Abstract
Tumors often respond favorably to initial chemotherapy but eventually relapse with drug resistance and increased metastatic potential. Cellular senescence is a major therapeutic outcome of cancer chemotherapy, which leads to tumor stasis or regression through immune clearance of senescent cells. However, senescent tumor cells have been shown to resume proliferation at low frequency. We found that subjecting arrested senescent tumor cells to cytotoxic treatments stimulates the clonogenic proliferation of remaining survivors. The senescence revertants showed a reduced rate of proliferation but increased migration and invasion potential in vitro, and increased tumorigenic potential in vivo. Gene expression profiling showed that the senescence revertants are distinct from both parental and senescent cells. A subset of senescence-activated genes remains active in the revertants. These genes are implicated in regulating cell motility, invasion, and metastasis, which may collectively contribute to the aggressiveness of the revertants. The findings suggest that although therapy-induced senescence has short-term benefits, the response also causes reprogramming of gene expression and activates invasion-related genes that accelerate tumor progression.
Similar content being viewed by others
Introduction
Post-treatment relapse and distant metastasis are major causes of mortality by solid tumors. There is extensive preclinical evidence in animal models that nearly all types of cancer treatments, from surgery to chemotherapy and targeted therapy, can stimulate tumor spread.1 Modification of the host microenvironment by the treatments plays an important role by providing survival niches for metastatic tumor cells. Furthermore, tumor cells are genetically unstable. Induction and subsequent Darwinian evolution of drug-resistant cells play important roles in therapy-induced progression. It is likely that additional mechanisms also contribute to this phenomenon.
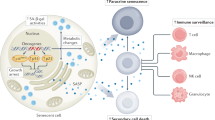
Replicative senescence was initially discovered as a cell culture phenomenon. Excessive cell division results in telomere erosion, activation of DNA damage signaling, and irreversible arrest mediated by the pRb and p53 tumor suppressors.2 Senescent cells in culture exhibit large flat cell morphology, increased p21/p16 levels, active metabolism, and increased lysosomal SA-β-galactosidase (SA-β-gal) activity. Radiation and chemotherapy drugs also induce premature senescence that is similar to replicative senescence.3 SA-β-gal staining suggests that senescence occurs in vivo in aging tissues, premalignant lesions, and in tumors treated with chemotherapy.4 Mouse models showed that tumor cells with defective apoptosis pathway respond to chemotherapy by entering senescence. Senescent tumor cells secrete many pro-inflammatory cytokines, growth factors, and matrix metalloproteases (the SASP phenotype). These factors act through paracrine and autocrine mechanisms to promote immune clearance and tumor remission. Therefore, senescence is an important tumor suppression mechanism.5
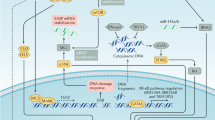
Senescence cells undergo self-sustaining cell-cycle arrest involving stable epigenetic silencing of proliferation genes.4 Silenced E2F target genes form heterochromatin foci (SAHF) in some senescent cells.6 Senescent cells also upregulate many pro-inflammatory genes.2 Presumably, senescence involves establishing and maintaining positive feedback loops in the heterochromatinization of cell-cycle genes and activation of senescence-specific genes. Heterochromatin proteins such as HP1 and SUV39H1 bind to dimethylated H3K9 and then promote further methylation of adjacent H3K9. Therefore, they help maintain self-perpetuating positive feedback loops and stable repression. In addition to transcription repression, senescent cells display constitutively active DNA damage signaling.7 Paracrine and autocrine effects from the SASP factors also play a role in maintaining positive feedback activation of gene expression and senescence arrest.8,9
Tumor cells that are resistant to apoptosis often respond to chemotherapy by entering premature senescence. Normal stromal fibroblasts also enter senescence after DNA-damaging treatment. Although senescence is generally perceived as a form of irreversible cell-cycle arrest, studies of drug-induced senescence showed that senescent tumor cells in culture spontaneously revert to active proliferation at low frequency.10 Inactivation of p53 or pRb in early stage senescent cells is often sufficient to stimulate cell-cycle re-entry.11 Our recent study showed that deficiency in nucleolar rRNA transcription repression significantly increases the frequency of senescence reversal.12 Therefore, after termination of drug treatment, senescent tumor cells may eventually produce proliferative clones and result in relapse.
In addition to causing relapse, senescence reversal of tumor cells may have other adverse effects. Recent studies suggest that tumor cells in culture that have undergone senescence arrest re-emerge with increased levels of certain tumor stem cell markers.13,14 Normal human fibroblasts undergoing replicative senescence acquire DNA hypomethylation/hypermethylation patterns similar to cancer cells.15 Furthermore, the cancer-like DNA methylation pattern is partially retained after the senescent fibroblasts are forced to proliferate by SV40 T antigen expression.15 Senescent fibroblasts forced to re-enter the cell cycle by p53 inactivation retain the expression of many genes associated with senescence.16 Therefore, senescence in fibroblasts creates long-lasting imprints on the epigenome and certain gene expression programs. Similar reprogramming may also occur in tumor cells that have undergone senescence reversal.
Chemotherapy promotes the emergence of drug-resistant and simultaneously more malignant tumor cells.17,18 Induction chemotherapy has been shown to significantly accelerate the re-growth of NSCLC compared with untreated tumors.19 Multiple mechanisms, such as selection of pre-existing mutant clones and activation of stress-resistant genes by epigenetic mechanisms, are responsible for some of the effects. Tumor stem cells that exist in a stress-resistant epigenetic state in the population may be enriched by the chemotherapy and contribute to relapse and metastasis.20,21 Stromal fibroblast senescence and production of SASP factors can promote tumor cell proliferation and invasion through paracrine mechanism, creating a microenvironment for metastasis.9 Whether tumor cell senescence response also promotes progression is unclear.
Results described in this report show that tumor cell senescence is frequently reversed after stimulation by a variety of stress signals. Reversal from senescence produces tumor cells that are distinct from the parental cells, exhibiting altered gene expression profile and increased invasiveness. The results suggest that senescence response to DNA damage by tumor cells may contribute to the phenomenon of therapy-induced progression.
Results
Stress treatment of senescent tumor cells promotes cell-cycle re-entry
Reversal from drug-induced senescence has been implicated as a mechanism of tumor recurrence.10 Therefore, we were interested in identifying secondary treatments that can reduce the frequency of senescence reversal. As a cell culture model of therapy-induced senescence, A549 lung tumor cells were treated 7 days with 100 nM doxorubicin, which resulted in cell-cycle arrest and SA-β-gal positive staining (Figure 1a). Following drug treatment, the cells were washed and cultured in drug-free medium for 20–30 days and stained for colony formation. The treatment resulted in >90% SA-β-gal-positive cells and >99% of the cells remained arrested 21 days after drug removal (Figure 1b).
Stress treatments stimulate senescence reversal. (a) A549 cells were treated with 100 nM doxorubicin for 7 days and stained for SA-β-galactosidase activity. Cell morphology was photographed under phase-contrast microscope. Bar=20 μm. (b–f) Senescent A549 cells induced by doxorubicin were mock treated (b) or treated with different stimuli, including 80 μM etopside (c), 2 μM ABT-737 (d), 500 nM camptothecin (e), and glucose deprivation (f) for indicated time. The cells were then washed and cultured in drug-free media for 3 weeks. Colonies were stained with crystal violet. Bar=20 μm.
To test the impact of different stress signals on senescence maintenance, the senescent cells were incubated with drugs or glucose-free medium. After removing the drugs by washing, the cells were incubated in drug-free medium for 21 days to determine colony formation efficiency. The results with several drugs showed drastically different outcomes. Secondary treatment with etoposide did not induce noticeable cell death or morphological change, but significantly reduced background colony formation efficiency (Figure 1c). In contrast, the Bcl2 inhibitor ABT-737 induced significant death in senescent cells, but surviving cells formed colonies at much higher frequency than background (Figure 1d). Camptothecin (CPT) and glucose deprivation induced significant cell rounding and detachment. The small number of surviving cells also formed colonies at high efficiency in drug-free medium (Figures 1e and f).
In similar experiments, the RNA Pol I inhibitor CX5461 was used to induce senescence in A549 cells.12,22 Secondary treatments with ABT-737, CPT, and glucose starvation also induced partial cell death accompanied by significant colony formation (Supplementary Figure S1). These results suggest that stress treatments can stimulate senescent cells to re-enter cell cycle.
Inhibition of Bcl2 promotes senescence escape
The ability of Bcl2 inhibitor ABT-737 to induce clonogenic growth of senescent cells was of interest because a derivative of this compound is used in clinical trials. Non-senescent A549 cells were not sensitive to apoptosis by ABT-737 (Figure 2a). However, senescent cells showed significant sensitivity to ABT-737-induced apoptosis and PARP cleavage (Figures 2a and b), consistent with recent report using senescent fibroblasts.23 Transient knockdown of Bcl2 using siRNA also induced colony formation in senescent A549 cells (Supplementary Figures S2a–d). Senescent H460 cells were also induced to proliferate by ABT-737 treatment (Supplementary Figure S2e), suggesting that the phenomenon is not restricted to A549 cells alone. To directly test whether activation of apoptotic caspase stimulates senescence reversal, senescent A549 cells were transiently transfected with Caspase 3, followed by colony formation analysis. Transfection of Caspase 3 induced partial cell death (Figure 2c). Following long-term culture, significant colony formation was observed (Figure 2d). This result suggests that direct activation of caspase 3 increases the probability of senescence escape.
Apoptosis stimulates senescence reversal. (a) Senescent A549 cells were treated with 2 μM Bcl2 inhibitor ABT-737 for 2 days and photographed to show apoptotic cell death. Bar=20 μm. (b) Senescent A549 cells induced by doxorubicin were treated with 2 μM ABT-737 in the presence of pan-specific caspase inhibitor QVD-OPh (20 μM) for indicated times. The cells were analyzed for the indicated markers by western blot. (c) Senescent A549 cells were transfected with pCMV-caspase-3. Cell morphology was photographed 5 days after the transfection. Bar=20 μm. (d) Senescent A549 cells were transfected with the indicated plasmids and cultured for 7 weeks. Colony formation was detected by crystal violet staining. (e) Senescent A549 cells were treated with 2 μM ABT-737 in the presence or absence of 20 μM QVD-OPh for 3 days, the cells were washed and cultured in drug-free medium until colonies formed. Colonies were stained with crystal violet. (f) Senescent A549 cells were treated with DMSO control or 2 μM ABT-737 for 3 days. The expressions of cell-cycle proteins in senescent cells after removing the stimuli for 24 h were analyzed by western blot.
To test whether ABT-737-mediated senescence reversal requires caspase activation, cells were treated with combination of ABT-737 and broad spectrum caspase inhibitor QVD-OPh. QVD-OPh completely blocked PARP cleavage and apoptosis induced by ABT-737 (Figure 2b) but only moderately reduced colony formation efficiency (Figure 2e), suggesting that both caspase-dependent and -independent mechanisms were involved. The cell death caused by CPT was not blocked by the caspase inhibitor QVD-OPh (Supplementary Figure S3a), suggesting that it was not apoptosis. Staining of the cells treated with CPT or glucose starvation with fluorescent dyes Hoechst 33342 and propidium iodide revealed that cells mainly undergo necrosis (Supplementary Figure S3b). The results above suggest that senescence arrest in tumor cells is susceptible to disruption by various stress signals. Necrosis is a clinical relevant form of stress in vivo since large tumors often contain necrotic regions due to poor blood supply.
Analysis of cell-cycle markers revealed that ABT-737 reduced p27 levels, while the level of p53 and p21 were not significantly affected (Figure 2f). The expression of p27 in proliferating A549 cells was not affected by ABT-737 (data not shown). Bcl2 has been shown to promote p27 expression, cell-cycle arrest, and senescence.24,25 ABT-737 may stimulate the proliferation of senescent cells partly by inhibiting Bcl2 and downregulating p27.
Senescence revertants exhibit increased migration and invasion
Senescent cells undergo extensive epigenetic and gene expression changes, which may not be completely erased when cells resume proliferation.15 To test whether the senescence revertants gained aggressive features, the revertant colonies were pooled and analyzed in wound healing assay and Matrigel invasion assay. The results revealed that starvation-induced revertants showed stronger migration (Figure 3a) and increased ability to traverse Matrigel barrier (Figures 3b and d). Spontaneous revertants also showed similar results (not shown), suggesting that senescence revertants are inherently more invasive irrespective of whether they arise spontaneously or induced by stress treatments.
Senescence revertants have increased migration and invasion potential. (a) Cell migration activity of A549 parental cells and senescence revertants was compared using wound healing assay. Upper panels show the identical gaps created at the beginning of the assay. Cells were cultured for 24 h and stained with crystal violet to determine the closure percentage. Bar=20 μm. (b and c) Glucose starvation-induced escape colonies from senescent A549 were cultured in Matrigel chambers; cell transition through the matrix was stained after 48 h. Bar=20 μm. (d) Quantitation of (b). Values are mean±S.D. of triplicates. (e and f) A549 doxorubicin-induced senescence revertants were treated with doxorubicin or camptothecin for 3 days; cell death was determined by MTT assay.
Since initial experiments were performed using A549 cells with unknown passage number, the senescence/reversal procedure may enrich a pre-existing subpopulation. Therefore, we derived a clonal A549 cell line from a single cell and repeated the experiments. Similar increase of invasiveness was observed from the senescence revertants (Figure 3c). Furthermore, the revertants showed no increase in resistance to different chemotherapy drugs in short-term survival test (Figures 3e and f), arguing against the selection of cancer stem cells or drug-resistant cells in our assay. Instead, the results suggested that the process of entering and exiting senescence produced reprogrammed aggressive cells.
Persistent activation of a subset of senescence genes in revertants
RT-PCR analysis using a focused cancer stem cell gene panel did not reveal increased expression of stem cell markers in the senescence revertants (Supplementary Figure S4). To further investigate the basis of increased invasiveness in the senescence revertants, RNA-seq was performed on proliferating A549, arrested senescent cells, and proliferating revertants (all derived from the same clonal A549 cell line). SA-β-gal staining confirmed there were no senescent cells in the proliferating revertant culture. Senescent A549 cells showed strong activation of large numbers of genes (449 genes activated >4-fold; Supplementary Table S1). PANTHER gene ontology analysis showed that 41% of the activated genes encode proteins that were membrane-associated, extracellular matrix, or secreted proteins (Figure 4a).26 DAVID analysis also suggested strong functional clustering in extracellular and immunity functions.27 There were 236 genes downregulated by >4-fold in senescent cells (Supplementary Table S2); 25% of these genes were involved in DNA replication (Figure 4b). DAVID analysis of the downregulated genes also suggested strong functional clustering in cell cycle and cell division. The results were consistent with the activation of pro-inflammatory pathways and stable cell-cycle arrest in senescent cells.
Senescence revertants retain activation of a subset of senescence genes. (a) Cellular component of genes activated in senescent cells based on PANTHER gene ontology analysis. (b) Pathways of genes downregulated in senescent cells based on PANTHER gene ontology analysis. (c) Overlap of genes activated in senescent cells and senescence revertants. Sixty-one of 77 genes activated in senescence revertants were also activated in senescence cells.
The revertants had 77 genes activated >2 fold compared with A549 (P<0.1), these genes were highly reproducible by RT-PCR analysis (35/39, see below). Interestingly, 61 (79%) of these genes were also activated >2-fold in senescent cells (Figure 4c and Supplementary Table S3). There were 63 genes repressed >2-fold in the revertants relative to A549 (P<0.1; Supplementary Table S4). Unlike the activated genes, the repressed genes in the revertants did not overlap with repressed genes in senescent cells, showed no clear functional clustering in DAVID analysis, and had poor confirmation rate by RT-PCR (0/7). Therefore they were not investigated further. The high degree of overlap in activated genes (Figure 4c) suggests that the revertants were originated from senescent cells and retained activation of a subset of senescence genes. Furthermore, there was no strong activation of other genes in the revertants that would suggest the selection of a unique population such as stem cells or drug-resistant cells.
Validation of revertant genes
To validate the RNA-seq results, 39 genes that were activated in both senescent cells and revertants were selected for analysis by RT-PCR. Activation was confirmed in both senescent cells (39/39, 100%) and in revertants (35/39, 90%; Table 1). In contrast, seven of seven repressed genes failed confirmation by RT-PCR (not shown).
To test the cell type specificity of senescence-mediated long-term gene activation, 19 A549 genes were selected for analysis in senescence revertants derived from H1299 and MCF7 cells. The results showed strong similarity between NSCLC cell lines A549 and H1299. Although some A549 genes (9/19) were not expressed in H1299, between the expressed genes senescent A549 and H1299 cells shared 100% similarity (10/10), and the revertants shared 80% similarity (8/10) (Table 2). In contrast, the similarity between A549 and MCF7 (breast tumor) was significantly weaker. Although most A549 genes were also expressed in MCF7 (18/19) and senescent A549 and MCF7 shared 72% (13/18) activated genes, the magnitude of activation in MCF7 was significantly lower. Furthermore, there was only 7% (3/18) similarity between A549 and MCF7 revertants (Table 2). The results suggest that cells of the same tumor type shared significant similarity in gene activation during senescence and reversal. There is significant cell type specificity in the genes that remain activated after senescence reversal.
In gene ontology analysis, the A549 revertant-activated genes also showed significant bias towards membrane-associated or extracellular proteins as expected from their significant overlap (79%) with senescence-activated A549 genes. Further literature search of these genes revealed that most were linked to cell adhesion, migration, and metastasis, suggesting that they may be responsible for the enhanced invasiveness in senescence revertants (Table 2). In general, the magnitude of activation of the revertant genes was significantly smaller compared with their expression level in senescent cells (e.g., NPTX1 was activated 40-fold in senescent cells versus 6-fold in revertants; Table 1).
Senescence reversal as a potential source of non-genetic heterogeneity
The RNA-seq analysis and RT-PCR validation was performed using pooled revertant colonies. To determine whether there was clonal variability of gene activation in the revertants, a clonal A549 was used to generate senescence revertants and clonal cell lines were established from individual revertant colonies. As control, the parental clonal A549 was also used to establish single-cell subclones. RT-PCR analysis of several genes from Table 1 suggested clonal variation in the expression of AMOT and IGF2 in the revertants (Figures 5a and b), whereas the same revertant clones showed relatively uniform increase of CPE and SP5 expression (Figures 5c and d). Therefore, senescence reversal may expand the range of expression level of a subset of genes, producing subclones with different combination of activated genes.
Gene expression heterogeneity in senescence revertant clones. A clonal A549 cell line was used to generate 14 clonal sublines by limited dilution. The same cell line was also treated with doxorubicin to generate senescence revertant colonies. Fourteen revertant colonies were cloned and expanded into cell lines. The expression of AMOT (a), IGF2 (b), CPE (c) and SP5 (d) in 14 A549 clonal sublines and 14 clonal revertant cell lines were analyzed by RT-qPCR. Values are mean±S.D. for experimental triplicates.
Senescence revertants have reduced proliferation rate in vitro
During routine culture, the pooled senescence revertants did not show noticeable difference in proliferation rate compared with parental cells. To detect more subtle changes, the clonal A549 cells and pooled senescence revertants were labeled with lentivirus expressing H2B-RFP and H2B-GFP respectively.28 The labeled cells were mixed at 1 : 1 ratio and cultured for eight passages. FACS analysis of RFP/GFP ratio showed that parental cells in the mixture out grew the senescence revertants during co-culture, irrespective of the fluorescence marker (Figures 6a and b). Therefore, although the revertants were more motile and invasive, they had a slight proliferative disadvantage compared with parental cells under normal culture conditions. When the fluorescent-labeled cells were co-cultured in the presence chemotherapy drugs for several passages, the revertants showed a growth advantage in the presence of cisplatin and CPT, whereas 5-FU and etoposide had no effect or even preferentially blocked the growth of revertants (Figures 6c and d). The results suggest that under certain stress conditions, the senescence revertants have a competitive advantage over the parental cells.
Senescence revertants have reduced proliferation potential in vitro. (a) A549 cells stably infected with lentivirus expressing GFP or RFP were used to generate senescence revertants from doxorubicin treatment. The RFP-labeled parental cells and GFP-labeled revertants were mixed at 1 : 1 ratio, co-cultured for eight passages, and analyzed by FACS. (b) GFP-labeled parental cells and RFP-labeled revertants were mixed at 1 : 1 ratio, co-cultured for 8 passages, and analyzed by FACS. (c and d) RFP and GFP-labeled cells were mixed 1 : 1 and treated with 50 μM 5-FU, 40 μM etoposide, 0.5 μM CPT and 20 μM cisplatin. The cells were treated for 7 days, followed by 14-day culture in drug-free medium. The treatment was repeated once and the surviving cells were analyzed by FACS.
In vivo competition between revertant and parental cells
To determine whether the senescence revertants can compete with parental cells under in vivo growth condition, 1 : 1 mixture of GFP-parental:RFP-revertant and RFP-parental:GFP-revertant cells were inoculated into nude mice subcutaneously (Figure 7a). The tumors formed by the cell mixture were dissociated into single cells and analyzed by FACS to determine the GFP/RFP ratio (Figures 7b and c). The results showed that in 13/16 tumors the revertant cells outgrew the parental cells (P=0.03; Figure 7d). Therefore, although the senescence revertants had a proliferative disadvantage in culture, they were able to out-compete the parental cells under more stressful in vivo tumor growth condition. We also inoculated the cell mixture through tail vein injection in an attempt to compare metastasis with the lung. However, the analysis was uninformative due to the low percentage of fluorescent tumor cells in the lung cell mixture (not shown).
Senescence revertants exhibit growth advantage in vivo. (a) A schematic diagram of the in vivo competition assay. (b and c) GFP/RFP-labeled parental A549 and senescence revertants were inoculated subcutaneously into nude mice. After the tumor nodules were formed, the tumors cells were isolated and analyzed for the GFP/RFP ratio by FACS. The ratios of cell mixtures before injection and from representative tumors are shown. (d) Summary of parental/revertant cell ratios in 16 tumors (8 from RFP-parental/GFP-revertant mixture and 8 from GFP-parental/RFP-revertant mixture).
Discussion
DNA-damaging chemotherapy induces senescence in both stromal and tumor cells. Senescent stromal fibroblasts have been shown to promote tumor growth and metastasis through secretion of SASP factors. The results described in this study suggest that senescence response by tumor cells may also contribute to therapy-induced progression by producing invasive revertants. The basis of this phenomenon is the ability of senescent tumor cells to re-enter the cell cycle when disturbed by various forms of stress, and the stable activation of a subset of genes implicated in cell motility and metastasis in the revertants.
Contrary to the common perception that senescence is an irreversible arrest, previous work showed that inactivation of p53 in human fibroblasts undergoing replicative senescence can lead to cell-cycle re-entry.29 Therefore, after cells have committed to senescence, disruption of cell-cycle regulators can cause reversion to active proliferation. Spontaneous escape from drug-induced senescence arrest has been suggested to be a mechanism of tumor relapse.10 Our current results showed that several cytotoxic treatments that induce apoptosis and necrosis can stimulate senescent tumor cells to re-enter cell cycle. This cell culture phenomenon is likely to be clinically relevant, since senescent cells in tumors are exposed to a variety of stresses due to chemotherapy treatments, hypoxia and nutrient deprivation.
Although the mechanism of senescence reversal by these treatments remains to be investigated in detail, previous studies showed that apoptotic cells activate neighboring cells through secretion of prostaglandin E2.30,31 Caspase 3 activation leads to cleavage of pRb, which is an important mediator of senescence.32 In addition to pRb, Caspase 3 cleaves a large number of proteins involved in chromatin modification.33 It is possible that lethal levels of Caspase 3 stimulates neighboring cells through paracrine mechanisms, whereas sublethal Caspase 3 activation causes changes in heterochromatin in senescent cells, increasing the probability of reactivating genes essential for cell cycle.
Classic drug resistance selection scheme using cytotoxic levels of drugs can enrich for survivors with cancer stem cell features.14,20,21 Previous studies suggested that tumor cells spontaneously reverted from drug-induced senescence were more stem-like.13,14 Our results suggest that the non-lethal levels of DNA damage in our experiments did not select for or induce stem cells, but instead induced stable changes in gene expression through activating the senescence response program. Establishing a senescence state involves downregulation of large number of genes involved in DNA replication and cell division, and activation of numerous genes in inflammation, motility, invasion, and cell–cell communication. The return to active cell cycle requires restoring normal expression of DNA replication and cell division genes. However, a subset (~10%) of senescence-activated genes remains activated. The degree of activation of these ‘revertant genes’ were generally weaker than in senescent cells, suggesting that the upstream signals that regulate their expression in senescent cells were decreased or absent in the revertants.
It is possible that epigenetic changes and positive feedback loops become self-perpetuating for a subset of senescence genes, resulting in continued activation without upstream signals in the revertants. Our comparison of lung and breast tumor cell lines suggests that different cell types may retain expression of distinct subset of senescence genes after reversal. Previous study revealed strong cell type-specific difference in gene expression profile when comparing the senescent state.34 The difference in chromatin state and tissue-specific gene expression may dictate what genes are activated during senescence, and remain active after re-entering the cell cycle. Interestingly, the revertants have a growth disadvantage compared with parental cells under normal cell culture condition, suggesting that there may be a tradeoff in proliferation rate for the continued expression of some senescence genes. However, in a tumor xenograft environment, the revertants were able to out-compete the parental cells, suggesting that proliferation rate alone does not determine tumorigenic potential in vivo.
Tumor cell heterogeneity presents a significant hurdle to curative treatment.35 Genetic and epigenetic heterogeneity facilitates Darwinian evolution, and predicts tumor progression.36,37 Non-genetic transient or semi-permanent gene expression changes also promote treatment resistance and progression.20 Understanding the cause of tumor cell heterogeneity is necessary for developing treatments that prevent its emergence. Our results suggest that senescence response by tumor cells may provide a non-genetic mechanism of increasing tumor heterogeneity, since a subset of the revertant genes were activated with significant degree of clonal variability. It is possible that in the absence of sustained upstream signaling, the revertant genes were turned off in a random fashion, producing clones with different combination of gene activation and heterogeneity.
Chemotherapy is often given in cycles with significant gaps in between to allow for recovery from side effects. The results of our study suggest that apoptotic or cytotoxic agents may have unintended activities that cause activation of senescent tumor cells from prior treatments, generating new variants with stable activation of genes that stimulate invasion and metastasis. Multiple cycles of senescence and reversal may further increase the number of activated genes and increase the level of clonal heterogeneity. This phenomenon underscores the complexities of cancer therapy and may contribute to the lack of overall survival benefit of certain treatments despite favorable short-term responses. The results suggest that blocking the initiation of senescence, or inhibiting reversal from senescence, may reduce the chance of tumor progression.
Materials and methods
Cell lines and reagents
The human NSCLC cell lines A549, H460, H1299 and human breast cancer cell line MCF7 were purchased from the ATCC (Manassas, VA, USA) from 1995 to 2000 and maintained in Dulbecco modified Eagle medium (DMEM) with 10% fetal bovine serum. Low passage (<5) original stocks were used for the experiments. The cells have not been reauthenticated, but were routinely monitored for morphological characteristics and p53 status by western blot (wild-type p53 for A549/H460/MCF7, p53-null for H1299). MCF7 was also monitored for p53 codon 72 polymorphism by western blot. Cells were cultured at a 37 °C incubator in a humidified atmosphere containing 5% CO2. A549 cells with stable expression of GFP and RFP were generated by infecting A549 cells with LV-GFP and LV-RFP viruses. The LV-GFP and LV-RFP plasmids were gifts from Elaine Fuchs (Addgene plasmid # 25999 and # 26001). Corning Biocoat Matrigel Invasion Chambers was obtained from Corning (Bedford, MA, USA). Mouse anti-PARP, mouse anti-p53 (DO1), mouse anti-p27, and mouse anti-p21 were purchased from BD Biosciences (San Jose, CA, USA). Mouse anti-β-actin was obtained from Sigma (St Louis, MO, USA). Rabbit anti-Bcl2 antibody was purchased from Cell Signaling Technology (Danvers, MA, USA). MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) was obtained from Sigma. Human cancer stem cell PCR array (PAHS-176ZA) was purchased from Qiagen (Germantown, MD, USA). Data processing was performed using the manufacturer website.
SA-β-galactosidase staining
A549 cells (3×104/well) were seeded to 24-well plates for overnight before being treated with 100 nM doxorubicin. After 7 days of the treatment, SA-β-galactosidase staining was carried out using the Senescence histochemical staining kit (Sigma) following the manufacturer's instruction. Cells were incubated at 37 °C for 18 h and photographed.
Colony formation assay
A549 cell senescence was induced by the treatment with 100 nM doxorubicin for 7 days. The cells were washed twice with PBS to remove doxorubicin. The senescent cells were later treated with different drugs, including 80 μM etopside, 500 nM camptothecin, 2 μM ABT-737, cultured in glucose-free medium for 5 days, or transfected with 2 μg of Caspase 3 plasmid for 24 h. After removing the stimuli, cells were cultured in medium for 3–7 weeks with re-feeding every 5 days until colonies were visible. Colonies were stained by 0.5% crystal violet.
RNA interference (RNAi)
Senescence cells induced by 0.1 μM doxorubicin for 7 days were transfected with 50 nM control siRNA ( AATTCTCCGAACGTGTCACGT), Bcl2 siRNA1 ( GGATGACTGAGTACCTGAA), and Bcl2 siRNA2 ( GGAGAACAGGGTACGATAA) (Invitrogen, Carlsbad, CA, USA) using RNAiMAX (Invitrogen) according to the instructions from the supplier. After 48 h of transfection, cells were cultured in normal DMEM medium to observe colonies formation. The knock down efficiency of Bcl2 was detected both on mRNA and protein level.
Cell growth inhibition assay
A549 parental and revertant cells (5×103/well) were seeded to 96-well plate. After 24 h incubation, cells were treated with different concentrations of doxorubicin or camptothecin for 72 h. Cell growth inhibition was detected by MTT assay. Briefly, 10 μl MTT (final concentration 5 mg/ml) was added to each well and after 3 h incubation in cell culture incubator, the culture medium was removed and 200 μl DMSO was added to each well to dissolve the formazan. The optical density was measured spectrophotometrically at 570 nm.
Wound healing assay
In vitro wound healing assay was used to measure the migration ability of A549 parental cells or A549 cells escaped from senescence. The cells (4×105/well) were plated in six-cell plates for 18 h to form a confluent monolayer. The cells were then maintained in serum-free medium for 18 h before scraping in a straight line with a P200 pipette tip. After removing the debris, cells were maintained in regular medium for 24–48 h, stained with 0.5% crystal violet, and photographed under a phase-contrast microscope.
Transwell assay
The Matrigel invasion chamber (Corning) was used to assess the invasive property of A549 parental cells or A549 revertants in vitro. The assay was based on the manufacturer protocol. The cells were cultured in serum-free medium for 18 h, trypsinized, and suspended in serum-free medium at 5×104 cells/ml for 24-well chambers. The chamber was then inserted to the wells containing chemoattractant and cultured in the humidified cell culture incubator for 24–48 h. Cell invasion was measured by crystal violet staining and photographed.
Western blot
After the indicated treatments, cells were lysed in lysis buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 0.5% NP-40, 1 mM phenylmethylsulfonyl fluoride, protease inhibitor cocktail) and centrifuged for 10 min at 14 000×g to remove insoluble debris. The supernatant was boiled in sample buffer for 5 min and subjected to SDS-PAGE and western blot to detect the expression of indicated proteins using their corresponding antibodies.
Necrosis assay
A549 cell senescence was induced by 7 days treatment with 100 nM doxorubicin. The cells were then treated with 500 nM camptothecin or cultured in glucose-free medium for 48 h. Hoechst 33342 (1 μg/ml) and propidium iodide (5 μg/ml) were added to the medium and incubated for 15 min. Both floating and attached cells were collected for the necrosis analysis. The cell pellets were washed with PBS and suspended in PBS. The cell suspension (0.1 ml, ~105 cells) was centrifuged onto glass slides in a cytospinner for 3 min at 300×g. The slides were washed with PBS once and fixed in pre-chilled methanol/acetone (3 : 1) for 20 min at −20 °C. The slides were washed with PBS and mounted in Antifade solution. Necrotic cells were examined by fluorescence microscopy at 340/425 nm (Hoechst 33342) and 580/630 nm (propidium iodide).
RNA sequencing
RNA samples were prepared by RNeasy min kit (Qiagen); a mixture of three biological RNA replicates was used to represent each condition (parental, senescent, revertant). RNA-seq analysis, including rRNA depletion, library preparation, multiplexing and cluster generation, sequencing on Illumina HiSeq2500, and differential gene expression analysis, was performed by Genewiz (South Plainfield, NJ, USA). Gene function analysis was performed using DAVID (Database for Annotation, Visualization and Integrated Discovery, https://david.ncifcrf.gov/) and PANTHER (Protein Analysis THrough Evolutionary Relationships, http://pantherdb.org) programs.
RNA-seq genes validation
To validate the activated genes identified by RNA-seq, a panel of primers were designed using the Primer3 (v. 0.4.0) software and synthesized by Integrated DNA Technologies (IDT, Coralville, IA, USA). The sequences of primers are listed in Supplementary Table S5. The RNA samples were prepared by RNeasy min kit (Qiagen). High capacity cDNA reverse transcription kit (Appliedbiosystems by Thermo Fisher Scientific, Waltham, MA, USA) was used to transcribe RNA to cDNA. The PowerUp SYBR green master mix kit was employed for the qPCR analysis.
Flow cytometry
A549 cells were infected with LV-GFP or LV-RFP lentivirus to generate the A549-GFP and A549-RFP cells. Despite the absence of selection markers on the viral vectors, the GFP/RFP labels were stably expressed through multiple generations. The infected cells were treated with 100 nM doxorubicin for 5 days to induce senescence, followed by culture in drug-free medium to generate revertant colonies. The pooled revertant colonies (RFP or GFP labeled) were mixed with the parental cells (GFP or RFP labeled) at 1 : 1 ratio, cultured for eight passages and subjected to FACS analysis using LSRII flow cytometer (BD BioSciences, San Jose, CA, USA). GFP or RFP-parental and RFP or GFP-revertants were mixed 1 : 1 and treated with chemotherapy drugs including 50 μM 5-FU, 40 μM etoposide, 0.5 μM CPT, and 20 μM of cisplatin. The cells were treated for 7 days with the drugs, followed by 2 weeks culture in drug-free medium. The treatment was repeated once and the surviving cells were analyzed by FACS. The data were analyzed using FlowJo software (TreeStar, Ashland, OR, USA).
In vivo tumor growth and metastasis assay
Experimental procedures involving animals were reviewed and approved by the Institutional Animal Care and USE committee of the University of South Florida. Athymic-NCr-nu/nu female mice at 6 weeks (Charles River) were inoculated subcutaneously on both flanks with 2×106 cells containing mixed GFP (parental) and RFP (revertants) or RFP (parental) and GFP (revertants) at 1 : 1 ratio. When the tumor volume reached ~0.3 cm3, the tumors were isolated, treated with collagenase D protease (Sigma) to generate single-cell suspension, and analyzed by FACS.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Ebos JM . Prodding the beast: assessing the impact of treatment-induced metastasis. Cancer Res 2015; 75: 3427–3435.
Campisi J . Senescent cells, tumor suppression, and organismal aging: good citizens, bad neighbors. Cell 2005; 120: 513–522.
d'Adda di Fagagna F . Living on a break: cellular senescence as a DNA-damage response. Nat Rev Cancer 2008; 8: 512–522.
Schmitt CA . Cellular senescence and cancer treatment. Biochim Biophys Acta 2007; 1775: 5–20.
Xue W, Zender L, Miething C, Dickins RA, Hernando E, Krizhanovsky V et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 2007; 445: 656–660.
Narita M, Nunez S, Heard E, Narita M, Lin AW, Hearn SA et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 2003; 113: 703–716.
Rodier F, Coppe JP, Patil CK, Hoeijmakers WA, Munoz DP, Raza SR et al. Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion. Nat Cell Biol 2009; 11: 973–979.
Rodier F, Munoz DP, Teachenor R, Chu V, Le O, Bhaumik D et al. DNA-SCARS: distinct nuclear structures that sustain damage-induced senescence growth arrest and inflammatory cytokine secretion. J Cell Sci 2011; 124: 68–81.
Coppe JP, Desprez PY, Krtolica A, Campisi J . The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu Rev Pathol 2010; 5: 99–118.
Roberson RS, Kussick SJ, Vallieres E, Chen SY, Wu DY . Escape from therapy-induced accelerated cellular senescence in p53-null lung cancer cells and in human lung cancers. Cancer Res 2005; 65: 2795–2803.
Chakradeo S, Elmore LW, Gewirtz DA . Is senescence reversible? Curr Drug Targets 2016; 17: 460–466.
Yang L, Song T, Chen L, Soliman H, Chen J . Nucleolar repression facilitates initiation and maintenance of senescence. Cell Cycle 2015; 14: 3613–3623.
Sabisz M, Skladanowski A . Cancer stem cells and escape from drug-induced premature senescence in human lung tumor cells: implications for drug resistance and in vitro drug screening models. Cell Cycle 2009; 8: 3208–3217.
Achuthan S, Santhoshkumar TR, Prabhakar J, Nair SA, Pillai MR . Drug-induced senescence generates chemoresistant stemlike cells with low reactive oxygen species. J Biol Chem 2011; 286: 37813–37829.
Cruickshanks HA, McBryan T, Nelson DM, Vanderkraats ND, Shah PP, van Tuyn J et al. Senescent cells harbour features of the cancer epigenome. Nat Cell Biol 2013; 15: 1495–1506.
Coppe JP, Patil CK, Rodier F, Sun Y, Munoz DP, Goldstein J et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol 2008; 6: 2853–2868.
Pisco AO, Huang S . Non-genetic cancer cell plasticity and therapy-induced stemness in tumour relapse: 'What does not kill me strengthens me'. Br J Cancer 2015; 112: 1725–1732.
Pisco AO, Brock A, Zhou J, Moor A, Mojtahedi M, Jackson D et al. Non-Darwinian dynamics in therapy-induced cancer drug resistance. Nat Commun 2013; 4: 2467.
El Sharouni SY, Kal HB, Battermann JJ . Accelerated regrowth of non-small-cell lung tumours after induction chemotherapy. Br J Cancer 2003; 89: 2184–2189.
Sharma SV, Lee DY, Li B, Quinlan MP, Takahashi F, Maheswaran S et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 2010; 141: 69–80.
Xu Y, Hu YD, Zhou J, Zhang MH, Yuan WW, Luo Y . shRNA targeting Bmi1 impedes the self-renewal of cisplatin-enriched stem-like cells in human A549 cells. Oncol Rep 2012; 28: 629–639.
Bywater MJ, Poortinga G, Sanij E, Hein N, Peck A, Cullinane C et al. Inhibition of RNA polymerase I as a therapeutic strategy to promote cancer-specific activation of p53. Cancer Cell 2012; 22: 51–65.
Chang J, Wang Y, Shao L, Laberge RM, Demaria M, Campisi J et al. Clearance of senescent cells by ABT263 rejuvenates aged hematopoietic stem cells in mice. Nat Med 2016; 22: 78–83.
Vairo G, Soos TJ, Upton TM, Zalvide J, DeCaprio JA, Ewen ME et al. Bcl-2 retards cell cycle entry through p27(Kip1), pRB relative p130, and altered E2F regulation. Mol Cell Biol 2000; 20: 4745–4753.
Crescenzi E, Palumbo G, Brady HJ . Bcl-2 activates a programme of premature senescence in human carcinoma cells. Biochem J 2003; 375: 263–274.
Mi H, Muruganujan A, Casagrande JT, Thomas PD . Large-scale gene function analysis with the PANTHER classification system. Nat Protoc 2013; 8: 1551–1566.
Huang, da W, Sherman BT, Lempicki RA . Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 2009; 37: 1–13.
Beronja S, Livshits G, Williams S, Fuchs E . Rapid functional dissection of genetic networks via tissue-specific transduction and RNAi in mouse embryos. Nat Med 2010; 16: 821–827.
Beausejour CM, Krtolica A, Galimi F, Narita M, Lowe SW, Yaswen P et al. Reversal of human cellular senescence: roles of the p53 and p16 pathways. EMBO J 2003; 22: 4212–4222.
Huang Q, Li F, Liu X, Li W, Shi W, Liu FF et al. Caspase 3-mediated stimulation of tumor cell repopulation during cancer radiotherapy. Nat Med 2011; 17: 860–866.
Li F, Huang Q, Chen J, Peng Y, Roop DR, Bedford JS et al. Apoptotic cells activate the ‘phoenix rising’ pathway to promote wound healing and tissue regeneration. Sci Signal 2010; 3: ra13.
Janicke RU, Walker PA, Lin XY, Porter AG . Specific cleavage of the retinoblastoma protein by an ICE-like protease in apoptosis. EMBO J 1996; 15: 6969–6978.
Fischer U, Janicke RU, Schulze-Osthoff K . Many cuts to ruin: a comprehensive update of caspase substrates. Cell Death Differ 2003; 10: 76–100.
Zhang H, Pan KH, Cohen SN . Senescence-specific gene expression fingerprints reveal cell-type-dependent physical clustering of up-regulated chromosomal loci. Proc Natl Acad Sci USA 2003; 100: 3251–3256.
Marusyk A, Almendro V, Polyak K . Intra-tumour heterogeneity: a looking glass for cancer? Nat Rev Cancer 2012; 12: 323–334.
Maley CC, Galipeau PC, Finley JC, Wongsurawat VJ, Li X, Sanchez CA et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat Genet 2006; 38: 468–473.
Park SY, Lee HE, Li H, Shipitsin M, Gelman R, Polyak K . Heterogeneity for stem cell-related markers according to tumor subtype and histologic stage in breast cancer. Clin Cancer Res 2010; 16: 876–887.
Acknowledgements
We would like to thank the Moffitt Flow Cytometry Core for assistance to the experiments. This work is supported in part by grants from the National Institutes of Health (CA141244, CA186917), Florida Department of Health (4KB12), and a Miles For Moffitt award from the Moffitt Foundation. H. Lee Moffitt Cancer Center & Research Institute is an NCI designated Comprehensive Cancer Center (P30-CA076292).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
edited by Isaac Harris
Supplementary Information accompanies the paper on the Cell Death Discovery website
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yang, L., Fang, J. & Chen, J. Tumor cell senescence response produces aggressive variants. Cell Death Discov. 3, 17049 (2017). https://doi.org/10.1038/cddiscovery.2017.49
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/cddiscovery.2017.49
This article is cited by
-
Targeting mTOR to overcome resistance to hormone and CDK4/6 inhibitors in ER-positive breast cancer models
Scientific Reports (2023)
-
Cellular uptake and retention studies of silica nanoparticles utilizing senescent fibroblasts
Scientific Reports (2023)
-
A three-marker signature identifies senescence in human breast cancer exposed to neoadjuvant chemotherapy
Cancer Chemotherapy and Pharmacology (2023)
-
Exploiting senescence for the treatment of cancer
Nature Reviews Cancer (2022)
-
Translocation of intracellular CD24 constitutes a triggering event for drug resistance in breast cancer
Scientific Reports (2021)