Abstract
Carotenoids, a class of natural pigments found in all photosynthetic organisms, are involved in a variety of physiological processes, including coloration, photoprotection, biosynthesis of abscisic acid (ABA) and chloroplast biogenesis. Although carotenoid biosynthesis has been well studied biochemically, the genetic basis of the pathway is not well understood. Here, we report the characterization of two allelic Arabidopsis mutants, spontaneous cell death1-1 (spc1-1) and spc1-2. The weak allele spc1-1 mutant showed characteristics of bleached leaves, accumulation of superoxide and mosaic cell death. The strong mutant allele spc1-2 caused a complete arrest of plant growth and development shortly after germination, leading to a seedling-lethal phenotype. Genetic and molecular analyses indicated that SPC1 encodes a putative ζ-carotene desaturase (ZDS) in the carotenoid biosynthesis pathway. Analysis of carotenoids revealed that several major carotenoid compounds downstream of SPC1/ZDS were substantially reduced in spc1-1, suggesting that SPC1 is a functional ZDS. Consistent with the downregulated expression of CAO and PORB, the chlorophyll content was decreased in spc1-1 plants. In addition, expression of Lhcb1.1, Lhcb1.4 and RbcS was absent in spc1-2, suggesting the possible involvement of carotenoids in the plastid-to-nucleus retrograde signaling. The spc1-1 mutant also displays an ABA-deficient phenotype that can be partially rescued by the externally supplied phytohormone. These results suggest that SPC1/ZDS is essential for biosynthesis of carotenoids and plays a crucial role in plant growth and development.
Similar content being viewed by others
Introduction
Carotenoids are a diverse and widespread group of isoprenoid pigments that are essential compounds in all photosynthetic organisms. In higher plants, carotenoids are largely tetraterpenoid pigments of chloroplast and chromoplast membranes, characterized as a C40 backbone with polyene chains. Carotenoids not only are important accessory pigments in the light-harvesting complex but also function as redox intermediates in the process of electron transfer in photosystem II (PS II) 1, 2, 3, 4, 5, 6. Moreover, carotenoids play important roles in photoprotection by quenching free radicals, singlet oxygen and other reactive oxygen species (ROS) derived from excessive light, thereby protecting plants from these oxidative insults 4, 7. These pigments are also known to provide the yellow, red and orange coloration of many fruits, flowers and vegetables, which, in nature, are beneficial for pollination and seed spreading 8, 9. In addition, carotenoids are precursors of the plant phytohormone abscisic acid (ABA). Therefore, defective carotenoid biosynthesis often results in reduced de novo synthesis of ABA, which, in turn, causes abnormal development as well as an altered response to various stresses 10, 11, 12.
During the past several decades, carotenoids have been well studied biochemically in various photosynthetic organisms. Genes encoding many enzymes of the pathway have been cloned from a variety of species, and some of them have been genetically characterized 9, 13, 14, 15. In higher plants, biosynthesis of carotenoids takes place in the plastid. However, all currently known enzymes of the pathway are encoded by nuclear genes, and then imported into the plastid 16. During carotenoid biosynthesis (Figure 4), the first rate-limiting step is the condensation of two molecules of geranylgeranyl pyrophosphate into the colorless phytoene catalyzed by phytoene synthase. Phytoene undergoes four desaturation reactions to form the pink pigment lycopene, catalyzed by two enzymes, phytoene desaturase (PDS) and ζ-carotene desaturase (ZDS) 15. In addition to PDS and ZDS, other components, such as Phytoene Desaturation 1 (PDS1, a p-hydroxyphenylpyruvate dioxygenase) 17 and PDS2, are also involved in the desaturation reactions 18. Lycopene is a branch point for two separate pathways. In one branch, lycopene undergoes two sequential cyclization reactions to form δ- and α-carotene, followed by the conversion into lutein as the end product. In a second branch, a series of reactions, including cyclization and hydroxylation, result in the production of γ-carotene, β-carotene and zeaxanthin, respectively 19, 20. Epoxidation of zeaxanthin leads to the formation of violaxanthin that can be converted into neoxanthin, the end product of the branch in the plastid. Finally, cleavage of violaxanthin and neoxanthin, followed by further modifications, produces ABA in the cytoplasm 21.
The biosynthesis pathway of carotenoids in Arabidopsis (adapted from 71).
Despite a wealth of knowledge on the carotenoid biochemistry, only a limited number of mutants in the carotenoid biosynthesis pathway have been characterized in higher plants. In Arabidopsis, several lutein-deficient mutants were identified by a specific genetic screen 20. Among these loci, CCR1 and CCR2/CRTISO are required in carotenoid desaturation by the isomerization of poly-cis-carotenoids to all-trans-carotenoids, which are involved in the formation of prolamellar bodies and normal photomorphogenesis 22. Downstream of this step of the pathway, LUT1 and LUT2, which encode ε-cyclase and ε-hydroxylase, respectively, are essential for lutein biosynthesis 20, 23. In sunflower, the nondormant-1 (nd-1) mutant, obtained by in vitro tissue culture, was characterized as an albino mutant showing defects in carotenoid biosynthesis 24, and its wild type (WT) allele likely encodes a ZDS 25.
In general, most carotenoid-deficient mutants of higher plants show abnormal chloroplast development and bleaching or variegated leaves. These developmental defects are partly attributed to the loss of or reduced carotenoid-mediated antioxidant activity, impaired chlorophyll biosynthesis and reduced PS II capacity 4, 26, 27, 28. Moreover, the ABA-deficient phenotype is often associated with precocious seed germination (viviparous), reduced seed dormancy and compromised stress response 10. Interestingly, in carotenoid-deficient mutant plants, the expression of many nuclear-encoded photosynthetic genes, including the Lhc gene family (light-harvesting chlorophyll a/b-binding protein) and the small subunit of ribulose-1,5-bisphosphate carboxylase/oxygenase (RbcS), is greatly reduced in the absence of functional chloroplasts 29, 30. The reduced expression of these genes does not appear to be correlated with phytochrome content and activity 31, suggesting that a signal from plastids is required for the light induction of these nuclear genes. This regulatory mechanism, known as retrograde signaling, has been shown to involve the transduction of a signal from the plastid to nucleus 32. Notably, the plastid-generated signals were shown to be chlorophyll biosynthetic precursors, chloroplast redox states and ROS 32, 33, 34, 35, 36, all of which are related to carotenoid metabolism. Therefore, carotenoids are involved in the regulation of a large repertoire of physiological processes in higher plants.
In this paper, we report the characterization of the Arabidopsis ZDS gene. We found that the Arabidopsis Spontaneous Cell Death1 (SPC1) gene encodes a ZDS. Mutations in the single-copied SPC1/ZDS gene cause impaired carotenoid biosynthesis, leading to severe developmental defects, impaired retrograde signaling, ABA-deficient phenotype and spontaneous cell death in the mutant plants. These results demonstrate that ZDS is essential for carotenoid biosynthesis, thereby playing a critical role in plant growth and development.
Materials and Methods
Plant materials and growth conditions
A collection of T-DNA-mutagenized Arabidopsis thaliana (ecotype Wassilewskija or WS) lines was generated by transformation of the pga22 mutant plants 37 with a binary vector pTA231 as described 38. The spc1-1 mutant was identified by screening of approximately 2 500 lines for lesions of spontaneous cell death on leaves. The putative spc1-1 mutant was outcrossed with WS twice to segregate out pga22 and other unrelated mutations. The pTA231 vector carries a bialaphos selection marker gene (Bar), and the spc1-1 mutant shows a Bar-resistant phenotype. The spc1-2 mutant (SALK_033039) was obtained from the Arabidopsis Biological Resources Center (ABRC) 39.
Unless indicated otherwise, seeds were germinated and grown under continuous white light (120 μmol m−2 s−1) at 22 °C on the MS medium 40 containing 3% sucrose and 0.8% agar or in soil. Transformation of plants was carried out by vacuum infiltration as described 41.
Detection of cell death and superoxide
Superoxide accumulation in leaves was visualized by staining with 0.1% nitroblue tetrazolium (NBT) according to a previous report 42. Cell death was examined by Evans blue staining as described 43 with minor modifications. Briefly, samples were vacuum-infiltrated in a 0.1% Evans blue (w/v) solution for 5 min and then stained at room temperature for 3 to 5 h. After the staining, samples were washed with a phosphate-buffered saline containing 0.05% (v/v) Tween-20.
Analysis of carotenoids and chlorophylls
Sample preparation, analysis of carotenoids and chlorophylls by high-performance liquid chromatography (HPLC) were performed essentially according to a previous report 44. The samples were subjected to HPLC analysis using a reverse-phase C30, 5 μm column (250 × 4.6 mm2) coupled to a 20 × 4.6 mm C30 guard column (YMC Inc., Wilmington, NC, USA). Carotenoids were identified by their characteristic absorption spectra and typical retention time as well as in comparison with authentic standards (purchased from Sigma, Hong Kong SAR, China).
Total chlorophylls were prepared from leaves by acetone extraction and quantatively analyzed as described 45.
Chlorophyll fluorescence measurements
Chlorophyll fluorescence was measured essentially as previously described 46, 47 using a PAM-2000 portable chlorophyll fluorometer (Heinz Walz, Effieltrich, Germany). The maximal photochemical efficiency of PS II was calculated from the ratio of variable (Fv) to maximum (Fm) fluorescence by the formula Fv/Fm=(Fm-Fo)/Fm.
Transmission electron microscopy
Samples were fixed, post-fixed and then embedded in Spurr's resin (Sigma) as described by Mou et al. 48. Ultrathin sections (50-60 nm) were stained with lead citrate and uranyl acetate, and then observed under a transmission electron microscope (Model JEM-1230, JEOL).
Molecular manipulations
A 5.5-kb SPC1 genomic clone was obtained by polymerase chain reaction (PCR) using KOD plus DNA polymerase (Toyobo Biochemicals, Japan) and primers SPC1F3 (5′-CGC TAG CGT CAA CCA AAT AGG TGG GTG AAG AGC-3′) and SPC1B3 (5′-GGG CGC GCC GTA AGA TTC AAA GCG GGA AGT GTG G-3′). The SPC1 genomic clone included a 1.6-kb promoter sequence upstream of the 5′ untranslated region (UTR) of At3g04870. The PCR product, digested with NheI and AscI (both sites were included in the primers), was cloned into the SpeI/AscI-digested pER14 vector that was derived from pER8 49. The resulting construct was transformed into the Agrobacterium tumefaciens strain GV3101, which was then used for the transformation of spc1-1/- and spc1-2/+ plants.
Genotyping of spc1 mutants was carried out by PCR with three primers. Two primers were specific to genomic sequences flanking the respective T-DNA insertion sites. A third primer was specific to the T-DNA, which, in combination with an Arabidopsis sequence-specific primer, should produce a fusion PCR fragment of the T-DNA and the Arabidopsis genomic sequence. This fusion DNA fragment had a different size from that of the Arabidopsis genomic PCR fragments.
Northern blot analysis and real-time PCR analysis were performed as previously described 37, 50.
Primers used in the PCR analysis (all sequences are from 5′- to 3′- end) were:
GTAIL4 (for spc1-1): ATC GCC CTT CCC AAC AGT TGC; SPC1F1 (for spc1-1): ATC AAT CCA AAT CCG CTT CAC; SPC1B1 (for spc1-1): CGA AGA CGA CTG AAG AAG CCA T; LBa1 (for spc1-2): TGG TTC ACG TAG TGG GCC ATC G; SPC1F2 (for spc1-2): CCT CTG CGT CAG GTT GAT TCC; SPC1B2 (for spc1-2): CAG CAC CCA CCT ACA TCA CCA; SPC1qRTF: CCG ACA TGA GTG TTA ACG CTC CA; SPC1qRTB: CCA CGT CTA TCT ACA AAA GAA CCC AC; ACTIN7F: GGA ACT GGA ATG GTG AAG GCT G; ACTIN7B: CGA TTG GAT ACT TCA GAG TGA GGA.
Results
Identification and genetic characterization of the spc1 mutants
The Arabidopsis spc1-1 mutant was identified in a genetic screen for mutants with spontaneous cell death lesions. Under the normal growth condition, spc1-1 had normal cotyledons and roots, but developed chlorotic lesions in true leaves (Figure 1A and 1B). The spc1-1 leaves were often yellowish with bleached spots, but appeared morphologically normal. Moreover, inflorescences, floral organs and siliques also became yellow, accompanying the formation of chlorosis (Figure 1C–1E). In later developmental stages, some yellowish leaves could turn into light green. Matured spc1-1 plants were slightly smaller than WT plants (Figure 1F) and were able to set seeds with reduced fertility.
The spc1 mutant phenotype. (A) A two-week-old WT (WS) seedling. (B) A two-week-old spc1-1 seedling. (C) Inflorescences of WS (left) and spc1-1 (right) plants. (D) Flowers of WS (left) and spc1-1 (right). (E) Siliques of WS (left) and spc1-1 (right). (F) Mature plants of WS (left) and spc1-1 (right). (G) Seven-day-old WT (Col-0; right) and spc1-2 (left) seedlings. (H) An eight-week-old spc1-2 plant germinated and grown on MS medium. All plants were grown under continuous white light. Bar: (A-G) 1 mm and (H) 5 mm.
The spc1-1 mutant was in the WS background. In a cross between spc1-1 and WT WS plants, all F1 progeny (n=27) showed a normal growth phenotype, indicating that the mutation was recessive. In F2 progeny obtained from self-pollinated F1 plants, the mutant phenotype segregated in a 3:1 ratio (WT: spc1-1=236:67, χ2=0.67), suggesting that spc1-1 was a recessive mutation in a single nuclear locus.
Upon characterization of the molecular lesion of spc1-1 (see below), we identified a second mutant allele spc1-2 from the ABRC. The spc1-2 mutant showed a severely stunted phenotype, with dark pink or purple cotyledons (Figure 1G). Under the tissue culture condition, growth and development of spc1-2 were eventually arrested shortly after germination. Occasionally, the mutant could produce bleached leaves and inflorescences (Figure 1H). The spc1-2 mutant was not viable when grown in soil. Because of the seedling-lethal phenotype of spc1-2, most studies described below were carried out by using the weak mutant allele spc1-1.
The spontaneous cell death phenotype of spc1-1
To assay the spontaneous cell death phenotype in spc1-1, we first stained leaves of the mutant and WT with Evans blue, a dye for the specific detection of dead or dying cells. A mosaic pattern of dead cells was observed in spc1-1 leaves, whereas no apparent positive staining was found in WT leaves (Figure 2A). Spontaneous cell death in plants is often accompanied by the generation of ROS. We then examined the accumulation of superoxide, a major species of ROS, by staining leaves with NBT. Under normal conditions, no superoxide was detected in WT leaves under the assay conditions. However, abundant superoxide was detected in spc1-1 leaves (Figure 2B). These results suggest that spc1-1 displays a spontaneous cell death phenotype in the absence of any biotic or abiotic stress.
The cell death phenotype of spc1-1. (A) Leaves of WS and spc1-1 grown under different conditions were stained with Evans blue. Dead cells were stained as blue. SD: 12/12 h light/dark; WL: continuous white light. Comp 1: leaves of the spc1-1 mutant carrying a wild-type SPC1 transgene. WS and Comp 1 leaves shown in the panel were grown in WL, and similar phenotype was obtained when SD-grown leaves were tested. (B) Leaves of WS and spc1-1 grown under different conditions were stained with NBT. Superoxide was stained as blue precipitates. See (A) for other technical details. (C) Two-week-old WS (left) and spc1-1 (right) seedlings germinated and grown under the short-day (SD; 12/12 h light/dark) condition. (D) Two-week-old WS (left) and spc1-1 (right) seedlings germinated and grown under continuous white light (WL).
The penetrance of the spc1-1 mutant phenotype appears to be light dependent. When grown under the short-day condition (12 h light/12 h dark), the chlorotic phenotype of spc1-1 was compromised (Figure 2C). However, when grown in the continuous white light, the spc1-1 mutant plants showed a stronger phenotype (Figure 2D). Similar results were also observed when WT and spc1-1 plants were grown under different light intensities (data not shown). As expected, more apparent cell death was observed in spc1-1 leaves grown under continuous white light compared with those from the short-day condition (Figure 2A). Consistent with these observations, spc1-1 plants accumulated a higher level of superoxide under continuous white light compared with those from the short-day condition (Figure 2B).
Molecular characterization of the SPC1 gene
The spc1-1 mutant was recovered from a T-DNA insertional population 38. Segregation analysis suggested that the spc1-1 mutant genome contained a single T-DNA insertion, which was tightly linked with the mutant phenotype. Using thermal asymmetric interlaced-PCR 51, a T-DNA insertion was found about 400 base pairs (bp) upstream of the putative ATG of At3g04870 of chromosome III (Figure 3A). Northern blot analysis showed that the expression of At3g04870 was reduced in spc1-1 plants (Figure 3B). PCR analysis indicated that the open reading frame (ORF) of At3g04870 remained intact. These data suggest that At3g04870 has partial function in spc1-1 plants.
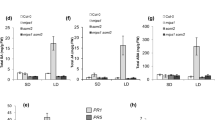
Molecular cloning of the SPC1 gene. (A) A schematic diagram showing the structure of SPC1. Filled box and solid lines indicate exons and introns, respectively. Dash lines denote untranscribed and untranslated regions. The insertion sites of the left border are shown by arrows and the insertion sites of the right border are unclear as indicated by question marks. (B) A northern blot analysis of SPC1 expression. Total RNA was prepared from 3-week-old seedlings of WT and spc1 mutant plants. Twenty micrograms of RNA were used for the northern blot analysis using a full-length SPC1 cDNA as the probe. Two cryptic transcripts in spc1-2 were indicated by arrows. (C) Molecular complementation of spc1-1. Comp 1: the spc1-1 mutant carrying a wild-type SPC1 transgene. (D) Molecular complementation of spc1-2. Comp 2: the spc1-2 mutant carrying a wild-type SPC1 transgene. The transformation was carried out using spc1-2/+ heterozygous plants, and T2 progeny showing WT phenotype and homozygous for the spc1-2 mutation was identified and confirmed by PCR (see Materials and Methods for details). (E) Allelism test of spc1-1 and spc1-2. F1 plants derived from a cross between spc1-1/- and spc1-2/+ were germinated and grown on MS medium under continuous white light. Arrows denote the spc1-1/spc1-2 F1 plants whose genotype was confirmed by PCR analysis. (F) Silique collected from WT (top) and spc1-2/+ (bottom) plants. Approximately one-fourth albino embryos/seeds (indicated by arrows) were found in the developing siliques of spc1-2/+ plants. (G) Real-time quantative RT-PCR analysis of expression of SPC1 in different tissues/organs of WT plants. Actin7 was used as an internal control and relative expression level of SPC1 was shown as percentage relative to that of Actin7. Mean values obtained from two independent experiments are shown in the histogram. Bars denote SE. All plants were grown under continuous white light. Bar: 1 mm.
To test whether At3g04870 represents SPC1, we carried out a molecular complementation experiment. A 5.5-kilo base pairs (kb) WT genomic DNA fragment, which encompassed the promoter region, 5′-UTR, the coding sequence region, 3′-UTR and 3′-untranscribed sequences of At3g04870, was cloned into a binary vector pER14 (a derivative of pER8; see 49). The resulting construct was transformed into spc1-1 plants. In all tested seven transgenic lines, the transgene was able to fully rescue the spc1-1 mutant phenotype (Figures 2A, 2B and 3C), demonstrating that At3g04870 indeed represents SPC1.
Data presented above suggest that spc1-1 is a weak mutant allele. To gain more insight into SPC1 function, we identified an additional mutant allele SALK_033039. In this mutant, a T-DNA was found to insert in exon 2 of SPC1 (Figure 3A). To confirm the allelism between these two mutants, we crossed spc1-1 homozygous plants with SALK_033039 plants heterozygous for the T-DNA insertion in SPC1. In F1 progeny, although approximate 50% plants (n=22) showed a WT phenotype, another half of plants (n=21) displayed an spc1-1-like phenotype (Figure 3E), consistent with the prediction that the F1 plants should show the phenotype of the weak allele. These results suggest that these two mutants are indeed allelic. Accordingly, this mutant was named as spc1-2. An SPC1 transgene was also able to fully rescue the spc1-2 mutant phenotype (Figure 3D), demonstrating that the observed mutant phenotype was caused by the disruption of SPC1 by the T-DNA insertion.
A northern blot analysis revealed that the SPC1 transcript with predicted size was absent in the spc1-2 mutant. Instead, two cryptic transcripts were detected in the mutant (Figure 3B). Although we could not confirm that spc1-2 was a null mutant allele, the above results suggested that spc1-2 was a stronger mutant allele. Consistent with this notion, spc1-2 displayed a more severe phenotype compared with spc1-1 (Figures 1 and 3). The heterozygous spc1-2 plants had no detectable phenotypes when grown under normal conditions. However, approximately one quarter (115 out of 494 examined embryos; 23.3%) developing embryos/young seeds showed an albino phenotype in siliques of spc1-2 heterozygous plants (Figure 3F), indicating that embryo development is also affected by the spc1-2 mutation.
Quantative real-time RT-PCR analysis revealed the highest SPC1 expression level in leaves, followed by flowers and siliques and a lower level in roots and stems (Figure 3G). This expression pattern is consistent with the mutant phenotype, of which leaves showed a stronger phenotype than other organs/tissues.
SPC1 encodes a ZDS in the carotenoid biosynthesis pathway
SPC1 is a single-copied gene in the Arabidopsis genome. Comparison of the genomic sequence and a cDNA sequence (Accession No. U38550) revealed that SPC1 contained 13 exons separated by 12 introns (Figure 3A). An ORF in the SPC1 gene encodes a protein of 558 amino-acid residues. Sequence comparison suggests that SPC1 encodes a putative ZDS or carotene 7, 8-desaturase (ZDS; EC 1.14.99.30), which acts to convert ζ-carotene into neurosporene, a key step in the biosynthesis of carotenoids (Figure 4). Similar to other enzymes in the carotenoid biosynthesis pathway, SPC1/ZDS appears to be highly conserved in both lower and higher plants.
Because SPC1 is a single-copied gene in Arabidopsis, one would expect that the spc1 mutations should cause impaired carotenoid biosynthesis. To test this hypothesis, we measured and compared several major species of carotenoids in WT and spc1-1 plants. Total carotenoids prepared from WT and spc1-1 plants were subjected to HPLC. In WT plants, ζ-carotene, the putative substrate of SPC1, and its precursor phytoene (Figure 4) were undetectable under the assay conditions. In spc1-1 plants, these two compounds were substantially increased (Figure 5A, 5B and 5D), suggesting that the carotenoid biosynthesis pathway is blocked by the mutation. Furthermore, several compounds downstream of ZDS were reduced, including lutein and β-carotene (Figure 5A–5C). Along with a previous study showing that a ZDS recombinant protein expressed in bacteria was enzymatically active 52, we conclude that SPC1 is a functional ZDS, and that mutations in SPC1 disrupt carotenoid biosynthesis in Arabidopsis.
Measurement of pigments in WT and spc1-1 plants. Total pigments were extracted from WT and spc1-1 plants and were subjected to HPLC analysis as described in Materials and Methods. (A) HPLC chromatogram recorded at 287 nm. (B) HPLC chromatogram recorded at 430 nm. (C) HPLC chromatogram recorded at 460 nm. (D) Derived spectra of phytoene and ζ-carotene. (E) Quantative analysis of chlorophyll a (Chl a) and chlorophyll b (Chl b). Peaks in (A) through (C) are (1) 15-cis-phytoene, (2) chlorophyll b, (3) α-carotene, (4) chlorophyll a, (5) β-carotene and (6) ζ-carotene.
Reduced PS II activity in spc1-1
Carotenoid deficiency often causes a reduced PS II activity 4. To examine the possible effects of the spc1-1 mutation on the PS II activity, we carried out a chlorophyll fluorescence imaging analysis, of which the ratio of variable fluorescence to maximum fluorescence (Fv/Fm) represents the maximum potential capacity of the PS II photochemical reactions 53. In spc1-1, maximal photochemical efficiency Fv/Fm is decreased (Figure 6), indicating that the mutation causes substantial photoinhibition in the mutant.
Chlorophyll fluorescence imaging of WT and spc1-1. Two-week-old seedlings of WT and spc1-1 were dark-adapted for 30 min before the measurements. The minimum fluorescence yield (Fo) was measured under the measuring light (650 nm) with very low intensity (0.8 μmol m−2 s−1), and the maximum fluorescence yield (Fm) was measured after a saturating pulse of white light (3 000 μmol m−2 s−1 for 1 s). The experiment was repeated three times. The mean values and the SE are given at the bottom of each image.
Defective chloroplast development in spc1-1
The bleaching leaf phenotype of the spc1 mutants suggests possible defects in chloroplast development. We therefore conducted a transmission electron microscopic analysis of the spc1 mutants in comparison with WT plants. In WT mesophyll cells, chloroplasts were well developed and organized (Figure 7A, 7B, 7E and 7F). By contrast, in spc1-1 mesophyll cells derived from the short-day-grown plants, chloroplasts were misshapen, and the thylakoid membranes were reduced and irregularly organized (Figure 7C and 7D). When grown under continuous white light, the spc1-1 mesophyll cells contained chloroplast-like structures, of which no distinctive thylakoid membranes could be observed (Figure 7G and 7H). In spc1-2, no plastid- or chloroplast-like structures were found under all tested conditions (data not shown), indicating that the mutation completely disrupted chloroplast development.
Abnormal chloroplast development in spc1-1. Leaves collected from WT and spc1-1 plants grown under different conditions as indicated at the right-hand side of the panels (SD: 12/12 h light/dark; WL: continuous white light). Ultrathin sections were prepared and analyzed by transmission electron microscopy. (A, C, E and G) An overview of mesophyll cells. Bar: 10 μm. (B, D, F and H) Enlarged views of chloroplast. In (F), s: starch granule. Bar: 1 μm.
Chlorophyll biosynthesis is downregulated in spc1
Because the spc1 mutations cause bleached leaves, it is likely that chlorophyll biosynthesis is also affected in the mutants. Moreover, carotenoid deficiency has been reported to cause sharply reduced levels of chlorophyll 54, 55. We therefore measured the chlorophyll levels in the leaves of WT, spc1-1 and spc1-2 plants. Levels of both chlorophylls a and b were reduced in spc1-1 plants, to about 45% and 50% relative to those in WT plants, respectively (Figure 5E). In the stronger spc1-2 mutant, chlorophylls were not detectable under our assay condition.
To explore possible mechanism of reduced chlorophyll levels in the spc1 mutants, we examined the expression of two key genes in the chlorophyll biosynthesis pathway, protochlorophyllide oxidoreductase B (PORB) and chlorophyllide a oxygenase (CAO), in the mutant plants. POR catalyzes the light-dependent reduction of protochlorophyllide a to chlorophyllide a, which is subsequently converted into chlorophyllide b by CAO 56, 57, 58, 59. The Arabidopsis genome has a single-copied CAO gene and three structurally related POR genes (A, B and C), of which only PORB is expressed throughout most developmental stages 56, 58, 59, 60. We found that expression of both PORB and CAO was downregulated in spc1-1 mutant plants and not detectable in spc1-2 (Figure 8A and data not shown), which likely accounts for the reduced chlorophyll level in the mutants.
Impaired retrograde signaling in spc1
The decreased expression of the nuclear genes PORB and CAO in spc1 mutants suggests the involvement of a plastid-to-nucleus retrograde signaling mechanism. To further test this possibility, we analyzed the expression of Lhcb1.1 (also known as CAB2), Lhcb1.4 and RbcS, which have been demonstrated as a class of nuclear genes regulated by retrograde signaling 32. In spc1-1, expression of these genes had no substantial reduction. However, essentially no expression of these genes was detected in the stronger mutant spc1-2 (Figure 8B). Collectively, the observation that expression of several nuclear genes, including PORB, CAO, Lhcb1.1, Lhcb1.4 and RbcS, is downregulated by the spc1 mutations suggests that spc1 mutations impair the plastid-to-nucleus retrograde signaling. In addition, these results also imply that PORB and CAO may be more sensitive to alterations of retrograde signaling.
ABA-deficient phenotype in spc1-1
The major biosynthetic pathway for ABA is dependent on carotenoids, of which violaxanthin and neoxanthin are critical precursors of ABA biosynthesis (Figure 4). To test if the spc1-1 mutant has an ABA-deficient phenotype, we carried out the following experiments. When germinated in MS medium, low concentrations of ABA were able to partially rescue the spc1-1 mutant phenotype (Figure 9A). Because ABA regulates a number of abiotic stress responses, ABA-deficient mutants are often hypersensitive to adverse growth conditions. When placed in the ambient condition with cut roots, spc1-1 seedlings had more rapid water loss than WT, thus showing less tolerance to dehydration (Figure 9B and 9C). These results suggest that the spc1-1 mutation caused an ABA-deficient phenotype.
The ABA-deficiency phenotype of spc1-1. (A) Three-week-old WT (WS) and spc1-1 seedlings germinated and grown on the MS medium in the absence (MS) or presence of 0.5 mM ABA (ABA) as indicated on the top of the panel. (B) Three-week-old WT (WS) and spc1-1 seedlings germinated and grown on the MS medium were removed from the medium and their roots were cut off. The aerial parts were then transferred into a Petri dish. Photos were taken before the cutting (0 min) and 20 min after the cutting. (C) Quantative analysis of water loss in WT (WS) and spc1-1 plants. The experiment was repeated three times, and mean values are given in the histogram. Bars indicate SE. At least 30-40 seedlings were used in each experiment. Bar: 2 mm.
Discussion
Carotenoids are a class of compounds that are synthesized de novo in the plastid, and have distinctive functions in a variety of physiological processes. In addition to their role for color decoration, carotenoids function as important accessory pigments in the light-harvesting complex, the redox intermediates of PS II and antioxidants against the photooxidative stress. Moreover, carotenoids are precursors of the de novo synthesized ABA. Thus, impaired biosynthesis of carotenoids is expected to have a broad spectrum of effects on plant growth and development. The spc1 mutants show a typical carotenoid-deficient syndrome, as characterized by bleached leaves, excessive production of superoxide, abnormal chloroplast development, reduced PS II activity and an ABA-deficient phenotype, suggesting that the mutations may affect carotenoid metabolism. SPC1 was found to encode a putative ZDS, a key enzyme in carotenoid biosynthesis catalyzing the desaturation of ζ-carotene to produce neurosporene. Biochemical analysis revealed that although carotenoids were not detectable in the strong mutant allele spc1-2, substantially altered levels of these compounds were found in the weak mutant allele spc1-1. Notably, the spc1-1 mutation causes the accumulation of a higher level of 15-cis-phytoene and ζ-carotene, the direct substrate of ZDS. The developmental and biochemical phenotype of spc1 is similar to that of a sunflower mutant nd-1, whose WT allele likely encodes a ZDS 24, 25. These results suggest that SPC1 acts downstream of ζ-carotene in the carotenoid biosynthesis pathway. Consistent with this notion, SPC1/ZDS expressed in Escherichia coli cells is able to catalyze the formation of lycopene in a light-dependent manner 52. These results indicate that SPC1 is a functional ZDS, and that mutations in SPC1 cause impaired biosynthesis of carotenoids.
SPC1/ZDS is a single-copied gene in the Arabidopsis genome. Considering the physiological importance of carotenoids, it would not be surprising to find that the strong mutant allele spc1-2 shows a seedling-lethal phenotype. Analysis of the non-lethal weak mutant allele spc1-1 has allowed us to gain more insight into the physiological function of carotenoids. Many species of carotenoids have been shown to act as antioxidants to quench excessive free radicals and ROS generated from photooxidation 61, 62. Because of the substantial loss of carotenoids, spc1-1 has accumulated a higher level of superoxide, which may, at least in part, contribute to the spontaneous cell death phenotype of the mutant. Similar to other carotenoid-deficient mutants 54, the penetrance of spc1-1 mutant phenotype is light dependent, with a stronger mutant phenotype under the high light condition. This light-dependent phenotype is presumably a consequence of the accumulation of more ROS under high light that cannot be efficiently removed owing to a lower level of carotenoid-type antioxidants. The abnormal development of the chloroplast in spc1-1 is thus likely caused by the photooxidative damage, which also appears to be light dependent.
The spc1-1 mutant was originally identified from a screen for mutants showing a spontaneous cell death phenotype. The accumulation of superoxide and other possible ROS, due to a reduced level of the carotenoid-type antioxidants, may be a major casual factor for the cell death phenotype in spc1. Notably, β-carotene, a major antioxidant for photooxidation protection in plants, was remarkably reduced in spc1-1, which may contribute in part to the spontaneous cell death phenotype of the mutant. It is interesting to notice that spc1-1 has a lower level of chlorophylls a and b, which appears to be correlated to the reduced expression of PORB and CAO. The impaired chlorophyll metabolism and abnormal chloroplast development have been attributed as an important mechanism to trigger spontaneous cell death in a number of mutants, including Arabidopsis-accelerated cell death2 (acd2) 63, 64, acd1 65, 66, 67 and maize lethal leaf spot-1 (lls1) 68, 69. ACD1 and LLS1 are believed to be ortholog genes in Arabidopsis and maize 65, 66, 69. These three genes, all encoding key enzymes for chlorophyll degradation, presumably function to prevent the formation of ROS or free radicals generated from photooxidation, thereby negatively regulating cell death 63, 64, 69. In spc1, however, the reduced chlorophyll level may not be a direct casual factor for the cell death phenotype, since spc1 and other three mutations (acd1, acd2 and lls1) appear to involve different biochemical mechanisms.
Although most plastid proteins are encoded by nuclear genes, plastid-derived signals, however, can also regulate expression of nuclear genes encoding plastid-localized proteins. The latter mechanism, termed as plastid-to-nucleus retrograde signaling, was initially appreciated in two carotenoid-deficiency mutants of barley 32. Expression of Lhcb1.1, Lhcb1.4 and RbcS appears to be unaltered in spc1-1, but is undetectable in the stronger mutant allele spc1-2. Interestingly, expression of CAO and PORB is downregulated in both mutant alleles, suggesting that these two genes are more sensitive to an altered retrograde signaling. The increased sensitivity of CAO and PORB to a plastid signal may be attributable to the fact that these two key genes for chlorophyll biosynthesis act in a linear pathway downstream of Mg-Protoporphyrin IX, the major signal molecule of retrograde signaling and the first intermediate in the chlorophyll branch of the tetrapyrrole biosynthetic pathway 32. Consistent with this notion, overexpression of PORA and PORB results in a genomes uncoupled-like mutant phenotype 70.
In summary, we have characterized the SPC1 gene that encodes a ZDS in the de novo carotenoids biosynthesis pathway in Arabidopsis. The spc1 mutations cause typical carotenoid-deficiency phenotype, including abnormal chloroplast development, reduced photoprotection, reduced PS II activity, ABA-deficient phenotype and impaired retrograde signaling. Collectively, these defects may trigger spontaneous cell death in spc1. The identification of the spc1 mutant alleles provides useful materials for more comprehensive studies on the physiological function of carotenoids by genetic, biochemical and molecular approaches.
Accession codes
References
Demmig-Adams B, Gilmore AM, Adams WW III . In vivo functions of carotenoids in higher plants. FASEB J 1996; 10:403–412.
Niyogi KK . Photoprotection revisited: genetic and molecular approaches. Annu Rev Plant Physiol Plant Mol Biol 1999; 50:333–359.
Yamamoto HY, Bugos RC, Hieber AD . Biochemistry and molecular biology of the xanthophyll cycle. In: Frank HA, Young AJ, Britton G, Cogdell RJ, eds. The photochemistry of carotenoids. Dordrecht, The Netherlands: Kluwer Academic Publishers, 1999:293–303.
Tracewell CA, Vrettos JS, Bautista JA, Frank HA, Brudvig GW . Carotenoid photooxidation in photosystem II. Arch Biochem Biophys 2001; 385:61–69.
Cuttriss AJ, Pogson BJ . Carotenoids. In: Davies KM, ed. Plant pigments and their manipulation. Boca Raton, FL, USA: CRC Press, 2004:57–91.
Pogson BJ, Rissler HM, Frank HA . The roles of carotenoids in energy quenching. In: Wydrzynski T, Satoh K, eds. Photosystem II: the water/plastoquinone oxidoreductase in photosynthesis. Dordrecht, The Netherlands: Springer 2006:515–537.
Siefermann-Harms D . The light harvesting protective functions of carotenoids in photosynthetic membranes. Physiol Plant 1987; 69:561–568.
Fraser PD, Bramley PM . The biosynthesis and nutritional uses of carotenoids. Prog Lipid Res 2004; 43:228–265.
Hirschberg J . Carotenoid biosynthesis in flowering plants. Curr Opin Plant Biol 2001; 4:210–218.
McCarty DR . Genetic control and integration of maturation and germination pathways in seed development. Annu Rev Plant Physiol Plant Mol Biol 1995; 46:71–93.tonn
Walton DC, Li Y . Abscisic acid biosynthesis and metabolism. In: Davies PJ, ed. Plant hormones. Dordrecht, The Netherlands: Kluwer Academic Publishers, 1995:140–157.
Zeevaart JAD, Creelman RA . Metabolism and physiology of abscisic acid. Annu Rev Plant Physiol Plant Mol Biol 1988; 39:439–473.
Armstrong GA, Hearst JE . Carotenoids 2: genetics and molecular biology of carotenoid pigment biosynthesis. FASEB J 1996; 10:228–237.
Cunningham FX, Gantt E . Genes and enzymes of carotenoid biosynthesis in plants. Annu Rev Plant Physiol Plant Mol Biol 1998; 49:557–583.
Sandmann G . Carotenoid biosynthesis and biotechnological application. Arch Biochem Biophys 2001; 385:4–12.
Bartley GE, Scolnik PA . Molecular biology of carotenoid biosynthesis in plants. Annu Rev Plant Physiol Plant Mol Biol 1994; 45:287–301.
Norris SR, Shen X, DellaPenna D . Complementation of the Arabidopsis pds1 mutation with the gene encoding p-hydroxyphenylpyruvate dioxygenase. Plant Physiol 1998; 117:1317–1323.
Norris SR, Barrette TR, DellaPenna D . Genetic dissection of carotenoid synthesis in Arabidopsis defines plastoquinone as an essential component of phytoene desaturation. Plant Cell 1995; 7:2139–2149.
Cunningham FX Jr, Pogson B, Sun Z, et al. Functional analysis of the β- and ε-lycopene cyclase enzymes of Arabidopsis reveals a mechanism for control of cyclic carotenoid formation. Plant Cell 1996; 8:1613–1626.
Pogson B, McDonald KA, Truong M, Britton G, DellaPenna D . Arabidopsis carotenoid mutants demonstrate that lutein is not essential for photosynthesis in higher plants. Plant Cell 1996; 8:1627–1639.
Nambara E, Marion-Poll A . Abscisic acid biosynthesis and catabolism. Annu Rev Plant Biol 2005; 56:165–185.
Park H, Kreunen SS, Cuttriss AJ, DellaPenna D, Pogson BJ . Identification of the carotenoid isomerase provides insight into carotenoid biosynthesis, prolamellar body formation, and photomorphogenesis. Plant Cell 2002; 14:321–332.
Pogson BJ, Rissler HM . Genetic manipulation of carotenoid biosynthesis and photoprotection. Philos Trans R Soc London 2000; 355:1395–1403.
Fambrini M, Pugliesi C, Vernieri P, Giuliano G, Baroncelli S . Characterization of a sunflower (Helianthus annuus L.) mutant deficient in carotenoid synthesis and abscisic-acid content induced by in vitro tissue culture. Theor Appl Genet 1993; 87:65–69.
Conti A, Pancaldi S, Fambrini M, et al. A deficiency at the gene coding for ζ-carotene desaturase characterizes the sunflower non dormant-1 mutant. Plant Cell Physiol 2004; 45:445–455.
Lee BM, Pack NC, Magill CW, Smith JD . Classification and characterization of the viviparous mutants of maize (Zea mays L.). Maydica 1997; 42:371–378.
Carol P, Stevenson D, Bisanz C, et al. Mutations in the Arabidopsis gene IMMUTANS cause a variegated phenotype by inactivating a chloroplast terminal oxidase associated with phytoene desaturation. Plant Cell 1999; 11:57–68.
Wu D, Wright DA, Wetzel C, Voytas DF, Rodermel S . The IMMUTANS variegation locus of Arabidopsis defines a mitochondrial alternative oxidase homolog that functions during early chloroplast biogenesis. Plant Cell 1999; 11:43–55.
Hess WR, Muller A, Nagy F, Borner T . Ribosome-deficient plastids affect transcription of light-induced nuclear genes: genetic evidence for a plastid-derived signal. Mol Gen Genet 1994; 242:305–312.
Rocca NL, Barbato R, Vecchia FD, Rascio N . CAB gene expression in bleached leaves of carotenoid-deficient maize. Photosynthesis Res 2000; 64:119–126.
Hess W, Schendel R, Borner T, Rudiger W . Reduction of mRNA level for two nuclear encoded light regulated genes in the barley mutant albostrians is not correlated with phytochrome content and activity. J Plant Physiol 1991; 138:292–298.
Nott A, Jung HS, Koussevitzky S, Chory J . Plastid-to-nucleus retrograde signaling. Ann Rev Plant Biol 2006; 57:739–759.
Karpinski S, Reynolds H, Karpinska B, et al. Systemic signaling and acclimation in response to excess excitation energy in Arabidopsis. Science 1999; 284:654–657.
Kropat J, Oster U, Rudiger W, Beck CF . Chloroplast signalling in the light induction of nuclear HSP70 genes requires the accumulation of chlorophyll precursors and their accessibility to cytoplasm/nucleus. Plant J 2000; 24:523–531.
Strand A, Asami T, Alonso J, Ecker JR, Chory J . Chloroplast to nucleus communication triggered by accumulation of Mg-protoporphyrinIX. Nature 2003; 421:79–83.
Fey V, Wagner R, Brautigam K, Pfannschmidt T . Photosynthetic redox control of nuclear gene expression. J Exp Bot 2005; 56:1491–1498.
Sun J, Niu QW, Tarkowski P, et al. The Arabidopsis AtIPT8/PGA22 gene encodes an isopentenyl transferase that is involved in de novo cytokinin biosynthesis. Plant Physiol 2003; 131:167–176.
Sun J, Hirose N, Wang X, et al. Arabidopsis SOI33/AtENT8 gene encodes a putative equilibrative nucleoside transporter that is involved in cytokinin transport in planta. J Integr Plant Biol 2005; 47:588–603.
Alonso JM, Stepanova AN, Leisse TJ, et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 2003; 301:653–657.
Murashige T, Skoog F . A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 1962; 15:473–497.
Clough SJ, Bent AF . Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 1998; 16:735–743.
Jabs T, Dietrich RA, Dangl JL . Initiation of runaway cell death in an Arabidopsis mutant by extracellular superoxide. Science 1996; 273:1853–1856.
Kim M, Ahn JW, Jin UH, et al. Activation of the programmed cell death pathway by inhibition of proteasome function in plants. J Biol Chem 2003; 278:19406–19415.
Fraser PD, Pinto ME, Holloway DE, Bramley PM . Technical advance: application of high-performance liquid chromatography with photodiode array detection to the metabolic profiling of plant isoprenoids. Plant J 2000; 24:551–558.
Lichtenthaler HK . Chlorophylls and carotenoids – pigments of photosynthetic biomembranes. Methods Enzymol 1987; 148:350–382.
Peng L, Ma J, Chi W, et al. Low PSII accumulation1 is involved in efficient assembly of photosystem II in Arabidopsis thaliana. Plant Cell 2006; 18:955–969.
Tang Y, Wen X, Lu Q, et al. Heat stress induces an aggregation of the light-harvesting complex of photosystem II in spinach plants. Plant Physiol 2007; 143:629–638.
Mou Z, He Y, Dai Y, Liu X, Li J . Deficiency in fatty acid synthase leads to premature cell death and dramatic alterations in plant morphology. Plant Cell 2000; 12:405–418.
Zuo J, Niu QW, Chua NH . Technical advance: an estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J 2000; 24:265–273.
Feng H, An F, Zhang S, et al. Light-regulated, tissue-specific, and cell differentiation-specific expression of the Arabidopsis Fe(III)-chelate reductase gene AtFRO6. Plant Physiol 2006; 140:1345–1354.
Liu YG, Mitsukawa N, Oosumi T, Whittier RF . Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J 1995; 8:457–463.
Bartley GE, Scolnik PA, Beyer P . Two Arabidopsis thaliana carotene desaturases, phytoene desaturase and ζ-carotene desaturase, expressed in Escherichia coli, catalyze a poly-cis pathway to yield pro-lycopene. Eur J Biochem/FEBS 1999; 259:396–403.
Krause GH, Weiss E . Chlorophyll fluorescence and photosynthesis: the basics. Annu Rev Plant Physiol Plant Mol Biol 1991; 42:313–349.
Wetzel CM, Jiang CZ, Meehan LJ, Voytas DF, Rodermel SR . Nuclear-organelle interactions: the immutans variegation mutant of Arabidopsis is plastid autonomous and impaired in carotenoid biosynthesis. Plant J 1994; 6:161–175.
Naested H, Holm A, Jenkins T, et al. Arabidopsis VARIEGATED 3 encodes a chloroplast-targeted, zinc-finger protein required for chloroplast and palisade cell development. J Cell Sci 2004; 117:4807–4818.
Armstrong GA, Runge S, Frick G, Sperling U, Apel K . Identification of NADPH:protochlorophyllide oxidoreductases A and B: a branched pathway for light-dependent chlorophyll biosynthesis in Arabidopsis thaliana. Plant Physiol 1995; 108:1505–1517.
Tanaka A, Ito H, Tanaka R, et al. Chlorophyll a oxygenase (CAO) is involved in chlorophyll b formation from chlorophyll a. Proc Natl Acad Sci USA 1998; 95:12719–12723.
Espineda CE, Linford AS, Devine D, Brusslan JA . The AtCAO gene, encoding chlorophyll a oxygenase, is required for chlorophyll b synthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 1999; 96:10507–10511.
Su Q, Frick G, Armstrong G, Apel K . PORC of Arabidopsis thaliana: a third light- and NADPH-dependent protochloro-phyllide oxidoreductase that is differentially regulated by light. Plant molecular biology 2001; 47:805–813.
Oosawa N, Masuda T, Awai K, et al. Identification and light-induced expression of a novel gene of NADPH-protochlorophyllide oxidoreductase isoform in Arabidopsis thaliana. FEBS Lett 2000; 474:133–136.
Hirayama O, Nakamura K, Hamada S, Kobayasi Y . Singlet oxygen quenching ability of naturally occurring carotenoids. Lipids 1994; 29:149–150.
Woodall AA, Britton G, Jackson MJ . Carotenoids and protection of phospholipids in solution or in liposomes against oxidation by peroxyl radicals: relationship between carotenoid structure and protective ability. Biochim Biophys Acta 1997; 1336:575–586.
Mach JM, Castillo AR, Hoogstraten R, Greenberg JT . The Arabidopsis accelerated cell death gene ACD2 encodes red chlorophyll catabolite reductase and suppresses the spread of disease symptoms. Proc Natl Acad Sci USA 2001; 98:771–776.
Yao N, Greenberg JT . Arabidopsis ACCELERATED CELL DEATH2 modulates programmed cell death. Plant Cell 2006; 18:397–411.
Tanaka R, Hirashima M, Satoh S, Tanaka A . The Arabidopsis-accelerated cell death gene ACD1 is involved in oxygenation of pheophorbide a: inhibition of the pheophorbide a oxygenase activity does not lead to the “stay-green” phenotype in Arabidopsis. Plant Cell Physiol 2003; 44:1266–1274.
Yang M, Wardzala E, Johal GS, Gray J . The wound-inducible Lls1 gene from maize is an orthologue of the Arabidopsis Acd1 gene, and the LLS1 protein is present in non-photosynthetic tissues. Plant Mol Biol 2004; 54:175–191.
Pruzinska A, Tanner G, Aubry S, et al. Chlorophyll breakdown in senescent Arabidopsis leaves. Characterization of chlorophyll catabolites and of chlorophyll catabolic enzymes involved in the degreening reaction. Plant Physiol 2005; 139:52–63.
Gray J, Close PS, Briggs SP, Johal GS . A novel suppressor of cell death in plants encoded by the Lls1 gene of maize. Cell 1997; 89:25–31.
Gray J, Janick-Buckner D, Buckner B, Close PS, Johal GS . Light-dependent death of maize lls1 cells is mediated by mature chloroplasts. Plant Physiol 2002; 130:1894–1907.
McCormac AC, Terry MJ . The nuclear genes Lhcb and HEMA1 are differentially sensitive to plastid signals and suggest distinct roles for the GUN1 and GUN5 plastid-signalling pathways during de-etiolation. Plant J 2004; 40:672–685.
Botella-Pavia P, Besumbes O, Phillips MA, et al. Regulation of carotenoid biosynthesis in plants: evidence for a key role of hydroxymethylbutenyl diphosphate reductase in controlling the supply of plastidial isoprenoid precursors. Plant J 2004; 40:188–199.
Acknowledgements
We thank the Arabidopsis Biological Resources Center for mutant seeds. We are grateful to Dr Sodmergen and Dr Quan Zhang (Peking University) for advice and help on transmission electron microscopy. This work is supported by grants from National Natural Science Foundation of China (Grant Nos. 30330360, 30125025 and 30221002), and Chinese Academy of Sciences (Grant No. KSCX2-YW-N-015) to Zuo J.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dong, H., Deng, Y., Mu, J. et al. The Arabidopsis Spontaneous Cell Death1 gene, encoding a ζ-carotene desaturase essential for carotenoid biosynthesis, is involved in chloroplast development, photoprotection and retrograde signalling. Cell Res 17, 458–470 (2007). https://doi.org/10.1038/cr.2007.37
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2007.37
Keywords
This article is cited by
-
Quantitative Trait Locus Mapping Combined with RNA Sequencing Reveals Candidate Genes for Chlorophyll Content in Oilseed Rape Leaves
Journal of Plant Growth Regulation (2024)
-
Molecular and biochemical characterization of rice developed through conventional integration of nDart1-0 transposon gene
Scientific Reports (2023)
-
Functional identification of ZDS gene in apple (Malus halliana) and demonstration of it's role in improving saline–alkali stress tolerance
Physiology and Molecular Biology of Plants (2023)
-
GWAS of grain color and tannin content in Chinese sorghum based on whole-genome sequencing
Theoretical and Applied Genetics (2023)
-
A foliar pigment-based bioassay for interrogating chloroplast signalling revealed that carotenoid isomerisation regulates chlorophyll abundance
Plant Methods (2022)