Abstract
The profile of polypeptides separated by SDS-PAGE from seed of major crop species such as pea (Pisum sativum) is complex, resulting from cleavage (processing) of precursors expressed from multiple copies of genes encoding vicilin and legumin, the major storage globulins. Translation in vitro of mRNAs hybrid-selected from mid-maturation pea seed RNAs by defined vicilin and legumin cDNA clones provided precursor molesules that were cleaved in vitro by a cell-free protease extract obtained from similar stage seed; the derived polypeptides were of comparable sizes be those observed in rivo. The feasibility of transcribing mRNA in vitro from a cDNA clone and cleavage in vitro of the derived translation products was established for a legumin clone, providing a method for determining polypeptide products of an expressed sequence. This approach will also be useful for characterising cleavage site requirements since modifications an readily be introduced at the DNA level.
Similar content being viewed by others
Introduction
The globulins stored in pea (Pisum sativum) seeds are encoded by gene families consisting of approximately 10 genes for legumins or 11 S proteins 1, 2, and approximalely 18 genes for vicilins or 7S proteins 2, 3, 4. However, considerable complexiby is revealed by polyacrylamide gel electrophoresis (PAGE) of the mature proleins. Endoproteolytic cleavage of precursor polypeptides 5, 6, 7, 8, followed by specific or quasi-specifio associations of the derivalives, appears to be the major cause of complexity at the protein level, although other co-or post-translational modifications such as glycosylation and deamidation may also contribute. Assessment of the relative contribution of any specific gene to total accumulation of storage protein requires direct correlation of the generate sequence, amounts of expressed mRNA and levels of both specific proteins synthesized and derivative peptides accumulated. These aspects are intrinsic to understanding the developmental biology of storage proteins and have practical value in that modification of highly expressed genes. It will be most effective in changing nutritional or functional properties of the grain.
Several approaches exist for determining the derivation of the seed pelypeptides. Comparisons of genomic and cDNA ,sequences with those of amino acid sequences of purified legumins have been carried out 9, 10. Although valuable, this approach is impractical for resolving the total complexity of the pea seed proteins because of the number of genes involved. Additionally, complete rigor is required to ensure that the protein is derived from the genomic sequence identified, and it is difficult to ensure that only one gene (encoding the protein sequence determined) exists. Genetical analyses have allowed the correlation of one class of legumin precursor with a class of α-subunit 11 and have eliminated several vicilin precursor classes from the possible precursors of the predominant vicilin polypeptide of Mr 50000 3.
Endoproteolytic cleavage of castor bean lectin precursors has been achieved in vitro using crude extracts of developing seeds; the results obtained with orude extracts mimicked those obtained with protein body extracts 12. In this paper, we have adapted these methods to assess the contribution of several seed protein gene classes to the profile seen on PAGE of pea proteins. The approach depends upon the cleavage of globulin precursors translated in vitro from hybrid-selected RNAs using a cell-free protease system and allows the correlation of precursor classes with subunit classes observed in vivo. Additionally we have transcribed a legumin cDNA clone in vitro and processed the derived polypeptide with the protease extract. An extension of the latter approach is its future application to understanding the amino acid determinants required for proteolytic cleavage.
Materials and Methods
Plant material
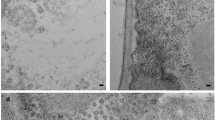
Immature seeds containing embryos of 200−350mg fresh weight were harvested from greenhouse grown plants of the Pisum sativum genotype BC1/7RR 13 or cv. Birte and immediately used in the preparation of processing extract. Liquid nitrogen frozen immature seeds of cv. Birte were used in the preparation of polyadenylated RNA.
Preparation of precursors
Polyadenylated RNA, isolated as previously described 14, was hybridized to the vicilin cDNA clones pCD70, pCD48 and pCD86 and to legumin cDNA clones pCD32, pCD40 and pCD43 2, 15 which were bound to diazobenzyloxymethyl (DBM) paper 16. The characterization of clone pCD48 has been described elsewhere 2, 16. The clone pCD70 contains a longer insert than, and is closely homologous to, pCD4 2, 16. The clone pCD86 is closely homologous to the plasmid pJC2−7 3 and represents a third class of vicilin precursor of Mr approx. 50000 in addition to those represented by pCD70 and pCD48. The legumin cDNA clone p10 represents a full-length cDNA corresponding to pCD43 sub-cloned into the transcription vector, pT7-1 (U. S. Biochemioals). Transcriptions were 0.5mM cap performed according to the manufacturer's instructions except that 50 μM GTP and analogue were used. RNasin (1U/μl) was included and reactions proceeded for 90 min. Reactions were extracted with phenol/chloroform and ethanol preipitated.
The RNA which hybridized to cDNA clones was released from the DBM paper, as previously described 16 ecoept thas the elution buffer contained 50% (v/v) deionized formamide, and was translated in a rabbit retioulocyte translation system using 20 μl messenger-dependent lysate and 9.105 Bq [3H] leucine or [35S] methionine.
Processing of precursors
The products of selected RNAs were recovered from cell-free translations by incubation wish anti-legumin or anti-vicilin IgG, as appropriate, and protein A-Sepharose 16. The pellet of protein A-Sepharose-IgG-preoursor was washed five times with 0.75M NaCl, 1% (v/v) Triton X-100, twice wish half-strength citrate-phosphate buffer, pH 5 17, suspended in 40 μl of enzyme extract (see later) and incubated at 30°C for 22h. The protein A-Sepharose-IgG-protein complex was pelleted and dissociated by boiling in twice-strength Laemmli 18 sample buffer. Unprocessed and processed products were analyzed by electrophoresis on 15% SDS polyaerylamide gels (PAGE) and fluorography. In one case, products were processed in situ after separation on SDS-PAGE. The translation products were first separated on a 10% SDS polyacrylamido gel. The gel track containing the separated polypeptides was cut out, fixed in two changes of 30% ethanol, 12% acetic acid for 20 min. each followed by several changes of half-strength citrate-phosphate buffer pH 5. The gel slice was incubated in 10ml enzyme extract at 30°C for 22h., equilibrated with twice-strength Laemmli 18 sample buffer and sealed on top of a second dimension gel (15%) using 1% (w/v) agarose in sample buffer.
Each processing experiment was performed at least twice and results were consistent between experiments.
Preparation of enzyme e∼tract
The testas were removed from seeds and the embryos extracted in 100 mM sodium phosphate buffer, pH 7.2 12. The cleared supernatant was dialyzed into half-strength citrate-phosphate buffer, pH 517 at 4°C for 5h. Insoluble material was removed from the dialysate by centrifugation at 16000g for 5 min.; the supernatant was used immediately for processing, or, was stored at −80°C.
Results
Vicilin precursor processing
The proteolytic processing of Pisum vioilin proteins is complex. Many of the smaller subunits observed in vivo are derived from a family of precursors of Mr approx. 50000 6, 7. Based on partial protein and cDNA sequence information, a scheme for the derivation of smaller vicilin subunits from precursors has been proposed [4, 19; see also Fig. l]. This scheme suggests the presence of two potential processing sites, with the subunits α, β and γ resulting from cleavage of precursors of the form NH2−α−β−γ−COOH at both sites; the α−, β−and γ−subunits are of Mr approx. 18000−19000, 13000−13500 and 12000−12500, respectively 4, 19. Processing at one of the sites alone has been suggested to give rise to subunits of intermediate sizes varying from Mr 25000 to 33000 4, 7. the derivation of minor subunibs of Mr 34000−35000 7, 19 is not clear from those sbudies.
Upper section; A diagrammatic representation of precursor−product relationships for vicilin based on published in vivo data (refs. 4, 6, 7, 19).Lower section: Results obtained from this work and diagrammatic representation thereof.An analysis by PAGE and fluorography or autoradiography of unprocessed (a, e, f) and processed (b, d, e, g) translation products of RNA selected by vicilin-related plasmids pCD70 (a, b), pCD48 (c, d, e) and pCD86 (f, g). Translations were performed in the presence of [3H] lencinc in all cases except (e) which shows the processed products of [35S] methionine labelled precursor. The positions adopted by [14C] labelled protein standards are shown.
Translation in vitro of RNA hybrid-selected by pCD70, pCD48 and pCD86 yielded polypeptides of Mr 47000−50000 (Fig. 1a, e, f, respectively). After protease treabment, the vicilin precursor of Mr 47000 (Fig. 1a) yielded subunits of Mr approx. 18000 (probably α) and 16000 (probably β and γ) (Fig. 1b). The similarity in size of the β-ant γ−vicilin subunits (Mr approx. 13000 in refs. 4 and 19) presumably prevents their resolution on the gels shown in Fig. 1 (Based on the vicilin precursors for which sequences are available, the intensity of [3H] leucine-labelled α-subunit would be expected to be equivalent to that of [3H] leucine-labelled β+γ-subunits). Processing of the vicilin precursor of Mr 50000 corresponding to pCD48 (Fig. 1c) also gave polypeptides of Mr approx. 18000 and 160(70 bah in addition gave pelypeptides of Mr approx. 30000−35000 (Fig. 1d). Vicilin precursors of Mr 50000 have previously boon shown to become labelled with [35S] methionine due to a number of methionine residues in the signal pepbide 20. Processing of [35S] methionine-labelled precursor corresponding to pCD48 (conbaining four methionine residues in the signal polypeptide 21) showed that the subunit of M, approx. 18000 was labelled whereas that of Mr approx. 16000 was not (Fig. 1e). Thus the methionine-containing subunit of Mr 18000 can be correlated with the N-terminal region of the vicilin precursor 4, 19 corresponding to the α-subunit 4 which in this study would have retained the signal peptide.
The products of RNA hybrid-selected by pCD86 (Fig, 1f) were of two size-classes: precursors of Mr 50000 (indistinguishable from those corresponding to pCD 48; Fig. lc) and precursors intermediate in size between those corresponding to pCD 70 (Fig. 1a) and pCD 48 (Fig. 1c). Processing of these two bands also gave polypeptides of Mr 18000 and 16000 but in addition yielded minor polypeptides of Mr approx. 25000−30000 (Fig. 1g).
The presence of polypeptides of Mr approx. 25000−35000 among the processed products of RNA corresponding to pGD48 and pGD86 (Fig. 1d, g) suggests that some of these precursors are processed at one but not at both of the two potential -processing sites. Processing at the second (βγ) processing site alone would be expected to give a subunit of Mr approx. 30000−33000 and a subunit of 12000−12500 4, 19; processing at the first (αβ) processing site alone would be expected to yield a subunit of 18000 -19000 and a subunit of 25000−30000 4, 7, 19. On this basis, the results of processing the precursors corresponding to pCD48 and pCD86 suggest that some of the former were processed at the second (βγ) site alone whereas some of the latler were processed at the first (αβ) site alone. Several minor bands are visible in the 40000 and 20000 range, and these are probably degradation products of the protein.
Legumin precursor processing
Legumin is synthesized as a set of precursor molecules that are endoprobeolytically cleaved to yield acidic (α) and basic (β) subunits that undergo disulfide-bonding to yield α/β dimers, six of which form a hexamer. The majority of legumin α-(Mr 40000) and β-(Mr 20000) subunits derive from Mr 60000 precursors (of the form NH2–α–β–COOH) 1. Smaller legumin dimers having α-subunits of Mr approx. 25000 bonded to Mr approx. 21000 β-subunits have been observed 22, 23; these might be expecbed to derive from Mr 45000 precursors, but the latter have not been observed. Conversely, precursors of Mr 80000 have been reported 24, 25 but their subunit derivatives have nob previously been identified.
Translation in vitro of mRNAs hybrid-selected by cDNA clones pCD32, pCD43 and pCD40 yielded polypeptides of Mr 80000, 60000 and 63000−65000, respectively (Fig. 2a, c and first-dimensional gel of Fig. 2e). After protease treatment, the Mr 80000 precursor yielded predominantly polypeptides of Mr approx. 25000, 23000 and 20000 (Fig. 2b); the sizes of two of these subunits correlate well with those derived from “small” legumin dimers of Mr 35000 that apparently consist of α-subunits of Mr 25000 and β-subunits of 21000 22, 23. The partial sequence of pCD32 predicts a β-subunit of Mr 19500 26, but no absolute date are available for the sizes or numbers of other subunits corresponding to this precursor. Incubation of precursors in buffer alone or in boiled extract did not result in any change of Mr (date not shown). After processing, the precursor of Mr 60000 yielded polypeptides of Mr approx. 40000 and 20000 (Fig. 2d) indicating that the products of pGD43 class constitute the majority of legumin. Due to considerable homology between pCD32 and pCD40 26, the products of RNA hybrid-selected by pCD40 also contain some precursors of Mr 80000 (25 and first dimension gel of Fig. 2e). To dis binguish between these products, they were processed after separation by eloctrophoresis and analyzed in a second dimension gel (Fig. 2e). Although the majority of the precursors remain unprocessed by this method, a fraction was processed to yield a heterogeneous group of polypeptides of Mr approx. 40000 and 18000−20000. Some differences in the products of the heterogeneous pCD40-related precursors were apparent (Fig. 2e). The presence of a signal peptide on the processed polypeptides of Mr 40000 precludes their correlation by two-dimensional isofocussing eloctrophoresis with αM and αm 27 subunits; attempts to remove signal peptides from these precursors using membranes prepared as described in 28 wer8 unsuccessful.
Upper section: A diagrammatic representation of precursor-product relationships for legumin based on published in vivo data. [(i)refs. 24, 25; (ii) refs. 22, 23; (iii) ref. 1]Lower section: Results obtained from this work and diagrammatic representation thereof.An analysis by PAGE and fluorography of unprocessed (a, c) and processed (b, d) translationtion products of RNA selected by the legumin-related plasmids pCD32 (a, b) and pCD43 (C, d). The asterisk(*) indicates that another processing site must exist in the N-terminal portion of the pCD32-related precursor. Unprocessed and processed translation products of RNA selected by the legumin-related plasmid pCD40 are shown in (e). The arrow shows the direction of electrophoresis in the first dimensional gel, which shows the unprocessed translation products. These products were treated with processing extract in the first dimensional gel and subsequently electrophoresed into the second dimensional gel. The translation product of RNA transcribed from the full-length legumin clone pl0 is shown in (f) and the procluets derived from this after processing in (g). Translations were performed in the presence of [3H] leucine. The positions adopted by [14C] labelled protein standards are shown.
Processing a legumin precursor corresponding to a single transcript
In vitro translation of the RNA derived by transcription of the clone pl0 (containing a full-length legumin cDNA)yielded potypeptides of Mr 60000 (Fig. 2f). After protease treatment, polypeptides of Mr 40000 and 20000 were obtained (Fig. 2g). The multiplicity of 20000 Mr subunits obtained from a single precursor species indicate either that cleavage between the two classes of polypeptide is "ragged" or thai some C-terminal processing of the M, 20000 subunits occurred. The former appears unlikely in that sequence data 10 have suggested that the cleavage of legumin precursors of Mr 60000 is a single proteolytic event which occurs between paired Asn-Gly residues resulting in a C-terminal Asn for the α-subunit and an N-terminal Gly for the β-subunit. This proteolytic site appears to be very conserved, even between different species (see 21), although a recent study reports an N-terminal Phe for a minor β-subunit in Pisum 29. Further processing of C-termini has been suggested to occur in the ease of some 11S proteins (glycinin) from Glycine max 30.
Discussion
The correlation of precursors and their constituent subunits has been a long-standing problem which, due to the numbers of genes involved 2, is not necessarily simplified by limited sequence analyses. In this paper, in vitro translation of hybrid-selected RNAs and processing of the translation products have shown that subunits similar in size to those observed in vivo can be correlated with precursor polypeptides representing different gone classes.
Differences in the extent of processing of the three vieilin precursor classes examined (Fig. 1) may reflect sequence variation at the putative processing sites, or, may be a result of conformation changes due to sequence differences elsewhere in the molecules. The fact that only a small proportion of vicilin precursors of Mr 50000 remained unprocessed in these experiments (although in vivo, polypeptides of Mr 50000 constitute a substantial portion of vioilin subunits) is not surprising as a cDNA class corresponding to these major Vtoilin subunits has not so far been identified (see 3). For both pCD48 and pCD86, some, precursors were apparently processed at both sites and sequence comparisons of cDNAs representative of the three classes do not clarify the picture. Comparisons of cDNA and protein sequence data have highlighted differences between sequences but have not proven which are processed and which are not 31. The lack of proteolytic cleavage of homologous proteins in Phaseolus vulgaris (phaseolin) and Glycine max (β-conglycinin) appears fo be a consequence of deletions existing in these, relative to the other sequences, in the areas of potential cleavage (see 32).
Genetieal analyses of, and assessment of the relative amounts of mRNA corresponding to, the legumin gone class represented by pCD43 have previously indicated that this class is responsible for the majority of legumin synthesized during seed development 11, 15; the similarify in size between the processed products of the pCD43-related RNA (Fig. 2d) and that of the major legumin subunits in vivo is in accord with this concept. The precursors of Mr 80000 were processed to give subunits similar in size (Fig. 2b) to those observed in vivo for “small” legumin dimers of the Lg-3 size-class [9.2, 23]. Clearly the size of this precursor is in excess of that required for one α- and one β-subunit. It is probable that each precursor is processed to give three or more subunits, a situation that may be analogous to the A5A4B3 glyeinin precursor 33. The partial predicted sequence of the precursor of Mr 80000 has shown that it contains a long polar tract of approx. 100 amino acids 26. The length and nature of this tract, which occurs just to the amino side of an αβ cleavage site, may cause anomalous migration on SDS gels.
To alleviate diffieulties intrinsic to the analysis of classes of precursors, we have begun an analysis of individual precursors. The results obtained with a single legumin precursor derived by transcription and translation from a full-length legumin clone are presented here. The multiplicity of the subunits derived from a single precursor is not surprising in that a multiplicity of products derived from a single vicilin or β-conglycinin gene has been observed in transgenic plants 34, 35. An extension of these studies using other full-length clones should further clarify the derivabion of legumin and vicilin polypeptides. As has been shown for processing of viral polyproteins 36, this system provides an approach to defining processing sites by in vitro mutagenesis of putative sites, and, subsequent analysis of the derived mutant polypeptides by in vitro processing.
References
Croy RRD, GW Lycett, JA Gatehouse, JN Yarwood and D Boulter . C1onlng and analysis of cDNAs encoding plant storage protein precursors. Nature 1982; 295: 76–79.
Domoney C and R Casey . Measurement of gone number for seed storage proteins in Pisum. Nucleic Acids Res 1985; 13: 687–699.
Ellis THN, C Domoney, J Castleton, W Cleary and DR Davies . Vicilin genes of Pisum. Mol Gen Genet 1986; 205: 164–169.
Gatehouse JA, GW Lycett, AJ Delauney, RRD Croy and D BouIter . Sequence specificity of the post-translational proteclytic cleavage of vicilin, a seed storage protein of pea (Pisum sativum L.). Biochem J 1983; 212: 427–432.
Croy RRD, JA Gatehouse, IM Evans and D Boulter . Characterization of the storage protein subunits synthesized in vitro by polyribosomes and RNA from developing pea (Pisum sativum L.). I. Legumin Planta 1980; 148: 49–56.
Gatehouse JA, RRD Croy, H Morton, M Tyler and D Boulter . Characterization and subunit structures of the vicilin storage proteins of pea (Pisum sativum L.). Eur J Biochem 1981; 118: 627–633.
Gatehouse JA, GW Lycett, RRD Croy and D Boulter . The post-translational proteolysis of the subunits of vicilin from pea (Pisum sativum L.). Biochem J 1982; 207: 629–632.
Spencer D and TJV Higgins . The biosynthesis of legumin in maturing pea seeds. Biochem Int 1980: 1: 502–509.
Gatehouse JA, et al. Two genes encoding 'minor' legumin polypeptides in pea (Pisum sativum L.). Biochem J 1988; 250: 15–24.
Lycett GW, RRD Groy, AD Shirsat and D Boulter . The complete nucleotide sequence of a legumin gene from pea (Pgsum sativum L.). Nucleic Acids Res 1984; 12: 4493–4506.
Domoney C, THN Ellis and DR Davies . Organization and mapping of legumin genes in Pisum. Mol Gen Genet 1986; 202: 280–285.
Harley SM and JM Lord . In vitro endoproteolytic cleavage of castor bean lectin precursors. Plant Science 1985; 41: 111–116.
Hedley CL, CM Smith, MJ Ambrose, S Cook and TL Wang . An analysis of seed development in Pisum sativum II. The effect of the r-locus on the growth and development of the seed. Ann Bot 1986; 58: 371–379.
Casey R, C Domoney and NC Nielsen . Isolation of a cDNA clone for pea (Pisum sativum) seed lipoxygenase. Biochem J 1985; 232: 79–85.
Domney C and R Casey . Changes in legumin messenger RNAs throughout seed development in Pisum sativum L. Planta 1987; 170: 562–566.
Domoney C and R Casey . Cloning and characterization of complementary DNA for convicilin, a major seed storage protein in Pisum sativum L. 1983; 159: 446–453.
Domori G . Preparation of buffer for use in enzyme studies. In: Methods in enzymology. 1955; 1: 138–146.
Laemmli UK . Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 1970; 227: 680–685.
Spencer D, PM Chandler, TJV Higgins, Inglis and M Rubira . Sequence interrelationships of the subunits of vicilia from pea seed. Plant Mol Biol 1983; 2: 259–267.
Spencer D . The physiological role of storage proteins in seeds. Phil Trans R Soc Lend B 1984; 304: 275–285.
Casey R, C Domoney and N Ellis . Legume storage proteins and their genes, In: BJ Miflin ed. Oxford surveys of plant molecular and cell biology Oxford University Press Oxford 1986; 3: 1–95.
Matta NK, JA Gateehouse and D Boulter . Molecular and subunit heterogeneity of legnmin of Pisum sativum L. (Garden pea)–a multidimensional gel electrophoretic study. J Exp Bet 1981; 32: 1295–1307.
Matta NK and JA Gatehouse . Inheritance and mapping of storage protein genes in Pisum sativum L. Heredity 1982; 48: 383–392.
Chrispeels MJ, TJV Higgins and D Spencer . Assembly of storage protein oligomers in the endoplasmic reticulum and processing of the polypeptides in the protein bodies of developing pea cotyledons. J Cell BioI 1982; 93: 306–313.
Domoney C and R Casey . Storage protein precursor polypeptides in cotyledons of Pisum sativum L. Identification of, and isolation of a cDNA clone for, an 80000-Mr legumin-related polypeptide. Eur J Biochem 1984; 139: 321–327.
Domoney C, D Barker and R Casey . The complete deduced amino acid sequences of legumin β–polypetides from different genetic loci in Pisum. Plant Mol Biol 1986; 7: 467–474.
Casey R . Genetic variability in the structure of the α-subunits of legumin from Pisum–a two-dimensional gel electrophoresis study, Heredity 1979; 43: 265–272.
Higgins TJV and D Spencer . Precursor forms of pea vicilin subunits. Modification by microsomal membranes during cell-free translation. Plant Physiol 1981; 67: 205–211.
March JF, DJC Pappin and R Casey . Isolation and charactrization of a minor legumin and its constituent polypeptides from Pisum sativum (pea). Biochem J 1988; 250: 911–915.
Marco YA, et al. Cloning and structural analysis of DNA encoding a A2 B1a subunit of glycinin. J Biol Chem 1984; 259: 13436–13441.
Lycett GW, AJ Delauney, JA Gatehouse, J Gilroy, RRD Croy and D Boulter . The vicilin gone family of pea (Pisum sativum L.)–a complete cDNA coding sequence for preprovicilin. Nucleic Acids Res 1983; 11: 2367–2380.
Doyle JJ, MA Schuler, WD Godette, V Zenger RN Beachy and JL Slightom . The glycosylated seed storage proteins of Glycine max and Phaseolus vulgaris. Structural homologies of genes and proteins. J Biol Chem 1986; 261: 9228–9238.
Momma T, et al. Glycinin A5A4B3 mRNA; cDNA cloning and nucleotide sequencing of a splitting storage protein subunit of soybean. Eut J Biochem 1985; 149: 491–496.
Beachy RN, et al. Accumulation and assembly of soybean β -conglycinin in seeds of transformed petunia plants. EMBO J 1985; 4: 3047–3053.
Higgins TJV, EJ Newbigin, D Spencer, DJ Llewellyn and S Craig . The Sequence of a pea vicilin gene and its expression in transgenic tobacco plants. Plant Mol Biol 1988; 11: 683–695.
Dougherty WG, JC Carrington, SM Cary and TD Parks . Biochemical and mutational analysis of a plant virua pelyprotein cleavage site. EMBO J 1988; 7: 1281–1287.
Acknowledgements
We thank S. Turner for the gift of antibody preparations, Dr. T. Wang for plants of the genotype BC1/7RR and Dr. N. ,Ellis, Dr. T. Wang and Professor D. R. Davies for their useful comments on the manuscript.
We gratefully acknowledge receipt of NATO grant 85/0742. This work was also supported by the Agricultural and Food Research Council via a grant-in-aid to the John Innes Institute.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Yang, L., Domoney, C., Casey, R. et al. Processing Pisum sativum seed storage protein precursors in vitro. Cell Res 1, 153–162 (1990). https://doi.org/10.1038/cr.1990.15
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/cr.1990.15