Abstract
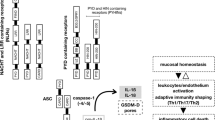
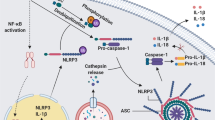
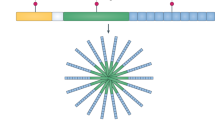
Nucleotide-binding oligomerization domain (NOD)-containing protein-like receptors (NLRs) are a recently discovered class of innate immune receptors that play a crucial role in initiating the inflammatory response following pathogen recognition. Some NLRs form the framework for cytosolic platforms called inflammasomes, which orchestrate the early inflammatory process via IL-1β activation. Mutations and polymorphisms in NLR-coding genes or in genetic loci encoding inflammasome-related proteins correlate with a variety of autoinflammatory diseases. Moreover, the activity of certain inflammasomes is associated with susceptibility to infections as well as autoimmunity and tumorigenesis. In this review, we will discuss how identifying the genetic characteristics of inflammasomes is assisting our understanding of both autoinflammatory diseases as well as other immune system-driven disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Takeuchi O, Akira S . Pattern recognition receptors and inflammation. Cell 2010; 140: 805–820.
Franchi L, Warner N, Viani K, Nunez G . Function of Nod-like receptors in microbial recognition and host defense. Immunol Rev 2009; 227: 106–128.
Wilkins C, Gale M Jr . Recognition of viruses by cytoplasmic sensors. Curr Opin Immunol 2010; 22: 41–47.
Garcia-Vallejo JJ, van Kooyk Y . Endogenous ligands for C-type lectin receptors: the true regulators of immune homeostasis. Immunol Rev 2009; 230: 22–37.
Kanneganti TD . Central roles of NLRs and inflammasomes in viral infection. Nat Rev Immunol 2010; 10: 688–698.
Allen IC, TeKippe EM, Woodford RMT, Uronis JM, Holl EK, Rogers AB et al. The NLRP3 inflammasome functions as a negative regulator of tumorigenesis during colitis-associated cancer. J Exp Med 2010; 207: 1045–1056.
Zaki MH, Vogel P, Body-Malapel M, Lamkanfi M, Kanneganti TD . IL-18 production downstream of the NLRP3 inflammasome confers protection against colorectal tumor formation. J Immunol 2010; 185: 4912–4920.
Okamoto M, Liu W, Luo Y, Tanaka A, Cai X, Norris DA et al. Constitutively active inflammasome in human melanoma cells mediating autoinflammation via caspase-1 processing and secretion of interleukin-1beta. J Biol Chem 2010; 285: 6477-6488.
Ghiringhelli F, Apetoh L, Tesniere A, Aymeric L, Ma Y, Ortiz C et al. Activation of the NLRP3 inflammasome in dendritic cells induces IL-1beta-dependent adaptive immunity against tumors. Nat Med 2009; 15: 1170–1178.
Lespinet O, Wolf YI, Koonin EV, Aravind L . The role of lineage-specific gene family expansion in the evolution of eukaryotes. Genome Res 2002; 12: 1048–1059.
Hibino T, Loza-Coll M, Messier C, Majeske AJ, Cohen AH, Terwilliger DP et al. The immune gene repertoire encoded in the purple sea urchin genome. Dev Biol 2006; 300: 349–365.
Proell M, Riedl SJ, Fritz JH, Rojas AM, Schwarzenbacher R . The Nod-like receptor (NLR) family: a tale of similarities and differences. PLoS One 2008; 3: e2119.
Bella J, Hindle KL, McEwan PA, Lovell SC . The leucine-rich repeat structure. Cell Mol Life Sci 2008; 65: 2307–2333.
Martinon F, Tschopp J . Inflammatory caspases: linking an intracellular innate immune system to autoinflammatory diseases. Cell 2004; 117: 561–574.
Faustin B, Lartigue L, Bruey JM, Luciano F, Sergienko E, Bailly-Maitre B et al. Reconstituted NALP1 inflammasome reveals two-step mechanism of caspase-1 activation. Mol Cell 2007; 25: 713–724.
Martinon F, Gaide O, Petrilli V, Mayor A, Tschopp J . NALP inflammasomes: a central role in innate immunity. Semin Immunopathol 2007; 29: 213–229.
McDonald C, Inohara N, Nunez G . Peptidoglycan signaling in innate immunity and inflammatory disease. J Biol Chem 2005; 280: 20177–20180.
Tattoli I, Travassos LH, Carneiro LA, Magalhaes JG, Girardin SE . The Nodosome: Nod1 and Nod2 control bacterial infections and inflammation. Semin Immunopathol 2007; 29: 289–301.
Hsu YM, Zhang Y, You Y, Wang D, Li H, Duramad O et al. The adaptor protein CARD9 is required for innate immune responses to intracellular pathogens. Nat Immunol 2007; 8: 198–205.
Liston P, Roy N, Tamai K, Lefebvre C, Baird S, Cherton-Horvat G et al. Suppression of apoptosis in mammalian cells by NAIP and a related family of IAP genes. Nature 1996; 379: 349–353.
Case CL, Shin S, Roy CR . Asc and Ipaf Inflammasomes direct distinct pathways for caspase-1 activation in response to Legionella pneumophila. Infect Immun 2009; 77: 1981–1991.
Martinon F, Burns K, Tschopp J . The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-beta. Mol Cell 2002; 10: 417–426.
Miao EA, Andersen-Nissen E, Warren SE, Aderem A . TLR5 and Ipaf: dual sensors of bacterial flagellin in the innate immune system. Semin Immunopathol 2007; 29: 275–288.
Shaw PJ, Lamkanfi M, Kanneganti TD . NOD-like receptor (NLR) signaling beyond the inflammasome. Eur J Immunol 2010; 40: 624–627.
Ren T, Zamboni DS, Roy CR, Dietrich WF, Vance RE . Flagellin-deficient Legionella mutants evade caspase-1- and Naip5-mediated macrophage immunity. PLoS Pathog 2006; 2: e18.
Zamboni DS, Kobayashi KS, Kohlsdorf T, Ogura Y, Long EM, Vance RE et al. The Birc1e cytosolic pattern-recognition receptor contributes to the detection and control of Legionella pneumophila infection. Nat Immunol 2006; 7: 318–325.
Hsu LC, Kobayashi KS, Kohlsdorf T, Ogura Y, Long EM, Vance RE et al. A NOD2–NALP1 complex mediates caspase-1-dependent IL-1beta secretion in response to Bacillus anthracis infection and muramyl dipeptide. Proc Natl Acad Sci USA 2008; 105: 7803–7808.
Ferwerda G, Ali SR, McGillivray S, Tseng PH, Mariathasan S, Humke EW et al. Engagement of NOD2 has a dual effect on proIL-1beta mRNA transcription and secretion of bioactive IL-1beta. Eur J Immunol 2008; 38: 184–191.
Poyet JL, Srinivasula SM, Tnani M, Razmara M, Fernandes-Alnemri T, Alnemri ES et al. Identification of Ipaf, a human caspase-1-activating protein related to Apaf-1. J Biol Chem 2001; 276: 28309–28313.
Muruve DA, Petrilli V, Zaiss AK, White LR, Clark SA, Ross J et al. The inflammasome recognizes cytosolic microbial and host DNA and triggers an innate immune response. Nature 2008; 452: 103–107.
Hornung V, Ablasser A, Charrel-Dennis M, Bauernfeind F, Horvath G, Caffrey DR et al. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature 2009; 458: 514–518.
Schroder K, Muruve DA, Tschopp J . Innate immunity: cytoplasmic DNA sensing by the AIM2 inflammasome. Curr Biol 2009; 19: R262–R265.
Mariathasan S, Newton K, Monack DM, Vucic D, French DM, Lee WP et al. Differential activation of the inflammasome by caspase-1 adaptors ASC and Ipaf. Nature 2004; 430: 213–218.
Gross O, Poeck H, Bscheider M, Dostert C, Hannesschläger N, Endres S et al. Syk kinase signalling couples to the NLRP3 inflammasome for anti-fungal host defence. Nature 2009; 459: 433–436.
Kanneganti TD, Body-Malapel M, Amer A, Park JH, Whitfield J, Franchi L et al. Critical role for Cryopyrin/Nalp3 in activation of caspase-1 in response to viral infection and double-stranded RNA. J Biol Chem 2006; 281: 36560–36568.
Pan Q, Mathison J, Fearns C, Kravchenko VV, Da Silva Correia J, Hoffman HM et al. MDP-induced interleukin-1beta processing requires NOD2 and CIAS1/NALP3. J Leukoc Biol 2007; 82: 177–183.
Martinon F, Agostini L, Meylan E, Tschopp J . Identification of bacterial muramyl dipeptide as activator of the NALP3/cryopyrin inflammasome. Curr Biol 2004; 14: 1929–1934.
Lightfield KL, Persson J, Brubaker SW, Witte CE, von Moltke J, Dunipace EA et al. Critical function for Naip5 in inflammasome activation by a conserved carboxy-terminal domain of flagellin. Nat Immunol 2008; 9: 1171–1178.
Mariathasan S, Weiss DS, Newton K, McBride J, O'Rourke K, Roose-Girma M et al. Cryopyrin activates the inflammasome in response to toxins and ATP. Nature 2006; 440: 228–232.
Craven RR, Gao X, Allen IC, Gris D, Bubeck Wardenburg J, McElvania-Tekippe E et al. Staphylococcus aureus alpha-hemolysin activates the NLRP3-inflammasome in human and mouse monocytic cells. PLoS One 2009; 4: e7446.
Gurcel L, Abrami L, Girardin S, Tschopp J, van der Goot FG . Caspase-1 activation of lipid metabolic pathways in response to bacterial pore-forming toxins promotes cell survival. Cell 2006; 126: 1135–1145.
Ozoren N, Masumoto J, Franchi L, Kanneganti TD, Body-Malapel M, Erturk I et al. Distinct roles of TLR2 and the adaptor ASC in IL-1beta/IL-18 secretion in response to Listeria monocytogenes. J Immunol 2006; 176: 4337–4342.
Boyden ED, Dietrich WF . Nalp1b controls mouse macrophage susceptibility to anthrax lethal toxin. Nat Genet 2006; 38: 240–244.
Martinon F, Petrilli V, Mayor A, Tardivel A, Tschopp J . Gout-associated uric acid crystals activate the NALP3 inflammasome. Nature 2006; 440: 237–241.
Eisenbarth SC, Colegio OR, O'Connor W, Sutterwala FS, Flavell RA . Crucial role for the Nalp3 inflammasome in the immunostimulatory properties of aluminium adjuvants. Nature 2008; 453: 1122–1126.
Dostert C, PÊtrilli V, van Bruggen R, Steele C, Mossman BT, Tschopp J . Innate immune activation through Nalp3 inflammasome sensing of asbestos and silica. Science 2008; 320: 674–677.
Cassel SL, Eisenbarth SC, Iyer SS, Sadler JJ, Colegio OR, Tephly LA et al. The Nalp3 inflammasome is essential for the development of silicosis. Proc Natl Acad Sci USA 2008; 105: 9035–9040.
Mayor A, Martinon F, de Smedt T, Petrilli V, Tschopp J . A crucial function of SGT1 and HSP90 in inflammasome activity links mammalian and plant innate immune responses. Nat Immunol 2007; 8: 497–503.
Petrilli V, Papin S, Dostert C Mayor A, Martinon F, Tschopp J . Activation of the NALP3 inflammasome is triggered by low intracellular potassium concentration. Cell Death Differ 2007; 14: 1583–1589.
Kahlenberg JM, Dubyak GR . Mechanisms of caspase-1 activation by P2X7 receptor-mediated K+ release. Am J Physiol Cell Physiol 2004; 286: C1100–C1108.
Laliberte RE, Eggler J, Gabel CA . ATP treatment of human monocytes promotes caspase-1 maturation and externalization. J Biol Chem 1999; 274: 36944–36951.
Kanneganti TD, Lamkanfi M, Kim YG, Chen G, Park JH, Franchi L et al. Pannexin-1-mediated recognition of bacterial molecules activates the cryopyrin inflammasome independent of Toll-like receptor signaling. Immunity 2007; 26: 433–443.
Locovei S, Bao L, Dahl G . Pannexin 1 in erythrocytes: function without a gap. Proc Natl Acad Sci USA 2006; 103: 7655–7659.
Cruz CM, Rinna A, Forman HJ, Ventura AL, Persechini PM, Ojcius OM et al. ATP activates a reactive oxygen species-dependent oxidative stress response and secretion of proinflammatory cytokines in macrophages. J Biol Chem 2007; 282: 2871–2879.
Zhou R, Tardivel A, Thorens B, Choi I, Tschopp J . Thioredoxin-interacting protein links oxidative stress to inflammasome activation. Nat Immunol 2010; 11: 136–140.
Shio MT, Eisenbarth SC, Savaria M, Vinet AF, Bellemare MJ, Harder KW et al. Malarial hemozoin activates the NLRP3 inflammasome through Lyn and Syk kinases. PLoS Pathog 2009; 5: e1000559.
Hornung V, Bauernfeind F, Halle A, Samstad EO, Kono H, Rock KL et al. Silica crystals and aluminum salts activate the NALP3 inflammasome through phagosomal destabilization. Nat Immunol 2008; 9: 847–856.
Rutault K, Alderman C, Chain BM, Katz DR . Reactive oxygen species activate human peripheral blood dendritic cells. Free Radic Biol Med 1999; 26: 232–238.
Dinarello CA . Immunological and inflammatory functions of the interleukin-1 family. Annu Rev Immunol 2009; 27: 519–550.
Hawkins PN, Lachmann HJ, McDermott MF . Interleukin-1-receptor antagonist in the Muckle–Wells syndrome. N Engl J Med 2003; 348: 2583–2584.
Chen CJ, Shi Y, Hearn A, Fitzgerald K, Golenbock D, Reed G et al. MyD88-dependent IL-1 receptor signaling is essential for gouty inflammation stimulated by monosodium urate crystals. J Clin Invest 2006; 116: 2262–2271.
Bauernfeind FG, Horvath G, Stutz A, Alnemri ES, MacDonald K, Speert D et al. Cutting edge: NF-kappaB activating pattern recognition and cytokine receptors license NLRP3 inflammasome activation by regulating NLRP3 expression. J Immunol 2009; 183: 787–791.
Guarda G, Dostert C, Staehli F, Cabalzar K, Castillo R, Tardivel A et al. T cells dampen innate immune responses through inhibition of NLRP1 and NLRP3 inflammasomes. Nature 2009; 460: 269–273.
Kool M, Petrilli V, de Smedt T, Rolaz A, Hammad H, van Nimwegen M et al. Cutting edge: alum adjuvant stimulates inflammatory dendritic cells through activation of the NALP3 inflammasome. J Immunol 2008; 181: 3755–3759.
Sutterwala FS, Ogura Y, Szczepanik M, Lara-Tejero M, Lichten-Berger GS, Grant EP et al. Critical role for NALP3/CIAS1/Cryopyrin in innate and adaptive immunity through its regulation of caspase-1. Immunity 2006; 24: 317–327.
Meylan E, Tschopp J, Karin M . Intracellular pattern recognition receptors in the host response. Nature 2006; 442: 39–44.
Sugawara S, Uehara A, Nochi T, Yamaguchi T, Ueda H, Sugiyama A et al. Neutrophil proteinase 3-mediated induction of bioactive IL-18 secretion by human oral epithelial cells. J Immunol 2001; 167: 6568–6575.
Puren AJ, Fantuzzi G, Dinarello CA . Gene expression, synthesis, and secretion of interleukin 18 and interleukin 1beta are differentially regulated in human blood mononuclear cells and mouse spleen cells. Proc Natl Acad Sci USA 1999; 96: 2256–2261.
Gattorno M, Tassi S, Carta S, Delfino L, Ferlito F, Pelagatti MA et al. Pattern of interleukin-1beta secretion in response to lipopolysaccharide and ATP before and after interleukin-1 blockade in patients with CIAS1 mutations. Arthritis Rheum 2007; 56: 3138–3148.
Shinkai K, McCalmont TH, Leslie KS . Cryopyrin-associated periodic syndromes and autoinflammation. Clin Exp Dermatol 2008; 33: 1–9.
Hoffman HM, Mueller JL, Broide DH, Wanderer AA, Kolodner RD . Mutation of a new gene encoding a putative pyrin-like protein causes familial cold autoinflammatory syndrome and Muckle–Wells syndrome. Nat Genet 2001; 29: 301–305.
Aganna E, Martinon F, Hawkins PN, Ross JB, Swan DC, Booth DR et al. Association of mutations in the NALP3/CIAS1/PYPAF1 gene with a broad phenotype including recurrent fever, cold sensitivity, sensorineural deafness, and AA amyloidosis. Arthritis Rheum 2002; 46: 2445–2452.
Touitou I, Lesage S, McDermott M, Cuisset L, Hoffman H, Dode C et al. Infevers: an evolving mutation database for auto-inflammatory syndromes. Hum Mutat 2004; 24: 194–198.
Masters SL, Simon A, Aksentijevich I, Kastner DL . Horror autoinflammaticus: the molecular pathophysiology of autoinflammatory disease (*). Annu Rev Immunol 2009; 27: 621–668.
Neven B, Callebaut I, Prieur AM, Feldmann J, Bodemer C, Lepore L et al. Molecular basis of the spectral expression of CIAS1 mutations associated with phagocytic cell-mediated autoinflammatory disorders CINCA/NOMID, MWS, and FCU. Blood 2004; 103: 2809–2815.
Goldbach-Mansky R, Dailey NJ, Canna SW, Gelabert A, Jones J, Rubin BI et al. Neonatal-onset multisystem inflammatory disease responsive to interleukin-1beta inhibition. N Engl J Med 2006; 355: 581–592.
Meng G, Zhang F, Fuss I, Kitani A, Strober W . A mutation in the NLRP3 gene causing inflammasome hyperactivation potentiates Th17 cell-dominant immune responses. Immunity 2009; 30: 860–874.
Aksentijevich I, Remmers EF, Goldbach-Mansky R, Reiff A, Kastner DL . Mutational analysis in neonatal-onset multisystem inflammatory disease: comment on the articles by Frenkel et al and Saito et al. Arthritis Rheum 2006; 54: 2703–2704; author reply 2704–2705.
Brydges SD, Mueller JL, McGeough MD, Pena CA, Misaghi A, Gandhi C et al. Inflammasome-mediated disease animal models reveal roles for innate but not adaptive immunity. Immunity 2009; 30: 875–887.
Fujisawa A, Kambe N, Saito M, Nishikomori R, Tanizaki H, Kanazawa N et al. Disease-associated mutations in CIAS1 induce cathepsin B-dependent rapid cell death of human THP-1 monocytic cells. Blood 2007; 109: 2903–2911.
Saito M, Nishikomori R, Kambe N, Fujisawa A, Tanizaki H, Takeichi K et al. Disease-associated CIAS1 mutations induce monocyte death, revealing low-level mosaicism in mutation-negative cryopyrin-associated periodic syndrome patients. Blood 2008; 111: 2132–2141.
Kambe N, Satoh T, Tanizaki H, Fujisawa A, Saito MK Nishikomori R . Enhanced NF-kappaB activation with an inflammasome activator correlates with activity of autoinflammatory disease associated with NLRP3 mutations outside of exon 3: comment on the article by Jeru et al. Arthritis Rheum 2010; 62: 3123–3124; author reply 3124–3125 .
Jeru I, Marlin S, Le Borgne G, Cochet E, Normand S, Duquesnoy P et al. Functional consequences of a germline mutation in the leucine-rich repeat domain of NLRP3 identified in an atypical autoinflammatory disorder. Arthritis Rheum 2010; 62: 1176–1185.
Arostegui JI, Lopez Saldaña MD, Pascal M, Clemente D, Aymerich M, Balaguer F et al. A somatic NLRP3 mutation as a cause of a sporadic case of chronic infantile neurologic, cutaneous, articular syndrome/neonatal-onset multisystem inflammatory disease: novel evidence of the role of low-level mosaicism as the pathophysiologic mechanism underlying mendelian inherited diseases. Arthritis Rheum 2010; 62: 1158–1166.
Villani AC, Lemire M, Fortin G, Louis E, Silverberg MS, Collette C et al. Common variants in the NLRP3 region contribute to Crohn's disease susceptibility. Nat Genet 2009; 41: 71–76.
Cummings JR, Cooney RM Clarke G, Beckly J, Geremia A, Pathan S, et al. The genetics of NOD-like receptors in Crohn's disease. Tissue Antigens 2010; 76: 48–56.
Schoultz I, Verma D, Halfvarsson J, Torkvist L, Fredrikson M, Sjöqvist U et al. Combined polymorphisms in genes encoding the inflammasome components NALP3 and CARD8 confer susceptibility to Crohn's disease in Swedish men. Am J Gastroenterol 2009; 104: 1180–1188.
Lewis GJ, Massey DC Zhang H Bredin F Tremelling M Lee JC, et al. Genetic association between NLRP3 variants and Crohn's disease does not replicate in a large UK panel. Inflamm Bowel Dis 2010 Oct 25. [Epub ahead of print] DOI: 10.1002/ibd.21499.
Aksentijevich I, Nowak M, Mallah M, Chae JJ, Watford WT, Hofmann SR et al. De novo CIAS1 mutations, cytokine activation, and evidence for genetic heterogeneity in patients with neonatal-onset multisystem inflammatory disease (NOMID): a new member of the expanding family of pyrin-associated autoinflammatory diseases. Arthritis Rheum 2002; 46: 3340–3348.
Jeru I, Duquesnoy P, Fernandes-Alnemri T, Cochet E, Yu JW, Lackmy-Port-Lis M et al. Mutations in NALP12 cause hereditary periodic fever syndromes. Proc Natl Acad Sci USA 2008; 105: 1614–1619.
Lich JD, Williams KL, Moore CB, Arthur JC, Davis BK, Taxman DJ et al. Monarch-1 suppresses non-canonical NF-kappaB activation and p52-dependent chemokine expression in monocytes. J Immunol 2007; 178: 1256–1260.
Pinheiro AS, Proell M, Eibl C, Page R, Schwarzenbacher R, Peti W et al. Three-dimensional structure of the NLRP7 pyrin domain: insight into pyrin–pyrin-mediated effector domain signaling in innate immunity. J Biol Chem 2010; 285: 27402–27410.
Kinoshita T, Wang Y, Hasegawa M, Imamura R, Suda T . PYPAF3, a PYRIN-containing APAF-1-like protein, is a feedback regulator of caspase-1-dependent interleukin-1beta secretion. J Biol Chem 2005; 280: 21720–21725.
Deveault C, Qian JH, Chebaro W, Ao A, Gilbert L, Mehio A . NLRP7 mutations in women with diploid androgenetic and triploid moles: a proposed mechanism for mole formation. Hum Mol Genet 2009; 18: 888–897.
Yu JW, Fernandes-Alnemri T, Datta P, Wu J, Juliana C, Solorzano L et al. Pyrin activates the ASC pyroptosome in response to engagement by autoinflammatory PSTPIP1 mutants. Mol Cell 2007; 28: 214–227.
Chae JJ, Komarow HD, Cheng J, Wood G, Raben N, Liu PP et al. Targeted disruption of pyrin, the FMF protein, causes heightened sensitivity to endotoxin and a defect in macrophage apoptosis. Mol Cell 2003; 11: 591–604.
Jeru I, Hayrapetyan H, Duquesnoy P, Sarkisian T, Amselem S . PYPAF1 nonsense mutation in a patient with an unusual autoinflammatory syndrome: role of PYPAF1 in inflammation. Arthritis Rheum 2006; 54: 508–514.
Master SS, Rampini SK, Davis AS, Keller C, Ehlers S, Springer B et al. Mycobacterium tuberculosis prevents inflammasome activation. Cell Host Microbe 2008; 3: 224–232.
Pontillo A, Brandao LA, Guimaraes RL, Segat L, Athanasakis E, Crovella S . A 3'UTR SNP in NLRP3 gene is associated with susceptibility to HIV-1 infection. J Acquir Immune Defic Syndr 2010; 54: 236–240.
Allen IC, Scull MA, Moore CB, Holl EK, McElvania-TeKippe E, Taxman DJ et al. The NLRP3 inflammasome mediates in vivo innate immunity to influenza A virus through recognition of viral RNA. Immunity 2009; 30: 556–565.
Ichinohe T, Lee HK, Ogura Y, Flavell R, Iwasaki A . Inflammasome recognition of influenza virus is essential for adaptive immune responses. J Exp Med 2009; 206: 79–87.
Thomas PG, Dash P, Aldridge JR, Jr, Ellebedy AH, Reynolds C, Funk AJ et al. The intracellular sensor NLRP3 mediates key innate and healing responses to influenza A virus via the regulation of caspase-1. Immunity 2009; 30: 566–575.
Lev-Sagie A, Prus D, Linhares IM, Lavy Y, Ledger WJ, Witkin SS . Polymorphism in a gene coding for the inflammasome component NALP3 and recurrent vulvovaginal candidiasis in women with vulvar vestibulitis syndrome. Am J Obstet Gynecol 2009; 200: 303–306.
Witkin SS, Bierhals K, Linhares I, Normand N, Dieterle S, Neuer A . Genetic polymorphism in an inflammasome component, cervical mycoplasma detection and female infertility in women undergoing in vitro fertilization. J Reprod Immunol 2010; 84: 171–175.
Wang W, Stassen FR, Surcel HMÖ hman H, Tiitinen A, Paavonen J et al. Analyses of polymorphisms in the inflammasome-associated NLRP3 and miRNA-146A genes in the susceptibility to and tubal pathology of Chlamydia trachomatis infection. Drugs Today (Barc) 2009; 45 Suppl B: 95–103.
McGonagle D, McDermott MF . A proposed classification of the immunological diseases. PLoS Med 2006; 3: e297.
Sutton C, Brereton C, Keogh B, Mills KH, Lavelle EC . A crucial role for interleukin (IL)-1 in the induction of IL-17-producing T cells that mediate autoimmune encephalomyelitis. J Exp Med 2006; 203: 1685–1691.
Chung Y, Chang SH, Martinez GJ, Yang XO, Nurieva R, Kang HS et al. Critical regulation of early Th17 cell differentiation by interleukin-1 signaling. Immunity 2009; 30: 576–587.
Acosta-Rodriguez EV, Napolitani G, Lanzavecchia A, Sallusto F . Interleukins 1beta and 6 but not transforming growth factor-beta are essential for the differentiation of interleukin 17-producing human T helper cells. Nat Immunol 2007; 8: 942–949.
van Beelen AJ, Zelinkova Z, Taanman-Kueter EW, Muller FJ, Hommes DW, Zaat SA et al. Stimulation of the intracellular bacterial sensor NOD2 programs dendritic cells to promote interleukin-17 production in human memory T cells. Immunity 2007; 27: 660–669.
Nistala K, Moncrieffe H, Newton KR, Varsani H, Hunter P, Wedderburn LR . Interleukin-17-producing T cells are enriched in the joints of children with arthritis, but have a reciprocal relationship to regulatory T cell numbers. Arthritis Rheum 2008; 58: 875–887.
Tzartos JS, Friese MA, Craner MJ, Palace J, Newcombe J, Esiri MM et al. Interleukin-17 production in central nervous system-infiltrating T cells and glial cells is associated with active disease in multiple sclerosis. Am J Pathol 2008; 172: 146–155.
Pernis AB . Th17 cells in rheumatoid arthritis and systemic lupus erythematosus. J Intern Med 2009; 265: 644–652.
Lubberts E . Th17 cytokines and arthritis. Semin Immunopathol 2010; 32: 43–53.
van den Berg WB, Miossec P . IL-17 as a future therapeutic target for rheumatoid arthritis. Nat Rev Rheumatol 2009; 5: 549–553.
Evans HG, Gullick NJ, Kelly S, Pitzalis C, Lord GM, Kirkham BW et al. In vivo activated monocytes from the site of inflammation in humans specifically promote Th17 responses. Proc Natl Acad Sci USA 2009; 106: 6232–6237.
Day TG, Ramanan AV, Hinks A, Lamb R, Packham J, Wise C et al. Autoinflammatory genes and susceptibility to psoriatic juvenile idiopathic arthritis. Arthritis Rheum 2008; 58: 2142–2146.
Agarwal S, Misra R, Aggarwal A . Interleukin 17 levels are increased in juvenile idiopathic arthritis synovial fluid and induce synovial fibroblasts to produce proinflammatory cytokines and matrix metalloproteinases. J Rheumatol 2008; 35: 515–519.
Rosengren S, Hoffman HM, Bugbee W, Boyle DL . Expression and regulation of cryopyrin and related proteins in rheumatoid arthritis synovium. Ann Rheum Dis 2005; 64: 708–714.
Kastbom A, Verma D, Eriksson P, Skogh T, Wingren G, Soderkvist P . Genetic variation in proteins of the cryopyrin inflammasome influences susceptibility and severity of rheumatoid arthritis (the Swedish TIRA project). Rheumatology (Oxford) 2008; 47: 415–417.
Hoffman HM, Gregory SG, Mueller JL, Tresierras M, Broide DH, Kolodner RD . Fine structure mapping of CIAS1: identification of an ancestral haplotype and a common FCAS mutation, L353P. Hum Genet 2003; 112: 209–216.
McGovern DP, Butler H, Ahmad T, Paolucci M, van Heel DA, Negoro K et al. TUCAN (CARD8) genetic variants and inflammatory bowel disease. Gastroenterology 2006; 131: 1190–1196.
Hitomi Y, Ebisawa M, Tomikawa M, Imai T, Komata T, Hirota T et al. Associations of functional NLRP3 polymorphisms with susceptibility to food-induced anaphylaxis and aspirin-induced asthma. J Allergy Clin Immunol 2009; 124: 779–785.
Kambe N, Nakamura Y, Saito M, Nishikomori R . The inflammasome, an innate immunity guardian, participates in skin urticarial reactions and contact hypersensitivity. Allergol Int 2010; 59: 105–113.
Nakamura Y, Kambe N, Saito M, Nishikomori R, Kim YG, Murakami M et al. Mast cells mediate neutrophil recruitment and vascular leakage through the NLRP3 inflammasome in histamine-independent urticaria. J Exp Med 2009; 206: 1037–1046.
Jin Y, Mailloux CM, Gowan K, Riccardi SL, LaBerge G, Bennett DC et al. NALP1 in vitiligo-associated multiple autoimmune disease. N Engl J Med 2007; 356: 1216–1225.
Jin Y, Birlea SA, Fain PR, Spritz RA . Genetic variations in NALP1 are associated with generalized vitiligo in a Romanian population. J Invest Dermatol 2007; 127: 2558–2562.
Magitta NF, Boe Wolff AS, Johansson S, Skinningsrud B, Lie BA, Myhr KM et al. A coding polymorphism in NALP1 confers risk for autoimmune Addison's disease and type 1 diabetes. Genes Immun 2009; 10: 120–124.
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 2007; 447: 661–678.
Todd JA, Walker NM, Cooper JD, Smyth DJ, Downes K, Plagnol V et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet 2007; 39: 857–864.
Omi T, Kumada M, Kamesaki T, Okuda H, Munkhtulga L, Yanagisawa Y et al. An intronic variable number of tandem repeat polymorphisms of the cold-induced autoinflammatory syndrome 1 (CIAS1) gene modifies gene expression and is associated with essential hypertension. Eur J Hum Genet 2006; 14: 1295–1305.
Rawat R, Cohen TV, Ampong B, Francia D, Henriques-Pons A, Hoffman EP et al. Inflammasome up-regulation and activation in dysferlin-deficient skeletal muscle. Am J Pathol 2010; 176: 2891–2900.
Masters SL, Dunne A, Subramanian SL, Hull RL, Tannahill GM, Sharp FA et al. Activation of the NLRP3 inflammasome by islet amyloid polypeptide provides a mechanism for enhanced IL-1beta in type 2 diabetes. Nat Immunol 2010; 11: 897–904.
Tsutsui H, Imamura M, Fujimoto J, Nakanishi K . The TLR4/TRIF-mediated activation of NLRP3 inflammasome underlies endotoxin-induced liver injury in mice. Gastroenterol Res Pract 2010; 2010: 641865.
Gu LQ, Li FY, Zhao L, Liu Y, Chu Q, Zang XX et al. Association of XIAP and P2X7 receptor expression with lymph node metastasis in papillary thyroid carcinoma. Endocrine 2010; 38: 276–282.
Ravaglia G, Paola F, Maioli F, Martelli M, Montesi F, Bastagli L et al. Interleukin-1beta and interleukin-6 gene polymorphisms as risk factors for AD: a prospective study. Exp Gerontol 2006; 41: 85–92.
Forlenza OV, Diniz BS, Nunes PV, Memória CM, Yassuda MS, Gattaz WF et al. Increased serum IL-1beta level in Alzheimer's disease and mild cognitive impairment. Dement Geriatr Cogn Disord 2009; 28: 507–512.
Halle A, Hornung V, Petzold GC, Stewart CR, Monks BG, Reinheckel T et al. The NALP3 inflammasome is involved in the innate immune response to amyloid-beta. Nat Immunol 2008; 9: 857–865.
Acknowledgements
We thank Lucy Robinson for critically reviewing the manuscript. This work was supported by Agency for Science, Technology and Research (A*STAR) of Singapore.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Conforti-Andreoni, C., Ricciardi-Castagnoli, P. & Mortellaro, A. The inflammasomes in health and disease: from genetics to molecular mechanisms of autoinflammation and beyond. Cell Mol Immunol 8, 135–145 (2011). https://doi.org/10.1038/cmi.2010.81
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cmi.2010.81
Keywords
This article is cited by
-
Allergen-induced CD11c + dendritic cell pyroptosis aggravates allergic rhinitis
Cell Communication and Signaling (2023)
-
An Atypical Autoinflammatory Disease Due to an LRR Domain NLRP3 Mutation Enhancing Binding to NEK7
Journal of Clinical Immunology (2022)
-
CRID3, a blocker of apoptosis associated speck like protein containing a card, ameliorates murine spinal cord injury by improving local immune microenvironment
Journal of Neuroinflammation (2020)
-
Inhibition of NLRP3 inflammasome activation by cell-permeable stapled peptides
Scientific Reports (2019)
-
Inflammasomes, the eye and anti-inflammasome therapy
Eye (2018)