Abstract
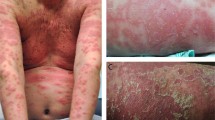
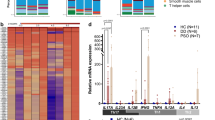
Psoriasis is a chronic inflammatory skin disease with a strong genetic background and is triggered by environmental factors. Available evidence supports CD6, a lymphocyte surface receptor mostly expressed by T cells, as a putative target in autoimmunity. Accordingly, a humanized anti-CD6 antibody has been assayed for the treatment of certain autoimmune disorders, including psoriasis. Here, we present novel evidence in mice and humans for a direct involvement of CD6 in psoriasis pathophysiology. First, an attenuated form of imiquimod-induced psoriasis-like skin inflammation was demonstrated in CD6-deficient mice, as deduced from lower epidermal thickness and local reduced production of pro-inflammatory cytokines, namely, interleukin-17A. Thus, isolated CD4+CD62L+ T cells from CD6-deficient mice displayed decreased in vitro T-helper type 17 polarization. Second, a statistically significant association between CD6 single-nucleotide polymorphisms (rs17824933, rs11230563 and rs12360861) and more severe forms of psoriasis was demonstrated in a cohort of 304 patients at three public hospitals from the metropolitan area of Barcelona. Taken together, these results provide new supportive evidence of the contribution of the CD6 lymphocyte receptor in psoriasis at both experimental and clinical levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Greb JE, Goldminz AM, Elder JT, Lebwohl MG, Gladman DD, Wu JJ et al. Psoriasis. Nat Rev Dis Prim 2016; 2: 16082.
Hwang ST, Nijsten T, Elder JT. Recent highlights in psoriasis research. J Invest Dermatol 2017; 137: 550–556.
Ray-Jones H, Eyre S, Barton A, Warren RB. One SNP at a time: moving beyond GWAS in psoriasis. J Invest Dermatol 2016; 136: 567–573.
Kofler DM, Farkas A, von Bergwelt-Baildon M, Hafler DA. The link between CD6 and autoimmunity: genetic and cellular associations. Curr Drug Targets 2016; 17: 651–665.
Hernández P, Moreno E, Aira LE, Rodríguez PC. Therapeutic targeting of CD6 in autoimmune diseases: a review of cuban clinical studies with the antibodies IOR-T1 and itolizumab. Curr Drug Targets 2016; 17: 666–677.
Krupashankar DS, Dogra S, Kura M, Saraswat A, Budamakuntla L, Sumathy TK et al. Efficacy and safety of itolizumab, a novel anti-CD6 monoclonal antibody, in patients with moderate to severe chronic plaque psoriasis: results of a double-blind, randomized, placebo-controlled, phase-III study. J Am Acad Dermatol 2014; 71: 484–492.
Sarukhan A, Martinez-Florensa M, Escoda-Ferran C, Carrasco E, Carreras E, Lozano F. Pattern recognition by CD6: a scavenger-like lymphocyte receptor. Curr Drug Targets 2015; 17: 640–650.
Brown MH. CD6 as a cell surface receptor and as a target for regulating immune responses. Curr Drug Targets 2016; 17: 619–629.
Sarrias M-R, Farnós M, Mota R, Sánchez-Barbero F, Ibáñez A, Gimferrer I et al. CD6 binds to pathogen-associated molecular patterns and protects from LPS-induced septic shock. Proc Natl Acad Sci USA 2007; 104: 11724–11729.
Martínez-Florensa M, Consuegra-Fernández M, Martínez VG, Cañadas O, Armiger-Borràs N, Bonet-Roselló L et al. Targeting of key pathogenic factors from gram-positive bacteria by the soluble ectodomain of the scavenger-like lymphocyte receptor CD6. J Infect Dis 2014; 209: 1077–1086.
Escoda-Ferran C, Carrasco E, Caballero-Baños M, Miró-Julià C, Martínez-Florensa M, Consuegra-Fernández M et al. Modulation of CD6 function through interaction with Galectin-1 and -3. FEBS Lett 2014; 588: 2805–2813.
Enyindah-Asonye G, Li Y, Ruth JH, Spassov DS, Hebron KE, Zijlstra A et al. CD318 is a ligand for CD6. Proc Natl Acad Sci USA 2017; 114: E6912–E6921.
Orta-Mascaró M, Consuegra-Fernández M, Carreras E, Roncagalli R, Carreras-Sureda A, Alvarez P et al. CD6 modulates thymocyte selection and peripheral T cell homeostasis. J Exp Med 2016; 213: 1387–1397.
Li Y, Singer NG, Whitbred J, Bowen MA, Fox DA, Lin F. CD6 as a potential target for treating multiple sclerosis. Proc Natl Acad Sci USA 2017; 114: 2687–2692.
Consuegra-Fernández M, Martínez-Florensa M, Aranda F, de Salort J, Armiger-Borràs N, Lozano T et al. Relevance of CD6-mediated interactions in the regulation of peripheral T-cell responses and tolerance. Front Immunol 2017; 8: 594.
Enyindah-Asonye G, Li Y, Xin W, Singer NG, Gupta N, Fung J et al. CD6 receptor regulates intestinal ischemia/reperfusion-induced injury by modulating natural IgM-producing B1a cell self-renewal. J Biol Chem 2017; 292: 661–671.
Kobarg J, Whitney GS, Palmer D, Aruffo A, Bowen MA. Analysis of the tyrosine phosphorylation and calcium fluxing of human CD6 isoforms with different cytoplasmatic domains. Eur J Immunol 1997; 27: 2971–2980.
Bonet L, Farnós M, Martínez-Florensa M, Martínez VG, Lozano F. Identification of functionally relevant phoshorylatable serine clusters in the cytoplasmic region of the human CD6 lymphocyte surface receptor. FEBS Lett 2013; 587: 2205–2213.
Ibáñez A, Sarrias M-R, Farnós M, Gimferrer I, Serra-Pagès C, Vives J et al. Mitogen-activated protein kinase pathway activation by the CD6 lymphocyte surface receptor. J Immunol 2006; 177: 1152–1159.
Breuning J, Brown MH. T cell costimulation by CD6 is dependent on bivalent binding of a GADS/SLP-76 complex. Mol Cell Biol 2017; 37; doi: 10.1128/MCB.00071-17.
Hem CD, Ekornhol M, Granum S, Sundvold-Gjerstad V, Spurkland A. CD6 and linker of activated T cells are potential interaction partners for T cell-specific adaptor protein. Scand J Immunol 2017; 85: 104–112.
Gimferrer I, Ibáñez A, Farnós M, Sarrias M-R, Fenutría R, Roselló S et al. The lymphocyte receptor CD6 interacts with syntenin-1, a scaffolding protein containing PDZ domains. J Immunol 2005; 175: 1406–1414.
Gimferrer I, Calvo M, Mittelbrunn M, Farnós M, Sarrias MR, Enrich C et al. Relevance of CD6-mediated interactions in T cell activation and proliferation. J Immunol 2004; 173: 2262–2270.
Santos RF, Oliveira L, Carmo AM. Tuning T cell activation: the function of CD6 at the immunological synapse and in T cell responses. Curr Drug Targets 2016; 17: 630–639.
van der Fits L, Mourits S, Voerman JSA, Kant M, Boon L, Laman JD et al. Imiquimod-induced psoriasis-like skin inflammation in mice is mediated via the IL-23/IL-17 axis. J Immunol 2009; 182: 5836–5845.
Cenit MC, Martínez-Florensa M, Consuegra M, Bonet L, Carnero-Montoro E, Armiger N et al. Analysis of ancestral and functionally relevant CD5 variants in systemic lupus erythematosus patients. PLoS One 2014; 9: e113090.
Potrony M, Carreras E, Aranda F, Zimmer L, Puig-Butille J-A, Tell-Martí G et al. Inherited functional variants of the lymphocyte receptor CD5 influence melanoma survival. Int J Cancer 2016; 139: 1297–1302.
Delgado J, Bielig T, Bonet L, Carnero-Montoro E, Puente XS, Colomer D et al. Impact of the functional CD5 polymorphism A471V on the response of chronic lymphocytic leukaemia to conventional chemotherapy regimens. Br J Haematol 2016; 177: 147–150.
Wagner M, Bilinska M, Pokryszko-Dragan A, Sobczynski M, Cyrul M, Kusnierczyk P et al. ALCAM and CD6—multiple sclerosis risk factors. J Neuroimmunol 2014; 276: 98–103.
Zhou P, Du L-F, Lv G-Q, Yu X-M, Gu Y-L, Li J-P et al. Functional polymorphisms in CD166/ALCAM gene associated with increased risk for breast cancer in a Chinese population. Breast Cancer Res Treat 2011; 128: 527–534.
Varadi V, Bevier M, Grzybowska E, Johansson R, Enquist-Olsson K, Henriksson R et al. Genetic variation in ALCAM and other chromosomal instability genes in breast cancer survival. Breast Cancer Res Treat 2012; 131: 311–319.
De Jager PL, Jia X, Wang J, de Bakker PIW, Ottoboni L, Aggarwal NT et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat Genet 2009; 41: 776–782.
Swaminathan B, Cuapio A, Alloza I, Matesanz F, Alcina A, García-Barcina M et al. Fine mapping and functional analysis of the multiple sclerosis risk gene CD6. PLoS One 2013; 8: e62376.
Ahlehoff O, Skov L, Gislason G, Lindhardsen J, Kristensen SL, Iversen L et al. Cardiovascular disease event rates in patients with severe psoriasis treated with systemic anti-inflammatory drugs: a Danish real-world cohort study. J Intern Med 2013; 273: 197–204.
Gelfand JM, Shin DB, Neimann AL, Wang X, Margolis DJ, Troxel AB. The risk of lymphoma in patients with psoriasis. J Invest Dermatol 2006; 126: 2194–2201.
Gelfand JM, Neimann AL, Shin DB, Wang X, Margolis DJ, Troxel AB. Risk of myocardial infarction in patients with psoriasis. JAMA 2006; 296: 1735.
Gelfand JM, Troxel AB, Lewis JD, Kurd SK, Shin DB, Wang X et al. The risk of mortality in patients with psoriasis: results from a population-based study. Arch Dermatol 2007; 143: 1493–1499.
Kurd SK, Troxel AB, Crits-Christoph P, Gelfand JM, Catalin MP, Hywel W et al. The risk of depression, anxiety, and suicidality in patients with psoriasis: a population-based cohort study. Arch Dermatol 2010; 146: 891–895.
Schmitt J, Wozel G. The psoriasis area and severity index is the adequate criterion to define severity in chronic plaque-type psoriasis. Dermatology 2005; 210: 194–199.
Henseler T, Christophers E. Psoriasis of early and late onset: characterization of two types of psoriasis vulgaris. J Am Acad Dermatol 1985; 13: 450–456.
Ferrándiz C, Pujol RM, García-Patos V, Bordas X, Smandía JA. Psoriasis of early and late onset: a clinical and epidemiologic study from Spain. J Am Acad Dermatol 2002; 46: 867–873.
Langley RGB, Krueger GG, Griffiths CEM. Psoriasis: epidemiology, clinical features, and quality of life. Ann Rheum Dis 2005; 64 (Suppl 2): ii18–ii23; discussion ii24–ii25.
Swindell WR, Michaels KA, Sutter AJ, Diaconu D, Fritz Y, Xing X et al. Imiquimod has strain-dependent effects in mice and does not uniquely model human psoriasis. Genome Med 2017; 9: 24.
Wu L, Schaid DJ, Sicotte H, Wieben ED, Li H, Petersen GM. Case-only exome sequencing and complex disease susceptibility gene discovery: study design considerations. J Med Genet 2015; 52: 10–16.
Sarrias MR, Gronlund J, Padilla O, Madsen J, Holmskov U, Lozano F. The scavenger receptor cysteine-rich (SRCR) domain: an ancient and highly conserved protein module of the innate immune system. Crit Rev Immunol 2004; 24: 1–38.
Krintel SB, Essioux L, Wool A, Johansen JS, Schreiber E, Zekharya T et al. CD6 and syntaxin binding protein 6 variants and response to tumor necrosis factor alpha inhibitors in Danish patients with rheumatoid arthritis. PLoS One 2012; 7: e38539.
Zheng M, Zhang L, Yu H, Hu J, Cao Q, Huang G et al. Genetic polymorphisms of cell adhesion molecules in Behcet’s disease in a Chinese Han population. Sci Rep 2016; 6: 24974.
Hamzaoui K. Th17 cells in Behçet’s disease: a new immunoregulatory axis. Clin Exp Rheumatol 2011; 29: S71–S76.
Dos Passos GR, Sato DK, Becker J, Fujihara K. Th17 cells pathways in multiple sclerosis and neuromyelitis optica spectrum disorders: pathophysiological and therapeutic implications. Mediat Inflamm 2016; 2016: 5314541.
Marinoni B, Ceribelli A, Massarotti MS, Selmi C. The Th17 axis in psoriatic disease: pathogenetic and therapeutic implications. Auto Immun Highlights 2014; 5: 9–19.
Egeberg A, Mallbris L, Gislason GH, Skov L, Hansen PR. Risk of multiple sclerosis in patients with psoriasis: a Danish Nationwide Cohort Study. J Invest Dermatol 2016; 136: 93–98.
Bughani U, Saha A, Kuriakose A, Nair R, Sadashivarao RB, Venkataraman R et al. T cell activation and differentiation is modulated by a CD6 domain 1 antibody Itolizumab. PLoS One 2017; 12: e0180088.
Zúñiga LA, Jain R, Haines C, Cua DJ. Th17 cell development: from the cradle to the grave. Immunol Rev 2013; 252: 78–88.
Hawkes JE, Gudjonsson JE, Ward NL. The snowballing literature on imiquimod-induced skin inflammation in mice: a critical appraisal. J Invest Dermatol 2017; 137: 546–549.
Kofler DM, Severson CA, Mousissian N, De Jager PL, Hafler DA. The CD6 multiple sclerosis susceptibility allele is associated with alterations in CD4+ T cell proliferation. J Immunol 2011; 187: 3286–3291.
Castro MAA, Oliveira MI, Nunes RJ, Fabre S, Barbosa R, Peixoto A et al. Extracellular isoforms of CD6 generated by alternative splicing regulate targeting of CD6 to the immunological synapse. J Immunol 2007; 178: 4351–4361.
Puig L, Bordas X, Carrascosa JM, Daudén E, Ferrándiz C, Hernanz JM et al. Consensus document on the evaluation and treatment of moderate-to-severe psoriasis. Spanish Psoriasis Group of the Spanish Academy of Dermatology and Venereology. Actas Dermosifiliogr 2009; 100: 277–286.
Solé X, Guinó E, Valls J, Iniesta R, Moreno V. SNPStats: a web tool for the analysis of association studies. Bioinformatics 2006; 22: 1928–1929.
Acknowledgements
We thank Francesc Calafell and Elena Bosch (Institute of Evolutionary Biology, University Pompeu Fabra, Barcelona) for statistical analysis support and critically reviews of the manuscript. The work is supported by grants from the Spanish Ministerio de Economía y Competitividad (Plan Nacional I+D+i, SAF2013-46151-R and SAF2016-80535-R to FL), co-financed by the European Development Regional Fund ‘A way to achieve Europe’ ERDF. FA is supported by the Sara Borrell fellowship CD15/00016 from Instituto de Salud Carlos III.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Consuegra-Fernández, M., Julià, M., Martínez-Florensa, M. et al. Genetic and experimental evidence for the involvement of the CD6 lymphocyte receptor in psoriasis. Cell Mol Immunol 15, 898–906 (2018). https://doi.org/10.1038/cmi.2017.119
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cmi.2017.119
Keywords
This article is cited by
-
Failure of a T cell regulator: CD6 contributes to the aggravation of autoimmune inflammation
Cellular & Molecular Immunology (2019)
-
Modulation of cell adhesion and migration through regulation of the immunoglobulin superfamily member ALCAM/CD166
Clinical & Experimental Metastasis (2019)