2000; 275:14013–14016.
Yasumoto K, Takeda K, Saito H, et al. Microphthalmia-associated transcription factor interacts with LEF-1, a mediator of Wnt signaling. EMBO J 2002; 21:2703–2714.
Schepsky A, Bruser K, Gunnarsson GJ, et al. The microphthalmia-associated transcription factor Mitf interacts with beta-catenin to determine target gene expression. Mol Cell Biol 2006; 26:8914–8927.
Steel KP, Davidson DR, Jackson IJ . TRP-2/DT, a new early melanoblast marker, shows that steel growth factor (c-kit ligand) is a survival factor. Development 1992; 115:1111–1119.
Blume-Jensen P, Jiang G, Hyman R, et al. Kit/stem cell factor receptor-induced activation of phosphatidylinositol 3'-kinase is essential for male fertility. Nat Genet 2000; 24:157–162.
Nocka K, Majumder S, Chabot B, et al. Expression of c-kit gene products in known cellular targets of W mutations in normal and W mutant mice – evidence for an impaired c-kit kinase in mutant mice. Genes Dev 1989; 3:816–826.
Matsui Y, Zsebo KM, Hogan BL . Embryonic expression of a haematopoietic growth factor encoded by the Sl locus and the ligand for c-kit. Nature 1990; 347:667–669.
Keshet E, Lyman SD, Williams DE, et al. Embryonic RNA expression patterns of the c-kit receptor and its cognate ligand suggest multiple functional roles in mouse development. EMBO J 1991; 10:2425–2435.
Motro B, van der Kooy D, Rossant J, Reith A, Bernstein A . Contiguous patterns of c-kit and steel expression: analysis of mutations at the W and Sl loci. Development 1991; 113:1207–1221.
Cable J, Jackson IJ, Steel KP . Mutations at the W locus affect survival of neural crest-derived melanocytes in the mouse. Mech Dev 1995; 50:139–150.
Mackenzie MA, Jordan SA, Budd PS, Jackson IJ . Activation of the receptor tyrosine kinase Kit is required for the proliferation of melanoblasts in the mouse embryo. Dev Biol 1997; 192:99–107.
Nishikawa S, Kusakabe M, Yoshinaga K, et al. In utero manipulation of coat color formation by a monoclonal anti-c-kit antibody: two distinct waves of c-kit-dependency during melanocyte development. EMBO J 1991; 10:2111–2118.
Yoshida H, Kunisada T, Kusakabe M, Nishikawa S, Nishikawa SI . Distinct stages of melanocyte differentiation revealed by analysis of nonuniform pigmentation patterns. Development 1996; 122:1207–1214.
Kunisada T, Yoshida H, Yamazaki H, et al. Transgene expression of steel factor in the basal layer of epidermis promotes survival, proliferation, differentiation and migration of melanocyte precursors. Development 1998; 125:2915–2923.
Kimura Y, Jones N, Kluppel M, et al. Targeted mutations of the juxtamembrane tyrosines in the Kit receptor tyrosine kinase selectively affect multiple cell lineages. Proc Natl Acad Sci USA 2004; 101:6015–6020.
Tsujimura T, Morii E, Nozaki M, et al. Involvement of transcription factor encoded by the mi locus in the expression of c-kit receptor tyrosine kinase in cultured mast cells of mice. Blood 1996; 88:1225–1233.
Lister JA, Robertson CP, Lepage T, Johnson SL, Raible DW . nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 1999; 126:3757–3767.
Sato S, Roberts K, Gambino G, et al. CBP/p300 as a co-factor for the Microphthalmia transcription factor. Oncogene 1997; 14:3083–3092.
Bismuth K, Skuntz S, Hallsson JH, et al. An unstable targeted allele of the mouse mitf gene with a high somatic and germline reversion rate. Genetics 2008; 178:259–272.
Imokawa G, Kobayasi T, Miyagishi M . Intracellular signaling mechanisms leading to synergistic effects of endothelin-1 and stem cell factor on proliferation of cultured human melanocytes. Cross-talk via trans-activation of the tyrosine kinase c-kit receptor. J Biol Chem 2000; 275:33321–33328.
Nataf V, Lecoin L, Eichmann A, Le Douarin NM . Endothelin-B receptor is expressed by neural crest cells in the avian embryo. Proc Natl Acad Sci USA 1996; 93:9645–9650.
Reid K, Turnley AM, Maxwell GD, et al. Multiple roles for endothelin in melanocyte development: regulation of progenitor number and stimulation of differentiation. Development 1996; 122:3911–3919.
Lee HO, Levorse JM, Shin MK . The endothelin receptor-B is required for the migration of neural crest-derived melanocyte and enteric neuron precursors. Dev Biol 2003; 259:162–175.
Opdecamp K, Kos L, Arnheiter H, Pavan WJ . Endothelin signalling in the development of neural crest-derived melanocytes. Biochem Cell Biol 1998; 76:1093–1099.
Pavan WJ, Tilghman SM . Piebald lethal (sl) acts early to disrupt the development of neural crest-derived melanocytes. Proc Natl Acad Sci USA 1994; 91:7159–7163.
Rice J, Doggett B, Sweetser DA, et al. Transgenic rescue of aganglionosis and piebaldism in lethal spotted mice. Dev Dyn 2000; 217:120–132.
Shin MK, Levorse JM, Ingram RS, Tilghman SM . The temporal requirement for endothelin receptor-B signalling during neural crest development. Nature 1999; 402:496–501.
Lecoin L, Sakurai T, Ngo MT, et al. Cloning and characterization of a novel endothelin receptor subtype in the avian class. Proc Natl Acad Sci USA 1998; 95:3024–3029.
Pla P, Alberti C, Solov'eva O, et al. Ednrb2 orients cell migration towards the dorsolateral neural crest pathway and promotes melanocyte differentiation. Pigment Cell Res 2005; 18:181–187.
Parichy DM, Mellgren EM, Rawls JF, et al. Mutational analysis of endothelin receptor b1 (rose) during neural crest and pigment pattern development in the zebrafish Danio rerio. Dev Biol 2000; 227:294–306.
Mayer TC . Enhancement of melanocyte development from piebald neural crest by a favorable tissue environment. Dev Biol 1977; 56:255–262.
Hou L, Pavan WJ, Shin MK, Arnheiter H . Cell-autonomous and cell non-autonomous signaling through endothelin receptor B during melanocyte development. Development 2004; 131:3239–3247.
Zhu L, Lee HO, Jordan CS, et al. Spatiotemporal regulation of endothelin receptor-B by SOX10 in neural crest-derived enteric neuron precursors. Nat Genet 2004; 36:732–737.
Hakami RM, Hou L, Baxter LL, et al. Genetic evidence does not support direct regulation of EDNRB by SOX10 in migratory neural crest and the melanocyte lineage. Mech Dev 2006; 123:124–134.
Yokoyama S, Takeda K, Shibahara S . SOX10, in combination with Sp1, regulates the endothelin receptor type B gene in human melanocyte lineage cells. FEBS J 2006; 273:1805–1820.
Moriyama M, Osawa M, Mak SS, et al. Notch signaling via Hes1 transcription factor maintains survival of melanoblasts and melanocyte stem cells. J Cell Biol 2006; 173:333–339.
Schouwey K, Delmas V, Larue L, et al. Notch1 and Notch2 receptors influence progressive hair graying in a dose-dependent manner. Dev Dyn 2007; 236:282–289.
Pinnix CC, Herlyn M . The many faces of Notch signaling in skin-derived cells. Pigment Cell Res 2007; 20:458–465.
Buac K, Pavan WJ . Stem cells of the melanocyte lineage. Cancer Biomark 2007; 3:203–209.
Nishimura EK, Granter SR, Fisher DE . Mechanisms of hair graying: incomplete melanocyte stem cell maintenance in the niche. Science 2005; 307:720–724.
Bharti K, Liu W, Csermely T, Bertuzzi S, Arnheiter H . Alternative promoter use in eye development: the complex role and regulation of the transcription factor MITF. Development 2008; 135:1169–1178.
Bernex F, De Sepulveda P, Kress C, et al. Spatial and temporal patterns of c-kit-expressing cells in WlacZ/+ and WlacZ/WlacZ mouse embryos. Development 1996; 122:3023–3033.
Lee TI, Rinaldi NJ, Robert F, et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 2002; 298:799–804.
Acknowledgements
We would like to thank Dr Heinz Arnheiter (NIH/NINDS) for generously contributing to the images, and Dr Laura Baxter and Dr Yingzi Yang (NIH/NHGRI) for thoughtful comments on the manuscript. We also acknowledge the support by the National Natural Science Foundation of China (30771149) and the Intramural Research Program of the National Human Genome Research Institute, National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hou, L., Pavan, W. Transcriptional and signaling regulation in neural crest stem cell-derived melanocyte development: do all roads lead to Mitf?. Cell Res 18, 1163–1176 (2008). https://doi.org/10.1038/cr.2008.303
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2008.303
Keywords
This article is cited by
-
Screening and expression analysis of genes related to skin coloration in pearlscale angelfish (Centropyge vrolikii)
Aquaculture International (2024)
-
A novel variant of the SOX10 gene associated with Waardenburg syndrome type IV
BMC Medical Genomics (2023)
-
Genetic insights, disease mechanisms, and biological therapeutics for Waardenburg syndrome
Gene Therapy (2022)
-
Melanocytotoxic chemicals and their toxic mechanisms
Toxicological Research (2022)
-
Using genomic resources for linkage analysis in Peromyscus with an application for characterizing Dominant Spot
BMC Genomics (2020)