Abstract
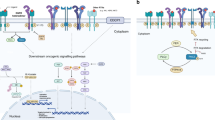
Lung cancer is one of the leading causes of death from cancer worldwide, with a poor prognosis in advanced cases. In the past decade, epidermal growth factor receptor (EGFR) inhibitors have shown significant efficacy towards treatment for EGFR mutant lung cancer. Expanding our knowledge of oncogenic EGFR signaling pathways is therefore of highly importance for the cancer field. Recently it has been proposed that mutant EGFR transcriptionally silences the TET1 (ten–eleven translocation methylcytosine dioxygenase 1) gene in cellular and animal models of lung cancer. Since TET1 is a known DNA demethylase, EGFR-mediated TET1 silencing therefore downregulates demethylation of tumor suppressor genes, which then leads to tumor growth inhibition, potentiating the role of TET1 as a tumor suppressor gene in NSCLC. In our study, we examined the role of EGFR-TET1 silencing in NSCLC patient samples. By independently analyzing the TCGA (The Cancer Genome Atlas) NSCLC data set as well as a cohort of patient samples from our hospital and a data set from publicly deposited databases, we did not observe the aforementioned mutant EGFR silencing of TET1. Conversely, in our cohort, TET1 expression levels were significantly elevated in EGFR mutant samples (P=0.007). Patients with higher TET1 levels showed a trend of better response rates to EGFR inhibitors compared to low TET1 staining levels, although the result was not significant (P=0.08). Furthermore, we did not observe a correlation between TET1 expression levels and patient survival. We conclude that while oncogenic EGFR suppression of TET1 is established in cellular and animal models of lung cancer, its role in patient outcome and prognosis remains inconclusive and warrants further investigation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Woodard GA, Jones KD, Jablons DM . Lung cancer staging and prognosis. Cancer Treat Res 2016; 170: 47–75.
Ciardiello F, Tortora G . EGFR antagonists in cancer treatment. N Engl J Med 2008; 358: 1160–1174.
Burotto M, Manasanch EE, Wilkerson J, Fojo T . Gefitinib and erlotinib in metastatic non-small cell lung cancer: a meta-analysis of toxicity and efficacy of randomized clinical trials. Oncologist 2015; 20: 400–410.
Sequist LV, Yang JC, Yamamoto N, O'Byrne K, Hirsh V, Mok T et al. Phase III study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J Clin Oncol 2013; 31: 3327–3334.
Janne PA, Yang JC, Kim DW, Planchard D, Ohe Y, Ramalingam SS et al. AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N Engl J Med 2015; 372: 1689–1699.
Hatzimichael E, Crook T . Cancer epigenetics: new therapies and new challenges. J Drug Deliv 2013; 2013: 529312.
Herman JG, Baylin SB . Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 2003; 349: 2042–2054.
Aribi A, Borthakur G, Ravandi F, Shan J, Davisson J, Cortes J et al. Activity of decitabine, a hypomethylating agent, in chronic myelomonocytic leukemia. Cancer 2007; 109: 713–717.
Fenaux P, Mufti GJ, Hellstrom-Lindberg E, Santini V, Gattermann N, Germing U et al. Azacitidine prolongs overall survival compared with conventional care regimens in elderly patients with low bone marrow blast count acute myeloid leukemia. J Clin Oncol 2010; 28: 562–569.
Kumar S, Cheng X, Klimasauskas S, Mi S, Posfai J, Roberts RJ et al. The DNA (cytosine-5) methyltransferases. Nucleic Acids Res 1994; 22: 1–10.
Robertson KD . DNA methylation, methyltransferases, and cancer. Oncogene 2001; 20: 3139–3155.
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009; 324: 930–935.
Ito S, D'Alessio AC, Taranova OV, Hong K, Sowers LC, Zhang Y . Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 2010; 466: 1129–1133.
Ko M, An J, Pastor WA, Koralov SB, Rajewsky K, Rao A . TET proteins and 5-methylcytosine oxidation in hematological cancers. Immunol Rev 2015; 263: 6–21.
Lian CG, Xu Y, Ceol C, Wu F, Larson A, Dresser K et al. Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 2012; 150: 1135–1146.
Yang H, Liu Y, Bai F, Zhang JY, Ma SH, Liu J et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 2013; 32: 663–669.
Hsu CH, Peng KL, Kang ML, Chen YR, Yang YC, Tsai CH et al. TET1 suppresses cancer invasion by activating the tissue inhibitors of metalloproteinases. Cell Rep 2012; 2: 568–579.
Taylor MA, Wappett M, Delpuech O, Brown H, Chresta CM . Enhanced MAPK signaling drives ETS1-mediated induction of miR-29b leading to downregulation of TET1 and changes in epigenetic modifications in a subset of lung SCC. Oncogene 2016; 35: 4345–4357.
Forloni M, Gupta R, Nagarajan A, Sun LS, Dong Y, Pirazzoli V et al. Oncogenic EGFR represses the TET1 DNA demethylase to induce silencing of tumor suppressors in cancer cells. Cell Rep 2016; 16: 457–471.
Cancer Genome Atlas Research N. Comprehensive molecular profiling of lung adenocarcinoma. Nature 2014; 511: 543–550.
Ni K, Dansranjavin T, Rogenhofer N, Oeztuerk N, Deuker J, Bergmann M et al. TET enzymes are successively expressed during human spermatogenesis and their expression level is pivotal for male fertility. Hum Reprod 2016; 31: 1411–1424.
Seo JS, Ju YS, Lee WC, Shin JY, Lee JK, Bleazard T et al. The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res 2012; 22: 2109–2119.
Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK . Non-small-cell lung cancers: a heterogeneous set of diseases. Nat Rev Cancer 2014; 14: 535–546.
Zhang J, Fujimoto J, Zhang J, Wedge DC, Song X, Zhang J et al. Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science 2014; 346: 256–259.
de Bruin EC, McGranahan N, Mitter R, Salm M, Wedge DC, Yates L et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 2014; 346: 251–256.
Midha A, Dearden S, McCormack R . EGFR mutation incidence in non-small-cell lung cancer of adenocarcinoma histology: a systematic review and global map by ethnicity (mutMapII). Am J Cancer Res 2015; 5: 2892–2911.
Dearden S, Stevens J, Wu YL, Blowers D . Mutation incidence and coincidence in non small-cell lung cancer: meta-analyses by ethnicity and histology (mutMap). Ann Oncol 2013; 24: 2371–2376.
Acknowledgements
We acknowledge the technical services provided by Clinical Medicine Research Laboratory of National Yang-Ming University Hospital, especially the technical support of Ping-Liang Hung, Peng-Chih Chiu, Chia-Yuan Huang for their assistance.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Lai, JI., Lai, YC., Chen, YC. et al. Clinical analysis of NSCLC patients reveals lack of association between EGFR mutation and TET1 downregulation. Cancer Gene Ther 24, 373–380 (2017). https://doi.org/10.1038/cgt.2017.26
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cgt.2017.26
This article is cited by
-
Advances in the DNA methylation hydroxylase TET1
Biomarker Research (2021)
-
DNA methylation instability by BRAF-mediated TET silencing and lifestyle-exposure divides colon cancer pathways
Clinical Epigenetics (2019)