Abstract
The saturated medium-chain fatty-acid lauric acid (LA) has been associated to certain health-promoting benefits of coconut oil intake, including the improvement of the quality of life in breast cancer patients during chemotherapy. As it concerns the potential to hamper tumor growth, LA was shown to elicit inhibitory effects only in colon cancer cells. Here, we provide novel insights regarding the molecular mechanisms through which LA triggers antiproliferative and pro-apoptotic effects in both breast and endometrial cancer cells. In particular, our results demonstrate that LA increases reactive oxygen species levels, stimulates the phosphorylation of EGFR, ERK and c-Jun and induces the expression of c-fos. In addition, our data evidence that LA via the Rho-associated kinase-mediated pathway promotes stress fiber formation, which exerts a main role in the morphological changes associated with apoptotic cell death. Next, we found that the increase of p21Cip1/WAF1 expression, which occurs upon LA exposure in a p53-independent manner, is involved in the apoptotic effects prompted by LA in both breast and endometrial cancer cells. Collectively, our findings may pave the way to better understand the anticancer action of LA, although additional studies are warranted to further corroborate its usefulness in more comprehensive therapeutic approaches.
Similar content being viewed by others
Introduction
Fatty acids are acyclic carboxylic acids with aliphatic tails of different lengths. Based on their carbon atom chain length, fatty acids are classified into the following three groups: short-chain fatty acids with <6 carbon atoms, medium-chain fatty acids (MCFA) and long-chain fatty acids that contain 6–12 carbons and >12 carbons, respectively.1 Fatty acids are major components of triacylglycerols, phospholipids and other complex lipids, therefore representing main contributors to dietary fat in humans.2 Plant oils like palm, coconut and olive oils, nuts, seeds and seed oils, cocoa butter and animal-derived fats as lard, tallow and butter, are rich of fatty acids that are important components of cell membranes and essential sources of energy.2 Previous studies have demonstrated that fatty acids are also involved in diverse transduction pathways, in gene transcription and relevant biological events as cell metabolism, inflammation, apoptosis and production of bioactive lipid mediators, thus contributing to multiple patho-physiological responses.2–7
Lauric acid (LA), which is a saturated MCFA with 12 carbon atoms and the primary fatty acid of coconut oil, has been associated with certain health benefits of coconut oil intake.8–10 LA is also contained in plant oils, fruits, seeds and in breast milk.11,12 LA has been shown to elicit diverse actions in various tissues, including a potent antimicrobial property.8 For instance, LA and the derivative monolaurin were reported to destroy cell membranes of gram-positive bacteria and lipid-coated viruses, to interfere with main cellular responses as the activation of transduction cascades and gene transcription, to stabilize cell membranes toward the prevention of bacterial resistance.8 In addition, LA promoted inflammatory processes activating the nuclear factor-κB transcription factor as well as stimulating the expression of cyclooxygenase-2 and pro-inflammatory cytokines.13 LA was also associated with beneficial effects on the cardiovascular system due to its ability to increase the high-density lipoproteins14 and to reduce the blood pressure and heart rate in both normotensive and hypertensive rats.15 Moreover, LA prevented the prostatic hyperplasia induced by testosterone in rats,16 triggered apoptosis in colon cancer cells through oxidative stress17 and improved the sensitization of the EGFR inhibitor cetuximab in KRAS/BRAF mutated colorectal cancer cells.18 It is worth mentioning that the consumption of virgin coconut oil during chemotherapy improved the global quality of life in patients with breast cancer.19
Here, we show for the first time that LA elicits antiproliferative and pro-apoptotic effects in breast and endometrial cancer cells promoting the generation of reactive oxygen species (ROS), the activation of transduction pathways and gene expression changes. In particular, the upregulation of the cyclin-dependent kinase inhibitor p21Cip1/WAF1 upon LA exposure was found to be required for its anticancer properties. Our findings shed new light on the molecular mechanisms through which LA induces antiproliferative and pro-apoptotic responses in both breast and endometrial cancer cells toward its usefulness in more comprehensive therapeutic approaches.
Results
LA inhibits cancer cell viability
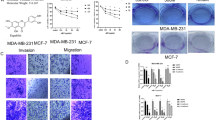
On the basis of previous findings showing that MCFAs may elicit apoptosis in certain cancer cells17,20 and considering that in our recent investigation LA exerted antiproliferative activity in diverse types of tumor cells,21 we began the present study evaluating whether LA (Figure 1a) and a further MCFA namely capric acid (CA) (Figure 1b) may affect the viability of SkBr3 breast and Ishikawa endometrial cancer cells, which were used as model system. Only LA inhibited the viability of both cancer cell types (Figures 1c and d) without altering the growth of MCF-10A normal breast epithelial cells (Figure 1e), thus suggesting its specific potential to trigger antiproliferative effects in malignant cells.
Lauric acid inhibits the proliferation of breast and endometrial cancer cells. (a, b) Chemical structures of lauric acid (LA) and capric acid (CA). (b–d) MTT growth assays in SkBr3 (c), Ishikawa (d) and MCF-10A (e) cells treated for 48 h with vehicle (−) or increasing concentrations of LA and CA, as indicated. Cell viability is expressed as the percentage of cells upon treatments respect to cells treated with vehicle. Values shown are mean±S.D. of three independent experiments performed in triplicate. (○) indicates P<0.05 for cells receiving vehicle versus treatments.
LA triggers ROS generation and EGFR, ERK and c-Jun phosphorylation
To evaluate the molecular mechanisms involved in the ability of LA to lower cancer cell viability, we ascertained that LA triggers the phosphorylation of EGFR, ERK and c-Jun in both SkBr3 and Ishikawa cells (Figures 2a and b). These responses were no longer observed in the presence of the EGFR inhibitor (AG) (Figures 2c and d), whereas ERK activation by LA was abolished using the MEK inhibitor (PD) and the Rho-associated kinase (ROCK) inhibitor (Y) but it still persisted using the JNK inhibitor (SP) (Figures 2c and d). The phosphorylation of c-Jun by LA was prevented in the presence of PD or SP, but not using the ROCK inhibitor Y (Figures 2c and d). Reminiscing previous data on the ability of LA to induce ROS levels in colon cancer cells,17 we found that LA triggers ROS generation in our model system, yet this response was no longer evident using the ROS scavenger N-acetyl-l-cysteine (NAC, Figure 3a). Thereafter, we established that the phosphorylation of EGFR, ERK and c-Jun upon LA exposure is strictly dependent on ROS generation, as ascertained using NAC in both cell types (Figures 3b and c).
Lauric acid triggers rapid responses in breast and endometrial cancer cells. (a, b) Phosphorylation of EGFR, ERK1/2 and c-Jun in SkBr3 (a) and Ishikawa (b) cells treated with vehicle (−) and 100 μM LA, as indicated. (c, d) EGFR, ERK1/2 and c-Jun activation in SkBr3 (c) and Ishikawa (d) cells treated for 60 min with vehicle or 100 μM LA alone or in combination with 10 μM EGFR inhibitor AG1478 (AG), 10 μM MEK inhibitor PD98089 (PD), 1 μM JNK inhibitor SP 600125 (SP) and 10 μM ROCK inhibitor Y-27632 (Y). EGFR, ERK2 and c-Jun were used as loading controls for pEGFR, pERK1/2 and pc-Jun, respectively. Results shown are representative of at least two independent experiments.
ROS generation by lauric acid is involved in the activation of transduction signaling observed in breast and endometrial cancer cells. (a) ROS production determined as DCF fluorescence in SkBr3 and Ishikawa cells treated for 60 min with vehicle (−) or 100 μM LA alone or in combination with 300 μM free radical scavenger NAC. DCF fluorescence obtained in cells treated with vehicle was set as onefold induction upon which ROS levels induced by treatments were calculated. Data shown are the mean±S.D. of three independent experiments performed in triplicate. (○) indicates P<0.05 for cells receiving vehicle versus treatments. EGFR, ERK1/2 and c-Jun activation in SkBr3 (b) and Ishikawa (c) cells treated for 60 min with vehicle or 100 μM LA alone or in combination with 300 μM NAC. EGFR, ERK2 and c-Jun were used as loading controls for pEGFR, pERK1/2 and pc-Jun, respectively. Results shown are representative of at least two independent experiments.
LA induces gene expression changes
Then, we assessed the expression levels of well-known cell cycle regulators as the member of the AP1 transcription factor complex namely c-fos, the tumor suppressor p53 and the cyclin-dependent kinase inhibitor p21Cip1/WAF1. In both SkBr3 and Ishikawa cells, LA upregulated the mRNA expression of c-fos and p21Cip1/WAF1, without altering the levels of p53 (Figures 4a and b). In addition, LA transactivated the AP1-luc responsive collagenase promoter construct that was transiently transfected in SkBr3 and Ishikawa cells and stimulated the transcriptional activity of reporter plasmids containing the c-fos and p21Cip1/WAF1 promoter sequences (Figures 4c and d). According to the results obtained in real-time PCR, LA did not modify the p53 protein levels, whereas it increased c-fos and p21Cip1/WAF1 protein expression in both cell types (Figures 4e and f). We next ascertained that the ROS scavenger NAC, the EGFR inhibitor AG, the MEK inhibitor PD and the ROCK inhibitor Y prevent c-fos induction by LA (Figures 5a and b). Likewise, these compounds together with the JNK inhibitor SP repressed the increase of p21Cip1/WAF1 protein levels elicited by LA (Figures 5a and b). As the upregulation of p21Cip1/WAF1 protein levels was no longer evident transfecting a dominant-negative form of c-fos (DN/c-fos) in both SkBr3 and Ishikawa cells (Figures 5c and d), we ascertained by chromatin immunoprecipitation assay that LA induces the recruitment of c-fos to the AP1 site located within the p21Cip1/WAF1 promoter sequence (Figure 5e). Overall, these data indicate that c-fos-AP1 transduction signaling is involved in the upregulation of p21Cip1/WAF1 induced by LA.
Lauric acid regulates the expression of cell cycle regulatory genes. The mRNA expression of c-fos, p53 and p21Cip1/WAF1 (p21) was evaluated by real-time PCR in SkBr3 (a) and Ishikawa (b) cells treated with vehicle (−) or 100 μM LA, as indicated. Data obtained from three independent experiments performed in triplicate were normalized to 18 S expression and shown as fold changes of mRNA expression induced by LA respect to cells treated with vehicle. Evaluation of c-fos, AP1 and p21 luciferase reporter genes in SkBr3 (c) and Ishikawa (d) cells treated for 18 h with vehicle or 100 μM LA. The luciferase activities were normalized to the internal transfection control, and values of cells receiving vehicle were set as onefold induction upon which the activity induced by treatments was calculated. Data shown are the mean±S.D. of three independent experiments performed in triplicate. (○) indicates P<0.05 for cells receiving vehicle versus treatments. c-fos, p53 and p21 protein levels in SkBr3 (e) and Ishikawa (f) cells treated with vehicle or 100 μM LA, as indicated. β-actin was used as a loading control. Results shown are representative of at least two independent experiments.
c-fos is involved in the upregulation of p21Cip1/WAF1 induced by lauric acid. Immunoblots of c-fos and p21Cip1/WAF1 (p21) in SkBr3 (a) and Ishikawa (b) cells treated for 4 h with vehicle (−) or 100 μM LA alone or in combination with 300 μM free radical scavenger NAC, 10 μM EGFR inhibitor AG1478 (AG), 10 μM MEK inhibitor PD98089 (PD), 1 μM JNK inhibitor SP 600125 (SP) and 10 μM ROCK inhibitor Y-27632 (Y). The expression vector encoding for a dominant-negative form of c-fos (DN/c-fos) blocked the upregulation of p21Cip1/WAF1 protein levels induced by 100 μM LA in SkBr3 (c) and Ishikawa (d) cells. β-actin was used as a loading control. Results shown are representative of at least two independent experiments. (e) Recruitment of c-fos induced by 100 μM LA to the AP1 site located within the p21Cip1/WAF1 promoter sequence in SkBr3 and Ishikawa cells, as indicated. In control samples non-specific IgG was used instead of the primary antibody. Each column represents the mean±S.D. of three independent experiments. (○) indicates P<0.05 for cells receiving vehicle versus treatments.
LA promotes stress fiber formation and apoptosis in cancer cells
Rho GTPases and their effectors as the Rho-associated protein kinase (ROCK) are key regulators of the cytoskeleton reorganization and the generation of the contractile force required for stress fiber formation.22 In line with the aforementioned findings regarding the capability of the ROCK inhibitor to prevent LA-induced responses, in both SkBr3 and Ishikawa cells LA promoted the formation of stress fibers in a ROCK-dependent manner as this effect was abrogated in the presence of its inhibitor (Figures 6a-f) that alone did not show any effects (data not shown). Then, we assessed that LA increases the percentage of SkBr3 (Figures 7a–c) and Ishikawa (Figures 7d-f) TdT-mediated dUTP nick-end-labeling (TUNEL)-positive cells, however this effect was prevented in the presence of the ROS scavenger NAC (Figure 7). In addition, the apoptotic effects induced by LA were blocked in the presence of the p21Cip1/WAF1 inhibitor UC2288 (Figure 7), suggesting that p21Cip1/WAF1is involved in the pro-apoptotic activity exerted by LA in breast and endometrial cancer cells.
Lauric acid promotes the formation of stress fibers. SkBr3 and Ishikawa cells were treated for 4 h with vehicle (−) (a, d) or 100 μM LA alone (b, e) or in combination with 10 μM ROCK inhibitor Y-27632 (Y) (c, f) and subjected to phalloidin staining to visualize F-actin. (g, h) Enlarged details of stress fibers shown in b and e, respectively. White arrows indicate stress fibers. Images shown are representative of 30 random fields obtained in three independent experiments. Scale bar: 12.5 μ m.
Lauric acid induces apoptotic cell death. (a, d) TdT-mediated dUTP nick-end-labeling (TUNEL) staining (green) in SkBr3 (a) and Ishikawa (d) cells treated for 18 h with vehicle (−) or 100 μM LA alone or in combination with 300 μM free radical scavenger NAC and 10 μM p21Cip1/WAF1 inhibitor UC2288, as indicated. Nuclei were stained by propidium iodide (PI, red). Magnification is indicated by bars (100 μm). Each experiment shown is representative of 20 random fields observed. (b, e) Bar graphs represent the percentage of TUNEL-positive cells upon treatments versus vehicle. Values are the mean of three independent experiments. (○) indicates P<0.05 for cells receiving vehicle versus treatments. (c, f) Efficacy of p21Cip1/WAF1 downregulation by UC2288. β-actin was used as a loading control. Results shown are representative of at least two independent experiments.
Discussion
The present study provides novel evidence regarding the molecular mechanisms through which LA elicits antiproliferative and pro-apoptotic effects in breast and endometrial cancer cells. In particular, we have ascertained that ROS generation induced by LA triggers the activation of the EGFR/ERK/AP1 transduction pathway, leading to the upregulation of p21Cip/WAF1 in a p53-independent manner.
Fatty acids are structural components of cellular membranes either alone or together with other molecules as phospholipids and triacylglycerides.23 In addition, fatty-acid oxidation occurring at the mitochondrial level plays a pivotal role in maintaining energy homeostasis during catabolic states.24 Nevertheless, fatty acids are currently no longer considered as mere membrane structure regulators or energy sources as they also influence diverse transduction signaling and cellular functions.2–7 For instance, regulating transcription factors involved in lipid metabolism and inflammation, saturated fatty acids as LA may influence the biosynthesis of cholesterol and triacylglycerols, the assembly, secretion and clearance of lipoproteins, various metabolic and inflammatory processes.23 Therefore, an increasing attention has been paid to the multifaceted role elicited by fatty acids on human health given that the amount and type of fatty acids contained in the diet are involved in the etiopathogenesis of diabetes, cancer and cardiovascular, immunity, inflammatory, renal, hepatic diseases.25 In this context, coconut oil that is one of the richest sources of saturated fatty acids as LA, has attracted interest for its potential health benefits.26–29 Furthermore, coconut oil has been shown to counteract the action of stimulatory agents in colon and mammary tumors in rats30,31 and to improve the quality of life of breast cancer patients undergoing chemotherapy.19 As it concerns LA, Fauser and co-workers17 firstly demonstrated its ability to induce apoptosis in colon cancer cells through the reduction of glutathione levels and the generation of oxidative stress. In accordance with these and other observations showing that fatty acids may induce ROS generation in diverse types of cells,32–34 we have extended these findings ascertaining that LA prompts ROS-mediated apoptosis also in breast and endometrial cancer cells through the subsequent activation of relevant transduction pathways. In this respect, it is worth mentioning that the EGFR and ERK signaling are mostly referred to as regulatory pathways of cell proliferation, migration and differentiation.35,36 Nevertheless, these two main transduction mediators can also trigger apoptotic signals especially in the context of tumor cells.35,36 For instance, EGF through the cognate receptor induced the expression of the caspase 1 enzyme and p21WAF1/CIP1 toward apoptosis and growth inhibition.35 In addition, it has been demonstrated that free radicals generated by radiation exposure may elicit the activation of the EGFR/ERK signaling in cancer cells.36 In line with these observations, we found that LA increases ROS levels that in turn trigger the EGFR/ERK transduction pathway and gene expression changes, therefore culminating in apoptotic responses in cancer cells.
Actin stress fibers have a pivotal role in many cellular functions, including cell adhesion, mobility, contraction and morphogenesis.37 Stress fibers are also required for membrane blebbing, nuclear disintegration and apoptosis.38–41 The small GTPase Rho and its main effector ROCK are involved in several cellular processes like the regulation of actin cytoskeleton, cell polarity, microtubule dynamics, gene transcription, cell cycle progression, differentiation, apoptosis and the formation of actin stress fibers.37,42–44 In this vein, ROCK was shown to mediate the generation of stress fibers that in turn trigger the p21Cip1/WAF1-dependent apoptosis upon phorbol 12-myristate 13-acetate exposure in prostate cancer cells.40 Further extending these data, our findings have determined for the first time that LA promotes in breast and endometrial cancer cells the formation of stress fibers through the ROCK transduction pathway, thus suggesting that LA might be included among the activators of the Rho/ROCK signaling.
The cyclin-dependent kinase inhibitor p21Cip1/WAF1 has an essential role in the cell cycle arrest, the transcriptional regulation, the inhibition of DNA replication, the DNA repair, the stress-induced premature senescence and the modulation of apoptosis.45–48 Numerous studies have shown that p21Cip1/WAF1 can mediate both pro- and anti-apoptotic functions depending on the type of stimulation and the cellular context.48 For instance, p21Cip1/WAF1 can prevent cells from undergoing apoptosis triggering cell cycle arrest, inactivating cyclin A/Cdk2 complexes, inhibiting the activity of procaspase 3, caspase 8 and 10, stress-activated protein kinases and apoptosis signal-regulating kinase 1.47,49 Likewise, several reports have also suggested that p21Cip1/WAF1 exerts a pro-apoptotic function under certain cellular stresses upregulating the pro-apoptotic protein Bax, activating the tumor necrosis factor family of death receptors and regulating components of the DNA repair machinery.47 It is worth mentioning that even though p21Cip1/WAF1 may represent a major p53 transcriptional target, it can promote apoptosis through both p53-dependent and independent mechanisms.47 In addition, p21Cip1/WAF1 can act as a tumor suppressor or an oncogene depending on the stimulations and the cellular context.45 In particular, various compounds eliciting an anticancer activity such as histone deacetylase inhibitors, cisplatin, phorbol 12-myristate 13-acetate and curcumin were shown to induce apoptotic cell death through the p21Cip1/WAF1 induction.40,50–54 Extending these findings, our data have ascertained that LA induces apoptosis in both breast and endometrial cancer cells upregulating the p21Cip1/WAF1 expression levels via an AP1-mediated pathway.
Overall, the present results provide novel insights into the potential of LA to activate the EGFR/ERK/AP1/p21Cip1/WAF1 transduction signaling toward antiproliferative and pro-apoptotic responses in tumor cells. Nevertheless, further experimental evidence are warranted to better define the action of LA alone or in the context of coconut oil consumption on tumor development as claimed by a current newsworthy debate.
Materials and methods
Reagents
LA, CA, NAC, Y-27632 (Y) and 2′,7′-dichlorofluorescin diacetate (DCFDA) were purchased from Sigma-Aldrich (Milan, Italy). Tyrphostin AG1478 (AG), PD98059 (PD), SP 600125 (SP) and UC2288 were obtained from Calbiochem (DBA, Milan, Italy). All compounds were dissolved in dimethyl sulfoxide except LA, CA, NAC and Y-27632 (Y), which were solubilized in water.
Cell cultures
SkBr3 breast cancer cells were obtained by ATCC, used <6 months after resuscitation and maintained in RPMI 1640 without phenol red supplemented with 10% FBS and 100 mg/ml penicillin/streptomycin (Life Technologies, Milan, Italy). Ishikawa endometrial cancer cells were obtained by D Picard (University of Geneva) and maintained in MEM supplemented with 10% FBS, 100 μg/ml penicillin/streptomycin, 2 mM L-glutamine and 1% Non-Essential Amino Acids Solution Cells (Life Technologies). Cells were switched to medium without serum the day before immunoblots and reverse transcription-PCR experiments.
Cell viability assay
Cell viability was determined by the MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide] assay, which is based on the conversion of MTT to MTT formazan by mitochondrial enzyme. Cells were seeded in quadruplicate in 96-well plates in regular growth medium and grown until 70–80% confluence. Cells were washed once they had attached and then treated with increasing concentrations of each compound for 48 h in regular medium supplemented with 1% FBS. Relative cell viability was determined by MTT assay according to the manufacturer’s protocol (Sigma-Aldrich). Mean absorbance of cells receiving vehicle (−) was set as onefold induction upon which the mean absorbance of treatments was calculated.
Plasmids, transfections and gene reporter assays
The luciferase reporter plasmid for c-fos encoding a 2.2-kb 5´ upstream fragment of human c-fos was a gift from Dr. K Nose (Hatanodai, Shinagawa-ku, Tokyo). The luciferase reporter plasmid for AP1 responsive collagen promoter was a kind gift from H Van Dam (Department of Molecular Cell Biology, Leiden University, Leiden, Netherlands). The human p21Cip1/WAF1 promoter-luciferase reporter was kindly provided by Dr T Sakai (Kyoto Prefectural University of Medicine, Kyoto, Japan). The Renilla luciferase expression vector pRL-TK (Promega, Milan, Italy) was used as internal transfection control in luciferase assays. Cells (1×105) were plated into 24-well plates with regular growth medium/well the day before transfection. Cell medium was replaced on the day of transfection with serum-free medium and transfection was performed using X-tremeGENE 9 DNA Transfection Reagent (Sigma-Aldrich) and a mixture containing 0.5 μg of each reporter plasmid and 5 ng of pRL-TK. After 6 h, treatments were added and cells were incubated for 18 h. Luciferase activity was measured using the Dual Luciferase Kit (Promega) according to the manufacturer’s recommendations. Firefly luciferase activity was normalized to the internal transfection control provided by the Renilla luciferase activity. Normalized relative light unit values obtained from cells treated with vehicle were set as onefold induction upon which the activity induced by treatments was calculated.
The plasmid DN/c-fos, which encodes a c-fos mutant that heterodimerizes with c-fos dimerization partners but does not allow DNA biding,55 was a kind gift from Dr C Vinson (NIH, Bethesda, MD, USA). Cells were plated onto 10-cm dishes and prior to treatments cells were transfected for 24 h using X-tremeGENE 9 DNA Transfection Reagent (Sigma-Aldrich) with a control vector and the plasmid DN/c-fos.
Gene expression studies
Total RNA was extracted and cDNA was synthesized by reverse transcription as previously described.56 The expression of selected genes was quantified by real-time PCR using Step One (TM) sequence detection system (Applied Biosystems Inc, Milan, Italy). Gene-specific primers were designed using Primer Express version 2.0 software (Applied Biosystems). For c-fos, p53, p21Cip1/WAF1 (p21) and the ribosomal protein 18S, which was used as a control gene to obtain normalized values, the primers were: 5′- CGAGCCCTTTGATGACTTCCT-3′ (c-fos forward), 5′- GGAGCGGGCTGTCTCAGA-3′ (c-fos reverse); 5′- GCTTCATGCCAGCTACTTC-3′ (p53 forward), 5′- GGCATTCTGGGAGCTTCATCT-3′ (p53 reverse); 5′- GCTTCATGCCAGCTACTTCC-3′ (p21 forward) and 5′- CTGTGCTCACTTCAGGGTCA-3′ (p21 reverse); 5′- GGCGTCCCCCAACTTCTTA-3′ (18S forward) and 5′- GGGCATCACAGACCTGTTATT-3′ (18S reverse), respectively. Assays were performed in triplicate and the results were normalized for 18S expression and then calculated as fold induction of RNA expression.
Chromatin immunoprecipitation assay
Cells were grown in 10-cm dishes to 70–80% confluence, shifted to serum-free medium for 24 h and then treated with vehicle (−) or 100 μM LA for 4 h. Thereafter, cells were cross-linked with 1% formaldehyde and sonicated. Supernatants were immunocleared with salmon DNA/protein A-agarose (Upstate Biotechnology, Inc., Lake Placid, NY, USA) and immunoprecipitated with anti c-fos (H-125) antibody or non-specific IgG (Santa Cruz Biotechnology, DBA, Milan, Italy). Pellets were washed, eluted with a buffer consisting of 1% SDS and 0.1 mol/l NaHCO3, and digested with proteinase K. DNA was obtained by phenol/chloroform extraction and precipitated with ethanol. A 4 μl volume of each sample was used as template to amplify an AP1-containing region located in the p21Cip1/WAF1 promoter by real-time PCR. The primers used were 5′- TCAGCTGCATTGGGTAAATCCT-3′ (forward) and 5′- CTGGACACATTTCCCCACGAA-3′ (reverse). Data were normalized to the input for the immunoprecipitation.
Western blot analysis
Cells were grown in 10-cm dishes, exposed to ligands, and then lysed as previously described.57 Equal amounts of whole-protein extract were resolved on a 10% SDS-polyacrylamide gel and transferred to a nitrocellulose membrane (Amersham Biosciences, Sigma-Adrich, Milan, Italy), which were probed with primary antibodies against pEGFR Tyr 1173, EGFR (1005), phosphorylated ERK1/2 (E-4), ERK2 (C-14), p-c-Jun S73, c-Jun (N), c-fos (E8), p53 (DO-1), p21 (H164) and β-actin (C2) (Santa Cruz Biotechnology) and then revealed using the chemiluminescent substrate for western blotting Westar Nova 2.0 (Cyanagen, Biogenerica, Catania, Italy).
ROS production
The non-fluorescent DCFDA probe, which becomes highly fluorescent on reaction with ROS, was used to evaluate intracellular ROS production. In brief, cells (2×105) were incubated with 10 μM DCFDA (Sigma-Aldrich) at 37 °C for 30 min, washed with PBS and then exposed to treatments, as indicated. Cells were washed with PBS and the fluorescent intensity of DCF was measured (excitation at 485 nm and emission at 530 nm).
Phalloidin staining
Cells were washed twice with PBS, fixed in 4% paraformaldehyde in PBS for 10 min, washed briefly with PBS, then incubated with Phalloidin-Fluorescent Conjugate (Santa Cruz Biotechnology) and visualized with the Olympus BX41 microscope and the images were taken with CSV1.14 software using a CAM XC-30 for images acquisition (Olympus Europa, Hamburg, Germany).
TUNEL assay
Cell apoptosis was determined by TUNEL assay, conducted using DeadEnd Fluorometric TUNEL System (Promega) and performed according to the manufacturer’s instructions. In brief, cells were treated for 18 h, as indicated, then were fixed in freshly prepared 4% paraformaldehyde solution in PBS (pH 7.4) for 25 min at 4 °C. After fixation, cells were permeabilized in 0.2% Triton X-100 solution in PBS for 5 min. After washing twice with washing buffer for 5 min, the cells were covered with equilibration buffer at room temperature for 5–10 min. The labeling reaction was performed using terminal deoxynucleotidyl transferase end-labeling TdT and fluorescein-dUTP cocktail for each sample and incubated for 1 h at 37 °C where TdT catalyses the binding of fluorescein-dUTP to free 3’OH ends in the nicked DNA. After rinsing, cells were washed with 2× SSC solution buffer and subsequently incubated with propidium iodide (Sigma-Aldrich) to stain nuclei and analyzed using the Cytation 3 Cell Imaging Multimode Reader (BioTek, Winooski, VT, USA).
Statistical analysis
Statistical analysis was done using ANOVA followed by Newman–Keuls’ testing to determine differences in means. P<0.05 was considered as statistically significant.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Layden BT, Angueira AR, Brodsky M, Durai V, Lowe WL Jr . Short chain fatty acids and their receptors: new metabolic targets. Transl Res 2013; 161: 131–140.
Calder PC . Functional roles of fatty acids and their effects on human health. JPEN J Parenter Enteral Nutr 2015; 39: 18S–32S.
Bocca C, Bozzo F, Gabriel L, Miglietta A . Conjugated linoleic acid inhibits Caco-2 cell growth via ERK-MAPK signaling pathway. J Nutr Biochem 2007; 18: 332–340.
Engelbrecht AM, Toit-Kohn JL, Ellis B, Thomas M, Nell T, Smith R . Differential induction of apoptosis and inhibition of the PI3-kinase pathway by saturated, monounsaturated and polyunsaturated fatty acids in a colon cancer cell model. Apoptosis 2008; 13: 1368–1377.
Jump DB, Clarke SD . Regulation of gene expression by dietary fat. Annu Rev Nutr 1999; 19: 63–90.
Risérus U . Fatty acids and insulin sensitivity. Curr Opin Clin Nutr Metab Care 2008; 11: 100–105.
Turcotte C, Chouinard F, Lefebvre JS, Flamand N . Regulation of inflammation by cannabinoids, the endocannabinoids 2-arachidonoyl-glycerol and arachidonoyl-ethanolamide, and their metabolites. J Leukoc Biol 2015; 97: 1049–1070.
Dayrit FM . The properties of lauric acid and their significance in coconut oil. J Am Oil Chem Soc 2015; 92: 1–15.
Kappally S, Shirwaikar A, Shirwaikar A . Coconut oil – a review of potential applications. Hygeia J D Med 2015; 7: 34–41.
Eyres L, Eyres MF, Chisholm A, Brown RC . Coconut oil consumption and cardiovascular risk factors in humans. Nutr Rev 2016; 74: 267–280.
Silva RB, Silva-Júnior EV, Rodrigues LC, Andrade LH, da Silva SI, Harand W et al. A comparative study of nutritional composition and potential use of some underutilized tropical fruits of Arecaceae. An Acad Bras Cienc 2015; 87: 1701–1709.
Silberstein T, Burg A, Blumenfeld J, Sheizaf B, Tzur T, Saphier O . Saturated fatty acid composition of human milk in Israel: a comparison between Jewish and Bedouin women. Isr Med Assoc J 2013; 15: 156–159.
Huang WC, Tsai TH, Chuang LT, Li YY, Zouboulis CC, Tsai PJ . Anti-bacterial and anti-inflammatory properties of capric acid against Propionibacterium acnes: a comparative study with lauric acid. J Dermatol Sci 2014; 73: 232–240.
Temme EH, Mensink RP, Hornstra G . Comparison of the effects of diets enriched in lauric, palmitic, or oleic acids on serum lipids and lipoproteins in healthy women and men. Am J Clin Nutr 1996; 63: 897–903.
Alves NF, de Queiroz TM, de Almeida Travassos R, Magnani M, de Andrade Braga V . Acute treatment with lauric acid reduces blood pressure and oxidative stress in spontaneously hypertensive rats. Basic Clin Pharmacol Toxicol 2017; 120: 348–353.
Veeresh Babu SV, Veeresh B, Patil AA, Warke YB . Lauric acid and myristic acid prevent testosterone induced prostatic hyperplasia in rats. Eur J Pharmacol 2010; 626: 262–265.
Fauser JK, Matthews GM, Cummins AG, Howarth GS . Induction of apoptosis by the medium-chain length fatty acid lauric acid in colon cancer cells due to induction of oxidative stress. Chemotherapy 2013; 59: 214–224.
Weng WH, Leung WH, Pang YJ, Hsu HH . Lauric acid can improve the sensitization of Cetuximab in KRAS/BRAF mutated colorectal cancer cells by retrievable microRNA-378 expression. Oncol Rep 2016; 35: 107–116.
Law KS, Azman N, Omar EA, Musa MY, Yusoff NM, Sulaiman SA et al. The effects of virgin coconut oil (VCO) as supplementation on quality of life (QOL) among breast cancer patients. Lipids Health Dis 2014; 13: 139.
Narayanan A, Baskaran SA, Amalaradjou MA, Venkitanarayanan K . Anticarcinogenic properties of medium chain fatty acids on human colorectal, skin and breast cancer cells in vitro. Int J Mol Sci 2015; 16: 5014–5027.
Abdullah FO, Hussain FH, Mannucci B, Lappano R, Tosi S, Maggiolini M et al. Composition, antifungal and antiproliferative activities of the hydrodistilled oils from leaves and flower heads of pterocephalus nestorianus Nab. Chem Biodivers 2017; 14: e1700009.
Jaffe AB, Hall A . Rho GTPases: biochemistry and biology. Annu Rev Cell Dev Biol 2005; 21: 247–269.
Ibarguren M, López DJ, Escribá PV . The effect of natural and synthetic fatty acids on membrane structure, microdomain organization, cellular functions and human health. Biochim Biophys Acta 2014; 1838: 1518–1528.
Wanders RJ, Ruiter JP, IJLst L, Waterham HR, Houten SM . The enzymology of mitochondrial fatty acid beta-oxidation and its application to follow-up analysis of positive neonatal screening results. J Inherit Metab Dis 2010; 33: 479–494.
Estadella D, da Penha Oller do Nascimento CM, Oyama LM, Ribeiro EB, Dâmaso AR, de Piano A . Lipotoxicity: effects of dietary saturated and transfatty acids. Mediators Inflamm 2013; 2013: 137579.
Babu AS, Veluswamy SK, Arena R, Guazzi M, Lavie CJ . Virgin coconut oil and its potential cardioprotective effects. Postgrad Med 2014; 126: 76–83.
Mumme K, Stonehouse W . Effects of medium-chain triglycerides on weight loss and body composition: a meta-analysis of randomized controlled trials. J Acad Nutr Diet 2015; 115: 249–263.
McCarty MF, DiNicolantonio JJ . Lauric acid-rich medium-chain triglycerides can substitute for other oils in cooking applications and may have limited pathogenicity. Open Heart 2016; 3: e000467.
Stanhope JM, Sampson VM, Prior IA . The Tokelau Island Migrant Study: serum lipid concentration in two environments. J Chronic Dis 1981; 34: 45–55.
Reddy BS, Maeura Y . Tumor promotion of dietary fat in azoxymethane-induced colon carcinogenesis in female F 344 rats. J Natl Cancer Inst 1984; 72: 745–750.
Cohen LA, Thompson DO, Maeura Y, Choi K, Blank ME, Rose DP . Dietary fat and mammary cancer. Promoting effects of different dietary fats on N-nitrosomethylurea- induced rat mammary tumorigenesis. J Natl Cancer Inst 1986; 77: 33–42.
Fukui M, Kang KS, Okada K, Zhu BT . EPA, an omega-3 fatty acid, induces apoptosis in human pancreatic cancer cells: role of ROS accumulation, caspase-8 activation, and autophagy induction. J Cell Biochem 2013; 114: 192–203.
Shin S, Jing K, Jeong S, Kim N, Song KS, Heo JY et al. The omega-3 polyunsaturated fatty acid DHA induces simultaneous apoptosis and autophagy via mitochondrial ROS-mediated Akt-mTOR signaling in prostate cancer cells expressing mutant p53. Biomed Res Int 2013; 2013: 568671.
Song Y, Li X, Li Y, Li N, Shi X, Ding H et al. Non-esterified fatty acids activate the ROS-p38-p53/Nrf2 signaling pathway to induce bovine hepatocyte apoptosis in vitro. Apoptosis 2014; 19: 984–997.
Lehto VP . EGF receptor: which way to go? FEBS Lett 2001; 491: 1–3.
Wada T, Penninger JM . Mitogen-activated protein kinases in apoptosis regulation. Oncogene 2004; 23: 2838–2849.
Pellegrin S, Mellor H . Actin stress fibres. J Cell Sci 2007; 120: 3491–3499.
Sebbagh M, Renvoizé C, Hamelin J, Riché N, Bertoglio J, Bréard J . Caspase-3-mediated cleavage of ROCK I induces MLC phosphorylation and apoptotic membrane blebbing. Nat Cell Biol 2001; 3: 346–352.
Huang CY, Liang CM, Chu CL, Peng JM, Liang SM . A fibrillar form of fibronectin induces apoptosis by activating SHP-2 and stress fiber formation. Apoptosis 2010; 15: 915–926.
Xiao L, Eto M, Kazanietz MG . ROCK mediates phorbol ester-induced apoptosis in prostate cancer cells via p21Cip1 up-regulation and JNK. J Biol Chem 2009; 284: 29365–29375.
Coleman ML, Sahai EA, Yeo M, Bosch M, Dewar A, Olson MF . Membrane blebbing during apoptosis results from caspase-mediated activation of ROCK I. Nat Cell Biol 2001; 3: 339–345.
Etienne-Manneville S, Hall A . Rho GTPases in cell biology. Nature 2002; 420: 629–635.
Del Re DP, Miyamoto S, Brown JH . RhoA/Rho kinase up-regulate Bax to activate a mitochondrial death pathway and induce cardiomyocyte apoptosis. J Biol Chem 2007; 282: 8069–8078.
Lai JM, Hsieh CL, Chang ZF . Caspase activation during phorbol ester-induced apoptosis requires ROCK-dependent myosin-mediated contraction. J Cell Sci 2003; 116: 3491–3501.
Abbas T, Dutta A . p21 in cancer: intricate networks and multiple activities. Nat Rev Cancer 2009; 9: 400–414.
Gartel AL . p21(WAF1/CIP1) and cancer: a shifting paradigm? Biofactors 2009; 35: 161–164.
Gartel AL, Tyner AL . The role of the cyclin-dependent kinase inhibitor p21 in apoptosis. Mol Cancer Ther 2002; 1: 639–649.
Karimian A, Ahmadi Y, Yousefi B . Multiple functions of p21 in cell cycle, apoptosis and transcriptional regulation after DNA damage. DNA Repair (Amst) 2016; 42: 63–71.
Dotto GP . p21(WAF1/Cip1): more than a break to the cell cycle? Biochim Biophys Acta 2000; 1471: M43–M56.
Ocker M, Schneider-Stock R . Histone deacetylase inhibitors: signalling towards p21cip1/waf1. Int J Biochem Cell Biol 2007; 39: 1367–1374.
Gogada R, Amadori M, Zhang H, Jones A, Verone A, Pitarresi J et al. Curcumin induces Apaf-1-dependent, p21-mediated caspase activation and apoptosis. Cell Cycle 2011; 10: 4128–4137.
Zhao B, He T . Chidamide, a histone deacetylase inhibitor, functions as a tumor inhibitor by modulating the ratio of Bax/Bcl-2 and P21 in pancreatic cancer. Oncol Rep 2015; 33: 304–310.
Kondo S, Barna BP, Kondo Y, Tanaka Y, Casey G, Liu J et al. WAF1/CIP1 increases the susceptibility of p53 non-functional malignant glioma cells to cisplatin-induced apoptosis. Oncogene 1996; 13: 1279–1285.
Lincet H, Poulain L, Remy JS, Deslandes E, Duigou F, Gauduchon P et al. The p21(cip1/waf1) cyclin-dependent kinase inhibitor enhances the cytotoxic effect of cisplatin in human ovarian carcinoma cells. Cancer Lett 2000; 161: 17–26.
Gerdes MJ, Myakishev M, Frost NA, Rishi V, Moitra J, Acharya A et al. Activator protein-1 activity regulates epithelial tumor cell identity. Cancer Res 2006; 66: 7578–7588.
Lappano R, Santolla MF, Pupo M, Sinicropi MS, Caruso A, Rosano C et al. MIBE acts as antagonist ligand of both estrogen receptor α and GPER in breast cancer cells. Breast Cancer Res 2012; 14: R12.
Santolla MF, Avino S, Pellegrino M, De Francesco EM, De Marco P, Lappano R et al. SIRT1 is involved in oncogenic signaling mediated by GPER in breast cancer. Cell Death Dis 2015; 6: e1834.
Acknowledgements
This work was supported by Associazione Italiana per la Ricerca sul Cancro (IG 16719 to MM; IG 19242 to AB).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by A Rufini
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lappano, R., Sebastiani, A., Cirillo, F. et al. The lauric acid-activated signaling prompts apoptosis in cancer cells. Cell Death Discov. 3, 17063 (2017). https://doi.org/10.1038/cddiscovery.2017.63
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/cddiscovery.2017.63
This article is cited by
-
Integrated transcriptomics and metabolomics analysis reveals the biomolecular mechanisms associated to the antitumoral potential of a novel silver-based core@shell nanosystem
Microchimica Acta (2023)
-
Metabolic disorders sensitise endometrial carcinoma through endoplasmic reticulum stress
Cellular & Molecular Biology Letters (2022)
-
Two species of Ulva inhibits the progression of cervical cancer cells SiHa by means of autophagic cell death induction
3 Biotech (2021)
-
Curcusone C induces apoptosis in endometrial cancer cells via mitochondria-dependent apoptotic and ERK pathway
Biotechnology Letters (2021)
-
Nutraceutical profiling of surimi gel containing β-glucan stabilized virgin coconut oil with and without antioxidants after simulated gastro-intestinal digestion
Journal of Food Science and Technology (2020)