Abstract
Inhibition of protein neddylation, particularly cullin neddylation, has emerged as a promising anticancer strategy, as evidenced by the antitumor activity in preclinical studies of the Nedd8-activating enzyme (NAE) inhibitor MLN4924. This small molecule can block the protein neddylation pathway and is now in clinical trials. We and others have previously shown that the antitumor activity of MLN4924 is mediated by its ability to induce apoptosis, autophagy and senescence in a cell context-dependent manner. However, whether MLN4924 has any effect on tumor angiogenesis remains unexplored. Here we report that MLN4924 inhibits angiogenesis in various in vitro and in vivo models, leading to the suppression of tumor growth and metastasis in highly malignant pancreatic cancer, indicating that blockage of angiogenesis is yet another mechanism contributing to its antitumor activity. At the molecular level, MLN4924 inhibits Cullin–RING E3 ligases (CRLs) by cullin deneddylation, causing accumulation of RhoA at an early stage to impair angiogenic activity of vascular endothelial cells and subsequently DNA damage response, cell cycle arrest and apoptosis due to accumulation of other tumor-suppressive substrates of CRLs. Furthermore, we showed that inactivation of CRLs, via small interfering RNA (siRNA) silencing of its essential subunit ROC1/RBX1, recapitulates the antiangiogenic effect of MLN4924. Taken together, our study demonstrates a previously unrecognized role of neddylation in the regulation of tumor angiogenesis using both pharmaceutical and genetic approaches, and provides proof of concept evidence for future development of neddylation inhibitors (such as MLN4924) as a novel class of antiangiogenic agents.
Similar content being viewed by others
Main
Neddylation, a type of protein posttranslational modifications, is a process of addition of the ubiquitin-like molecule Nedd8 to a protein substrate. This process is catalyzed by a cascade of the Nedd8-activating enzyme E1, Nedd8-conjugating enzyme E2 (UBC12) and substrate-specific Nedd8-E3 ligases.1, 2 This modification can alter the conformation, stability or localization of a protein, consequently regulating its function.1, 2 The best-known substrates of neddylation are a group of proteins called cullins that serve as the essential components of multiunit cullin–RING E3 ligases (CRLs).3, 4 CRLs are the largest cellular ubiquitin ligase family that controls the turnover of numerous proteins with fundamental roles in cancer biology.5, 6 Tumor suppressor p53 and oncoprotein Hu antigen R (HuR) were recently identified as new neddylation substrates.7, 8 Nedd8 conjugation inhibited p53 transcriptional activity,7 but stabilized HuR.8 These findings highlight a critical role of neddylation in tumor development and progression, but the molecular basis involved in this process still remains to be studied.
Inhibition of protein neddylation has emerged as a novel anticancer strategy.9, 10, 11 The Nedd8-activating enzyme (NAE) inhibitor MLN4924, that inhibits protein neddylation, was discovered as the first anticancer agent in this class by high-throughput screening.12, 13 Mechanistically, MLN4924 inhibits NAE activity by binding to NAE at its active site to form a covalent Nedd8-MLN4924 adduct that cannot move forward to subsequent intraenzyme reactions.14 As a result, MLN4924 can abrogate cullin neddylation, inactivate CRL and thus cause accumulation of CRL substrates that could eventually trigger a DNA damage response, cell cycle arrest, apoptosis and/or senescence in a broad panel of cancer cells.12, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25 In addition, MLN4924 has also been identified as a promising chemoradiosensitizer in preclinical studies.18, 19, 26, 27, 28 Because of its significant anticancer efficacy and well-tolerated toxicity in preclinical studies, MLN4924 is currently in several phase I clinical trials for cancer therapy.10, 11
Abnormal angiogenesis, a feature of solid tumors,29, 30 promotes tumor growth and metastasis, and thus disrupting this process serves as a promising strategy for cancer therapy.29, 31, 32 Although the effect of MLN4924 on the malignant phenotypes of cancer cells has been intensively investigated, its efficacy on tumor angiogenesis and the mechanism(s) underlying this effect remain unexplored. In an attempt to test this idea, we carried out a set of angiogenesis assays in vitro and in vivo, and found that MLN4924 indeed suppresses tumor angiogenesis by inhibiting cullin neddylation and inactivating CRL E3 ligases. As detailed below, our study identified neddylation as a molecular event important for tumor angiogenesis and also provided evidence for targeting this process, such as with MLN4924, as a novel antiangiogenic cancer therapy.
Results
MLN4924 inhibits angiogenesis in classical in vitro and in vivo angiogenesis assays
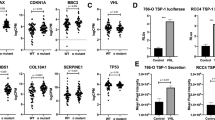
To investigate the role of neddylation in the regulation of angiogenesis, we first determined the effect of neddylation inactivation with MLN4924 using the in vitro rat aortic ring assay that recapitulates all of the key steps of angiogenesis (matrix degradation, migration, proliferation and reorganization). As shown in Figure 1a, new blood vessel growth from rat aortic rings was strongly inhibited upon treatment with MLN4924. We then performed a chick embryo chorioallantoic membrane (CAM) angiogenesis assay. The formation of capillary vessels was significantly suppressed by MLN4924 with a decrease of visible blood vessel branch points (Figure 1b). Finally, we evaluated the antiangiogenic effect of MLN4924 using the Matrigel plug assay that is widely used to detect newly formed blood vessels in vivo. As shown in Figure 1c, MLN4924 strongly inhibited VEGF-induced Matrigel plug angiogenesis. Immunostaining for CD31, a specific marker of vascular endothelial cells, further demonstrated that MLN4924 significantly reduces the microvessel density of the embedded Matrigel plug (Figure 1c). These findings observed from multiple classical angiogenic assays demonstrate that MLN4924 supresses angiogenesis by inhibiting protein neddeylation.
MLN4924 inhibits angiogenesis in multiple angiogenic assays. (a) Rat aortic ring angiogenesis assay. The aortic rings from 6-week-old male Sprague–Dawley rats were randomly seeded into Matrigel-coated wells and sealed with an overlay of Matrigel. VEGF in serum-free ECGM, with or without MLN4924, was added into the wells. After 6 days, microvessel sprouting was fixed and photographed using an inverted microscope (Olympus; magnification × 100) and vascular areas of each treatment were calculated (n=6). (b) Chick embryo chorioallantoic membrane (CAM) assay. The top air sac portions of fertilized chicken eggs were opened and sterilized filter disks with various concentrations of MLN4924 were placed on regions of the CAM relatively deficient in preexisting blood vessels. At the end of analysis, CAMs were photographed with a digital camera and angiogenesis inhibition was assessed by the number of visible blood vessel branch points in each group and expressed as a percentage of negative control (n=6). (c) In vivo Matrigel plug assay. Matrigel containing indicated amounts of MLN4924, VEGF and heparin were subcutaneously injected into the ventral area of 6-week-old C57BL/6 mice. After 6 days, Matrigel plugs were collected and photographed (c, upper panel). Plug sections were immunostained with a specific anti-CD31 antibody for microvessel density assay (c, bottom panel) (n=6). Data were presented as mean±S.E.M. *P<0.05, **P<0.01 and ***P<0.001
MLN4924 suppresses tumor angiogenesis and progression in a footpad xenograft model of pancreatic cancer
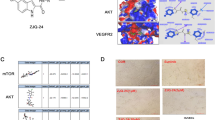
After establishing the antiangiogenic efficacy of MLN4924 in multiple classical assays, we evaluated the efficacy of MLN4924 on in vivo tumor angiogenesis and growth. To allow the noninvasive fluorescent imaging of tumor angiogenesis and progression, we inoculated RFP-expressing pancreatic MiaPaCa2 cells (MiaPaCa2-RFP) into the footpad of GFP transgenic nude mice, as described previously.33 As shown in Figure 2a, neovessels were clearly observed on the surface of control tumors, but not in MLN4924-treated tumors, indicating the strong inhibition of tumor angiogenesis by MLN4924 treatment. As a result, MLN4924 significantly suppressed the growth of primary pancreatic tumors (Figure 2b). At the end of the treatment, tumor tissues were collected, photographed and weighed (Figure 2c). Consistently, control tumors were much larger and weighed more than MLN4924-treated tumors (Figure 2c). Microvessel density analysis by CD31 staining further revealed that MLN4924 significantly inhibits tumor angiogenesis (Figure 2d). These findings indicate that MLN4924 exerts a strong suppressive effect on tumor angiogenesis and tumor growth of highly malignant pancreatic cancer.
MLN4924 suppresses tumor angiogenesis and progression in a mouse footpad model of human pancreatic cancer. Human MiaPaCa-2-RFP pancreatic cancer cells were inoculated into the footpads of GFP transgenic nude mice, treated with 60 mg/kg MLN4924 or 10% HPBCD as negative control subcutaneously twice a day, and subjected to angiogenesis and tumor growth assays over time. At 10 days post treatment, the status of tumor angiogenesis of treated mice was determined by noninvasive real-time optical imaging (white arrows show blood vessels) (a) and the tumor volume was measured (b). At the end of treatment, the tumors were collected, photographed and weighed (left, bright field; right, fluorescent imaging) (c) and tumor tissue sections were immunostained with a specific anti-CD31 antibody for microvessel density assay (d). Data are presented as mean±S.E.M. **P<0.01 and ***P<0.001
MLN4924 suppresses tumor angiogenesis, progression and metastasis in orthotopic models of pancreatic cancer
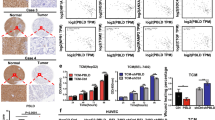
We then evaluated the efficacy of MLN4924 on tumor angiogenesis and tumor progression in clinically relevant orthotopic models of MiaPaCa2-RFP pancreatic cancer. The genetically manipulated fluorescent MiaPaCa2-RFP tumors allow us to determine the kinetic growth and progression of tumors in real time by external and noninvasive whole-body optical imaging.34, 35 As shown in Figure 3a, MLN4924 significantly suppressed pancreatic tumor growth. Consistently, control pancreatic tumors were significantly weighed more than that of MLN4924-treated tumors at the end of the treatment (Figure 3b). Moreover, we found that lymph node metastasis of the pancreatic tumors was also inhibited by MLN4924 drastically (Figure 3c). Finally, the antiangiogenic activity of MLN4924 on orthotopic pancreatic cancer was further verified by the significant reduction of tumor vessel density in MLN4924-treated tumors as compared with control tumors (Figure 3d). During the entire treatment, no obvious treatment-related toxicity to animals, such as loss of body weight, was observed (data not shown). These findings demonstrate the potent antiangiogenic and anticancer efficacy of MLN4924 in orthotopic pancreatic cancer mouse models.
MLN4924 suppresses tumor angiogenesis, progression and metastasis in an orthotopic pancreatic cancer model. To establish the orthotopic pancreatic cancer model, small pieces of human MiaPaCa-2-RFP pancreatic cancer tissue that originated from subcutaneous tumors of nude mice were inserted into the pancreases of nude mice, as described in the Materials and Methods. The tumor-bearing mice were then treated with MLN4924 or 10% HPBCD as a negative control. Tumor growth, metastasis and angiogenesis were determined (n=10 for each group). (a) Real-time optical imaging of the progression of MiaPaCa-2-RFP orthotopic pancreatic cancer (left panel) and tumor growth curves derived from the intensity of tumor fluorescence signals by ModFit LT software (right panel). (b) At the end of treatment, the primary pancreatic tumors were harvested, photographed under fluorescence (left panel) and weighed (right panel). (c) Lymph node metastases were harvested, photographed (left panel) and weighed (right panel). (d) CD31 staining of primary tumor sections for microvessel density analysis. Data are presented as mean±S.E.M. *P<0.05 and ***P<0.001
MLN4924 inhibits protein neddylation and multiple angiogenic phenotypes of HUVECs
To investigate how MLN4924 affects angiogenesis, we next used human umbilical vein endothelial cells (HUVECs), the most commonly used and easily manipulated cell line for phenotypic and mechanistic studies of angiogenesis. First, we found that all the protein components of the neddylation pathway, including Nedd8-activating enzyme (E1, NAE1 and UBA3 heterodimer), Nedd8-conjugating enzyme (E2, UBC12) and substrate-specific Nedd8-E3 ligase (ROC1 and Dcn-1),36, 37, 38 are expressed (Figure 4a) and functional (Figure 4b) in HUVECs. In addition, MLN4924 exerted its suppressive effect on protein neddylation in cells, as indicated by a significant decrease in the expression of total Nedd8-conjugated (neddylated) proteins, cullin neddylation and by the increase in free Nedd8 (Figure 4b), without obvious disturbance of the expression of neddylation enzymes (Figure 4a). Consequently, the formation of capillary-like tube networks (Figure 4c), transwell migration (Figure 4d) and the migrated distance (Figure 4e) of HUVECs were strongly inhibited by MLN4924 in a dose-dependent manner. These findings indicate that MLN4924 inhibits protein neddylation and angiogenic phenotypes of vascular endothelial cells.
MLN4924 inhibits protein neddylation and multiple angiogenic phenotypes of HUVECs. HUVECs were treated with MLN4924 for 12 h and subjected to immunoblotting analysis of indicated proteins using GAPDH as a loading control. (a) The expression of neddylation enzymes in HUVECs. (b) The inhibition of global protein neddylation and cullin neddylation by MLN4924 in HUVECs. (c) Capillary tube formation of HUVECs was inhibited by MLN4924. (d) MLN4924 inhibited the transwell migration of HUVECs. (e) MLN4924 inhibited HUVEC motility, recorded using time-lapse microscopy. Data are presented as mean±S.E.M. *P<0.05, **P<0.01 and ***P<0.001
Short-term treatment of HUVECs with MLN4924 induces RhoA accumulation to impair the angiogenic activity of cells
To investigate the molecular mechanisms for the antiangiogenic activity of MLN4924, we first measured the expression level of several cellular angiogenesis inhibitors neurofibro matosis type 1 (NF1),39 zinc-finger transcription factor Krüppel-like factor 2 (KLF2),40 and NF-κB inhibitor IκB-α17 upon MLN4924 treatment, as these angiogenic regulators have been reported as potential CRL substrates and may accumulate upon the inhibition of cullin neddylation by MLN4924 in HUVECs. We found that the degradation of IκB-α, but not of NF1 or KLF2, was blocked by MLN4924 treatment, as demonstrated by the accumulation of pIκB-α that led to functional inactivation of the NF-κB signaling as revealed by the suppression of transactivation of classical NF-κB targets IL-6 and TNF-α (Supplementary Figure 1). However, the knockdown of IκB-α by small interfering RNA (siRNA) silencing had no rescue effect on MLN4924-mediated inhibition of capillary tube formation (Supplementary Figure 2), suggesting that the NF-κB signaling is not directly involved in the antiangiogenic effect of MLN4924.
RhoA, a member of the Rho GTPase family, has been recently identified as a new substrate of CRL,41 and its accumulation upon CRL inactivation impaired cellular migration. In consideration of our above observation that MLN4924 inhibits the migration of HUVECs (Figure 4), we hypothesized that MLN4924 may regulate angiogenesis by modulating RhoA expression upon CRL inactivation. To test this idea, we determined the level of RhoA upon MLN4924 treatment for 12 h when cell viability was not obviously disturbed (Figure 5a). As shown in Figure 5b, RhoA, but not another Rho family member Cdc42, accumulated upon MLN4924-induced CRL inactivation. More importantly, knockdown of RhoA by siRNA (Figure 5c) significantly rescued the inhibitory effect of MLN4924 on capillary tube formation (Figure 5d) and transwell migration (Figure 5e) of HUVECs, suggesting a critical role of RhoA in MLN4924-induced angiogenesis inhibition.
Short-term treatment with MLN4924 induces RhoA accumulation which inhibits angiogenic activity of HUVECs. (a) Effect of short-term MLN4924 treatment on cell viability. HUVECs seeded in 96-well plates were cultured overnight and treated with MLN4924 at indicated concentrations for 12 h, followed by the ATPlite cell viability assay. (b) MLN4924 treatment induced accumulation of RhoA. HUVECs were treated with MLN4924 for 12 h and subjected to immunoblotting analysis of indicated proteins. (c) Downregulation of RhoA expression via siRNA silencing (siRhoA). (d and e) Knockdown of RhoA rescued the inhibitory effects of MLN4924 on capillary tube formation (d) and transwell migration (e) of HUVECs. HUVECs transfected with siRhoA or siControl were treated with MLN4924 and subjected to the capillary tube formation assay and transwell migration assay
Prolonged exposure of HUVECs to MLN4924 induces cell cycle arrest and apoptotic cell death
To further determine the potential mechanisms of antiangiogenic effect of MLN4924, the responses of HUVECs to prolonged inactivation of neddylation were investigated. We detected the status of Cullin neddylation and the expression of several classical CRL substrates at 48 h post MLN4924 treatment. As shown in Figure 6a, prolonged exposure of cells to MLN4924 induced significant accumulation of cell cycle inhibitors (p21, p27 and wee1) and DNA replication licensing proteins (CDT1 and ORC1) that triggered DNA damage response (DDR), as demonstrated by the appearance of p-CHK1 and p-H2AX as classical DDR markers. As the accumulation of cell cycle inhibitors and the induction of DNA damage response often trigger cell cycle defect and even induce cell death, we assessed the cell cycle and programmed cell death of HUVECs by performing propidium iodide (PI) staining and fluorescence-activated cell sorting (FACS) analysis. We found that MLN4924 can trigger G2 arrest, as demonstrated by the accumulation of G2/M-phase transition inhibitor Wee1 (Figure 6a) and increased cell populations in G2 phase (Figure 6b). MLN4924 also induced apoptosis as indicated by the appearance of a sub-G1 peak (Figure 6b) and cleaved PARP and cleaved Caspase 3 (Figure 6c) in treated cells. As a result, cell viability was significantly reduced by MLN4924 at 48 h post treatment in a dose-dependent manner (Figure 6d). To further define the molecular events by which MLN4924 induced apoptosis, we analyzed the expression of classical apoptosis-regulatory proteins and found that proapoptotic proteins NOXA, BIK and BIM increase, whereas antiapoptotic proteins C-IAP2 and Bcl-2 decrease significantly upon MLN4924 treatment (Figure 6e). Given the fact that NOXA could serve as a potential substrate of CRL,42 we further investigated whether NOXA knockdown could rescue the effect of MLN4924-induced apoptosis. As shown in Figures 6f and g, siRNA silencing of NOXA markedly attenuated MLN4924-mediated apoptosis and rescued cell survival. Taken together, these results demonstrate that the prolonged use of this inhibitor of protein neddylation can reduce cell viability by inducing cell cycle arrest and apoptosis as well.
Long-term inactivation of neddylation with MLN4924 induces cell cycle arrest, reduces cell viability and increases apoptosis in HUVECs. (a) The accumulation of cell cycle regulators and DNA replication licensing proteins upon MLN4924 treatment for 48 h. (b) HUVECs were treated with MLN4924 for 48 h and subjected to PI staining and FACS analysis. The percentage of cells in G2 phase and sub-G1 phase was indicated. (c) HUVECs were treated with MLN4924 for 48 h and subjected to IB analysis using antibodies against cleaved PARP and cleaved Caspase 3 with GAPDH as a loading control. (d) HUVECs seeded in 96-well plates were cultured overnight and treated with MLN4924 at different concentrations for 48 h, followed by the ATPlite cell viability assay. (e) HUVECs were treated with MLN4924 for 48 h and subjected to IB analysis using antibodies against apoptosis-regulatory proteins with GAPDH as a loading control. (f and g) Knockdown of NOXA attenuated apoptotic activation (f) and proliferation inhibition (g) upon MLN4924 treatment. HUVECs transfected with siNOXA or siControl were treated with MLN4924 for 48 h and subjected to IB analysis for indicated proteins and the ATPlite cell viability assay. Data are presented as mean±S.E.M. **P<0.01 and ***P<0.001
Genetic inactivation of CRL via ROC1/RBX1 siRNA silencing recapitulates the antiangiogenic effect of MLN4924 in HUVECs
Our present data suggest that the antiangiogenic effect of MLN4924 is mainly mediated by inactivation of the CRL E3 ligase, the best-known neddylation substrate. The core structure of CRL is a complex of ROC1/RBX1-cullins, in which ROC1/RBX1 interacts with all seven cullin family members to activate CRL.43, 44 To further validate the role of CRL inactivation in facilitating the antiangiogenic effect of MLN4924, we inactivated CRL by knocking down the expression of ROC1/RBX1 via specific siRNAs43, 44 and determined whether ROC1 knockdown could recapitulate the antiangiogenic effect of MLN4924. Similar to MLN4924 treatment, knockdown of ROC1 significantly inhibited capillary tube formation and transwell migration of HUVECs (Figures 7a and b). Moreover, ROC1 siRNA silencing also induced G2 arrest and apoptosis (Figures 7c and d), thus inhibiting the proliferation of HUVECs (Figure 7e). Finally, we found that CRL inactivation by ROC1 knockdown similarly led to the accumulation of CRL substrates that include (1) RhoA that inhibits cell migration; (2) cell cycle inhibitors p21, p27 and wee1 that lead to cell cycle arrest; and (3) DNA replication licensing proteins CDT1 and ORC1 that can cause DNA damage response (Figure 7f). These results demonstrate that the antiangiogenic effect of MLN4924 is indeed mediated through inactivation of CRL, but not other neddylation substrates.
CRL inactivation via ROC1/RBX1 siRNA silencing inhibits angiogenic activity of HUVECs. HUVECs were transfected with siROC1 or siControl and subjected to the capillary tube formation assay (a), transwell migration assay (b), PI staining and FACS analysis (c), PARP activation detection (d), ATPlite cell viability assay (e) and IB analysis of indicated proteins (f). Data was presented as mean±S.E.M. **P<0.01 and ***P<0.001
Discussion
Tumor angiogenesis, as a feature of solid cancers29, 30 that promotes tumor progression and metastasis, and is tightly regulated by a number of molecular events.45, 46, 47 Here, we describe a study that shows that protein neddylation also plays a role in abnormal angiogenesis, since targeting this protein modification with pharmacological and siRNA agents can suppress tumor angiogenesis. First, we demonstrate that inactivation of neddylation by the NAE inhibitor MLN4924 not only significantly inhibits abnormal angiogenesis in various classical models in vitro and in vivo, but also suppresses tumor angiogenesis, progression and metastasis in human pancreatic cancer xenografts, a highly vascularized and malignant neoplasm. In addition, our finding that the activation of the entire neddylation pathway (expression of neddylation enzymes and global neddylation) in vascular endothelial cells further renders the feasibility to develop new neddylation inhibitors, such as inhibitors against neddylation E2 or E3, as novel antiangiogenic agents.
MLN4924, a first-in-class inhibitor of NAE that inhibits protein neddylation, has been proved effective in numerous preclinical studies as a single anticancer agent or one component in combination with chemoradiotherapy.10, 11, 12, 18, 19 Previous mechanism studies focused on the efficacy of MLN4924 on cancer cells, leading to the findings that MLN4924 treatment induced DNA damage response, cell cycle arrest, autophagy, apoptosis and/or senescence in vitro and in vivo.12, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25 In this study, we provide direct evidence that demonstrates that MLN4924 can also suppress angiogenesis as a new mechanism of its antitumor activity. While our article was under review, Tan et al.48 also reported the inhibitory effects of MLN4924 on tumor angiogenesis to support their major findings that genetic deletion of Sag/Rbx2/Roc2 E3 ubiquitin ligase blocked angiogenesis. In their study, mouse endothelial MS-1 cells and the B16F10 mouse melanoma model were used, whereas in our study, the more physiologically relevant HUVECs and orthotopic models of human pancreatic cancer were applied. Moreover, we also performed several other classical in vivo angiogenesis assays (aortic ring, CAM and Matrigel plug) and intensive mechanism investigations to thoroughly determine the antiangiogenesis effects of MLN4924. Taken together, these findings offer an impetus for further development of MLN4924, the first investigational neddylation inhibitor, as a potential antiangiogenic agent.
At the molecular level, the activity of MLN4924 against angiogenesis is largely mediated by inactivating CRL E3 ligase and triggering several antiangiogenic pathways through the accumulation of different sets of CRL substrates (Figure 8). At early stages post treatment (e.g., 12 h), when cell viability is not obviously disturbed, the antiangiogenic activity of MLN4924 is to a large extent attributed to the accumulation of RhoA, a CRL substrate, since knockdown of RhoA markedly rescued MLN4924-induced angiogenic inhibition. In contrast, prolonged exposure of HUVECs to MLN4924 triggered DNA damage response, cell cycle arrest and apoptosis by inducing the accumulation of other tumor-suppressive CRL substrates. Importantly, the antiangiogenic effect of MLN4924 could be recapitulated by genetic inactivation of CRL via siRNA knockdown of ROC1/RBX1, an essential cullin-associated subunit of CRL. Together with previous reports that MLN4924 suppresses the growth of cancer cells through inactivation of CRL,9, 10, 11 we conclude that similar mechanisms of growth suppression are shared by cancer cells and vascular endothelial cells upon neddylation inhibition, which provides a molecular basis for the further development of MLN4924 as a dual-targeting anticancer agent.
Working model. Inhibition of neddylation pathway with MLN4924 suppresses angiogenesis by inducing cullin deneddylation, CRL inactivation and accumulation of CRL substrates that impairs migration, proliferation and survival of vascular endothelial cells. Genetic inactivation of CRL via ROC1/RBX1 siRNA silencing recapitulates the antiangiogenic effect of MLN4924. N8, Nedd8; SR, substrate receptor; DDR, DNA damage response
MLN4924 is currently in several phase I clinical trials for cancer therapy due to its significant anticancer efficacy and well-tolerated toxicity in preclinical studies.10, 11 The potential mechanisms for the tolerated toxicity of MLN4924 still remain elusive. Previous studies demonstrated that proliferating cells are much more sensitive to MLN4924 when compared with quiescent cells, and this may provide a potential explanation for the low side effects of MLN4924.12, 23 Future extensive investigations are directed to address why normal tissues and vasculature well tolerate MLN4924 treatment.
In conclusion, our study reveals a critical role of posttranslational neddylation in the regulation of tumor angiogenesis and validates this pathway as a novel antiangiogenic target. We also provide the experimental evidence for future development of neddylation inhibitors, such as MLN492, as a new class of antiangiogenic agents. In addition, the elucidation of the action of MLN4924 by regulating CRL activity provides a molecular basis for the possible development of new neddylation inhibitors as anticancer therapies. Our study also offers clues for further studying the regulatory role of MLN4924 and its detailed mechanisms in normal and cancer angiogenesis. The application of animal models with genetic deletion of neddylation components, specifically in vascular endothelial cells within tumor microenvironment may facilitate future research on the role of neddylation in the development and progression of solid tumors. Finally, our study suggests that it might be necessary to further evaluate the status of tumor angiogenesis upon MLN4924 treatment in clinical trials.
Materials and Methods
Cell line and reagents
MLN4924 was synthesized as previously described.15, 49 For in vitro studies, MLN4924 stock solution (10 mM) was prepared in dimethyl sulfoxide (DMSO) and stored at −20°C as small aliquots until needed. For in vivo studies, MLN4924 was freshly dissolved in 10% 2-hydroxypropyl-β-cyclodextrin (HPBCD) and stored in the dark at room temperature before use.15 HUVECs were purchased from Allcells Biotech Co. (Shanghai, China) and cultured in a complete endothelial cell medium (Allcells Biotech Co.) at 37°C with 5% CO2.
Rat aortic ring assay
The rat aortic ring assay was performed as described previously.40, 50 In brief, each well of a 48-well plate was coated with 120 μl Matrigel and polymerized in an incubator. Aortas isolated from 6-week-old male Sprague–Dawley rats were cleaned of periadventitial fat and connective tissues in cold phosphate-buffered saline (PBS) and then cut into rings with a circumference of 1–1.5 mm. The aortic rings were randomly placed into wells and sealed with a 100 μl overlay of Matrigel. VEGF in 500 μl of serum-free endothelial cell growth medium (ECGM), with or without MLN4924, was added into each well. Fresh medium was exchanged every 2 days. After 6 days, microvessel sprouting was fixed and imaged using an inverted microscope (Olympus, Tokyo, Japan; magnification × 100).
CAM assay
The CAM assay was performed according to the methods of Brooks et al.51, 52 with some modifications. Briefly, the top air sac portions of 10-day-old fertilized chicken eggs were opened for access to the CAM. Sterilized 5-mm diameter filter (Whatman, Kent, UK) disks with various concentrations of MLN4924 (0, 0.33 and 1 μM) were applied to the areas of the CAM relatively deficient in preexisting blood vessels (n=6 for each group). At the end of the assay, the embryos were fixed in 3.7% formaldehyde at 37°C for 30 min and associated discs were cut out and immersed in PBS. For analysis, CAMs were photographed by a digital camera. Angiogenesis inhibition was assessed by the number of visible blood vessel branch points in each group and expressed as a percent of negative control (DMSO).
In vivo Matrigel plug assay
The Matrigel plug assay was performed as described previously.40, 53 Briefly, 0.5 ml Matrigel containing indicated amounts of MLN4924, 100 ng of VEGF1 (Peprotech, Rocky Hill, NJ, USA) and 20 units of heparin was subcutaneously injected into the ventral area of 6-week-old C57BL/6 mice (n=6 for each group). After 6 days, the skin of each mouse was pulled back to expose an intact Matrigel plug that was collected and photographed. Plug sections of 5 μm were immunostained with a specific anti-CD31 antibody to determine the formation of blood vessels.
Pancreatic xenograft nude mouse footpad model
Human MiaPaCa2 pancreatic cancer cells, stably transfected with red fluorescent protein (RFP), were inoculated into the footpad of the GFP transgenic nude mouse. On the day of inoculation, HPBCD or 60 mg/kg MLN4924 was applied subcutaneously twice a day, lasting for 15 days (4 mice/group). RFP-expressing tumors and neovessels were photographed by Flour vivo model 300 (INDEC BioSystems, Santa Clara, CA, USA) at indicated days.
Orthotopic xenograft model of pancreatic cancer
The orthotopic xenograft mouse model of cancer was established by AntiCancer Biotech (Beijing, China) as described previously.54 Briefly, MiaPaCa-2-RFP pancreatic cancer tissue, that previously grew subcutaneously in nude mice, was harvested and carefully inspected to remove necrotic tissue. The harvested tumor tissue was then equally divided into small pieces of 1 mm3. For MiaPaCa-2-RFP pancreatic cancer models, one 1-mm piece of the above tumor tissue fragment was inserted into an incision on the distal pancreatic part of each mouse. Each tumor-bearing mouse model was randomized into two groups (10 animals/group) and treated with 10% HPBCD or MLN4924 (30 mg/kg), respectively, twice a day on a 3-days-on/2-days-off schedule.12
Tumor growth and progression were measured by a whole-body fluorescence imaging system twice a week, as described.34, 35 Whole-body imaging of tumor-bearing animals was conducted with an Olympus OV100 imaging system with 470 nm excitation light originating from an MT-20 light source. Emitted fluorescence was collected through appropriate filters configured on a filter wheel with a DP70 CCD camera, and processed for contrast and brightness with Paint Shop Pro 8 (Corel, Ottawa, ON, Canada). At the end of the study, tumor tissues of mice were collected, photographed and weighed. Parts of the tumor tissues were fixed in paraformaldehyde, embedded with paraffin and sectioned for CD31 immunostaining. All procedures were performed in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Real-time live cell imaging
HUVECs were cultured in a 37°C chamber on a TCS SP5 confocal microscope (Leica, Bensheim, Germany) equipped with a live-cell imaging workstation. The acquired image sequences were analyzed by ImageJ (National institute of Health, Bethesda, MD, USA).40 To examine cell migration, migrating cells were imaged at 5 min intervals to generate plots of cell migration distance over time.
Endothelial cell capillary-like tube formation assay
Growth factor-reduced Matrigel (BD Biosciences, San Jose, CA, USA) was pipetted into prechilled 24-well culture plates (100 μl/well) and polymerized for 30 min at 37°C. HUVECs (4 × 104) with various concentrations of MLN4924 were seeded onto the surface of the Matrigel. After 12 h of incubation, capillary tube formation was observed and digitized with an inverted photomicroscope (Olympus). The tubular networks were quantified using ImageJ.40
Endothelial cell transwell migration assay
The chemotactic motility of the HUVECs was assayed using transwell chambers (Corning, Corning, NY, USA). Briefly, the top chambers were seeded with 2 × 104 HUVECs in 100 μl serum-free medium containing different concentrations of MLN4924. The bottom chambers were filled with 600 μl complete medium containing 10% FBS. Cells were allowed to migrate for 12 h. Stationary cells on the top surface of the membrane were scraped with a cotton swab, and the cells that migrated were fixed with methanol for 30 min. After being washed three times with PBS, the cells were stained with 1% crystal violet. Images were taken using an inverted microscope (Olympus), and the cells that migrated to four independent areas per filter were counted manually.40
PI staining and FACS analysis
Cells treated with MLN4924 or DMSO were harvested and fixed in 70% ethanol at −20°C overnight, and stained with PI (36 μg/ml, Sigma, St. Louis, MO, USA) containing RNAase (50 μg/ml, Sigma) for 15 min, and then analyzed for apoptosis and cell cycle profile by CyAn ADP (Beckman Coulter, Fullerton, CA, USA). Apoptosis was measured by the percentage of cells in the sub-G1 population. Data were analyzed with ModFit LT software (Verity Software House, Topsham, ME, USA).43
ATPlite cell viability assay
Cells seeded in 96-well plates with 2500 cells per well were treated with MLN4924 or DMSO for the indicated times. Cell viability was determined using the ATPlite kit (PerkinElmer, Norwalk, CT, USA) following the manufacturer’s instructions.55
SiRNA silencing
HUVECs were transfected with siRNA oligonucleotides using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). The sequences of siRNAs are as follows: For RhoA, siRhoA: 5′-GAAGUCAAGCAUUUCUGUCTT-3′; for NOXA, siNOXA: 5′- GGUGCACGUUUCAUCAAUUUGTT-3′;56, 57 for ROC1: siROC1-1: 5′-GACUUUCCCUGCUGUUACCUAATT-3′, siROC1-2: 5′-GGACAACAGAGAGUGGGAATT-3′;43 for siIκB-α: siIκB-α-1: 5′-GGCUUUCCUCAACUUCCAGAA-3′, siIκB-α-2: 5′-CUCCGAGACUUUCGA GGAA-3′;58 for control scrambled siRNA, siControl: 5′-UUCUCCGAACGUGUCACGUTT-3′.43 All above siRNAs were purchased from GenePharma (Shanghai, China).
Immunoblotting
Cell lysates were prepared for immunoblotting using antibodies against Wee1, cullin-1, NF1 (Santa Cruz Biotechnology, Santa Cruz, CA, USA), ORC1, P27, cleaved Caspase 3, cleaved PARP, CDT1, H2AX, p-H2AX, CHK1, p-CHK1, Bcl-2,Bcl-xL, Mcl-1, c-IAP1, c-IAP2, Survivin, XIAP, BAX, Bik, Bim, BID, Bak, ROC1 (Cell Signaling Technology, CST, Danvers, MA, USA), IκB-α, p-IκB-α, P21, RhoA, Cdc42 (Epitomics, Burlingame, CA, USA), NOXA (CALBIOCHEM, San Diego, CA, USA) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH; Sigma).
Statistical analysis
Data are presented as mean±S.E.M. and the statistical significance of differences between groups was assessed using GraphPad Prism5 software (GraphPad Software, Inc., San Diego, CA, USA). Student’s t-test was used for the comparison of parameters between two groups. A P-value of <0.05 was considered to be significant. For all tests, three levels of significance (*P<0.05, **P<0.01, ***P<0.001) were applied.
Abbreviations
- CAM:
-
chick embryo chorioallantoic membrane
- CRL:
-
cullin–RING E3 ligases
- DDR:
-
DNA damage response
- DMSO:
-
dimethyl sulfoxide
- FACS:
-
fluorescence-activated cell sorting
- HUVEC:
-
human umbilical vein endothelial cell
- KLF2:
-
zinc-finger transcription factor Krüppel-like factor 2
- NAE:
-
Nedd8-activating enzyme
- NF1:
-
neurofibro matosis type 1
- PI:
-
propidium iodide
- siRNA:
-
small interfering RNA
References
Rabut G, Peter M . Function and regulation of protein neddylation. 'Protein modifications: beyond the usual suspects' review series. EMBO Rep 2008; 9: 969–976.
Xirodimas DP . Novel substrates and functions for the ubiquitin-like molecule NEDD8. Biochem Soc Trans 2008; 36 (Pt 5): 802–806.
Sakata E, Yamaguchi Y, Miyauchi Y, Iwai K, Chiba T, Saeki Y et al. Direct interactions between NEDD8 and ubiquitin E2 conjugating enzymes upregulate cullin-based E3 ligase activity. Nat Struct Mol Biol 2007; 14: 167–168.
Wu JT, Lin HC, Hu YC, Chien CT . Neddylation and deneddylation regulate Cul1 and Cul3 protein accumulation. Nat Cell Biol 2005; 7: 1014–1020.
Nalepa G, Rolfe M, Harper JW . Drug discovery in the ubiquitin-proteasome system. Nat Rev Drug Discov 2006; 5: 596–613.
Jia L, Sun Y . SCF E3 ubiquitin ligases as anticancer targets. Curr Cancer Drug Targets 2011; 11: 347–356.
Xirodimas DP, Saville MK, Bourdon JC, Hay RT, Lane DP . Mdm2-mediated NEDD8 conjugation of p53 inhibits its transcriptional activity. Cell 2004; 118: 83–97.
Embade N, Fernandez-Ramos D, Varela-Rey M, Beraza N, Sini M, Gutierrez de Juan V et al. Murine double minute 2 regulates Hu antigen R stability in human liver and colon cancer through NEDDylation. Hepatology 2012; 55: 1237–1248.
Tanaka T, Nakatani T, Kamitani T . Inhibition of NEDD8-conjugation pathway by novel molecules: potential approaches to anticancer therapy. Mol Oncol 2012; 6: 267–275.
Wang M, Medeiros BC, Erba HP, DeAngelo DJ, Giles FJ, Swords RT . Targeting protein neddylation: a novel therapeutic strategy for the treatment of cancer. Expert Opin Ther Target 2011; 15: 253–264.
Soucy TA, Dick LR, Smith PG, Milhollen MA, Brownell JE . The NEDD8 conjugation pathway and its relevance in cancer biology and therapy. Genes Cancer 2010; 1: 708–716.
Soucy TA, Smith PG, Milhollen MA, Berger AJ, Gavin JM, Adhikari S et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 2009; 458: 732–736.
Nawrocki ST, Griffin P, Kelly KR, Carew JS . MLN4924: a novel first-in-class inhibitor of NEDD8-activating enzyme for cancer therapy. Expert Opin Investig Drugs 2012; 21: 1563–1573.
Brownell JE, Sintchak MD, Gavin JM, Liao H, Bruzzese FJ, Bump NJ et al. Substrate-assisted inhibition of ubiquitin-like protein-activating enzymes: the NEDD8 E1 inhibitor MLN4924 forms a NEDD8-AMP mimetic in situ. Mol Cell 2010; 37: 102–111.
Luo Z, Yu G, Lee HW, Li L, Wang L, Yang D et al. The Nedd8-activating enzyme inhibitor MLN4924 induces autophagy and apoptosis to suppress liver cancer cell growth. Cancer Res 2012; 72: 3360–3371.
Jia L, Li H, Sun Y . Induction of p21-dependent senescence by an NAE inhibitor, MLN4924, as a mechanism of growth suppression. Neoplasia 2011; 13: 561–569.
Milhollen MA, Traore T, Adams-Duffy J, Thomas MP, Berger AJ, Dang L et al. MLN4924, a NEDD8-activating enzyme inhibitor, is active in diffuse large B-cell lymphoma models: rationale for treatment of NF-{kappa}B-dependent lymphoma. Blood 2010; 116: 1515–1523.
Nawrocki ST, Kelly KR, Smith PG, Espitia CM, Possemato A, Beausoleil SA et al. Disrupting protein NEDDylation with MLN4924 is a novel strategy to target cisplatin resistance in ovarian cancer. Clin Cancer Res 2013; 19: 3577–3590.
Wei D, Li H, Yu J, Sebolt JT, Zhao L, Lawrence TS et al. Radiosensitization of human pancreatic cancer cells by MLN4924, an investigational NEDD8-activating enzyme inhibitor. Cancer Res 2012; 72: 282–293.
Pan WW, Zhou JJ, Yu C, Xu Y, Guo LJ, Zhang HY et al. Ubiquitin E3 ligase CRL4CDT2/DCAF2 as a potential chemotherapeutic target for ovarian surface epithelial cancer. J Biol Chem 2013; 288: 29680–29691.
Blank JL, Liu XJ, Cosmopoulos K, Bouck DC, Garcia K, Bernard H et al. Novel DNA damage checkpoints mediating cell death induced by the NEDD8-activating enzyme inhibitor MLN4924. Cancer Res 2013; 73: 225–234.
Mackintosh C, Garcia-Dominguez DJ, Ordonez JL, Ginel-Picardo A, Smith PG, Sacristan MP et al. WEE1 accumulation and deregulation of S-phase proteins mediate MLN4924 potent inhibitory effect on Ewing sarcoma cells. Oncogene 2013; 32: 1441–1451.
Milhollen MA, Narayanan U, Soucy TA, Veiby PO, Smith PG, Amidon B . Inhibition of NEDD8-activating enzyme induces rereplication and apoptosis in human tumor cells consistent with deregulating CDT1 turnover. Cancer Res 2011; 71: 3042–3051.
Swords RT, Kelly KR, Smith PG, Garnsey JJ, Mahalingam D, Medina E et al. Inhibition of NEDD8-activating enzyme: a novel approach for the treatment of acute myeloid leukemia. Blood 2010; 115: 3796–3800.
Lin JJ, Milhollen MA, Smith PG, Narayanan U, Dutta A . NEDD8-targeting drug MLN4924 elicits DNA rereplication by stabilizing Cdt1 in S phase, triggering checkpoint activation, apoptosis, and senescence in cancer cells. Cancer Res 2010; 70: 10310–10320.
Jazaeri AA, Shibata E, Park J, Bryant JL, Conaway MR, Modesitt SC et al. Overcoming platinum resistance in preclinical models of ovarian cancer using the neddylation inhibitor MLN4924. Mol Cancer Ther 2013; 12: 1958–1967.
Yang D, Tan M, Wang G, Sun Y . The p21-dependent radiosensitization of human breast cancer cells by MLN4924, an investigational inhibitor of NEDD8 activating enzyme. PLoS One 2012; 7: e34079.
Kee Y, Huang M, Chang S, Moreau LA, Park E, Smith PG et al. Inhibition of the Nedd8 system sensitizes cells to DNA interstrand cross-linking agents. Mol Cancer Res 2012; 10: 369–377.
Folkman J . Role of angiogenesis in tumor growth and metastasis. Semin Oncol 2002; 29 (6 Suppl 16): 15–18.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Weis SM, Cheresh DA . Tumor angiogenesis: molecular pathways and therapeutic targets. Nat Med 2011; 17: 1359–1370.
Kerbel RS . Tumor angiogenesis. N Engl J Med 2008; 358: 2039–2049.
Yang M, Jiang P, Hoffman RM . Whole-body subcellular multicolor imaging of tumor-host interaction and drug response in real time. Cancer Res 2007; 67: 5195–5200.
Yang M, Baranov E, Moossa AR, Penman S, Hoffman RM . Visualizing gene expression by whole-body fluorescence imaging. Proc Natl Acad Sci USA 2000; 97: 12278–12282.
Hoffman RM, Yang M . Whole-body imaging with fluorescent proteins. Nat Protoc 2006; 1: 1429–1438.
Kurz T, Chou YC, Willems AR, Meyer-Schaller N, Hecht ML, Tyers M et al. Dcn1 functions as a scaffold-type E3 ligase for cullin neddylation. Mol Cell 2008; 29: 23–35.
Kurz T, Ozlu N, Rudolf F, O'Rourke SM, Luke B, Hofmann K et al. The conserved protein DCN-1/Dcn1p is required for cullin neddylation in C. elegans and S. cerevisiae. Nature 2005; 435: 1257–1261.
Morimoto M, Nishida T, Nagayama Y, Yasuda H . Nedd8-modification of Cul1 is promoted by Roc1 as a Nedd8-E3 ligase and regulates its stability. Biochem Biophys Res Commun 2003; 301: 392–398.
Tan M, Zhao Y, Kim SJ, Liu M, Jia L, Saunders TL et al. SAG/RBX2/ROC2 E3 ubiquitin ligase is essential for vascular and neural development by targeting NF1 for degradation. Dev Cell 2011; 21: 1062–1076.
Wang R, Wang Y, Liu N, Ren C, Jiang C, Zhang K et al. FBW7 regulates endothelial functions by targeting KLF2 for ubiquitination and degradation. Cell Res 2013; 23: 803–819.
Chen Y, Yang Z, Meng M, Zhao Y, Dong N, Yan H et al. Cullin mediates degradation of RhoA through evolutionarily conserved BTB adaptors to control actin cytoskeleton structure and cell movement. Mol Cell 2009; 35: 841–855.
Jia L, Yang J, Hao X, Zheng M, He H, Xiong X et al. Validation of SAG/RBX2/ROC2 E3 ubiquitin ligase as an anticancer and radiosensitizing target. Clin Cancer Res 2010; 16: 814–824.
Yang D, Li L, Liu H, Wu L, Luo Z, Li H et al. Induction of autophagy and senescence by knockdown of ROC1 E3 ubiquitin ligase to suppress the growth of liver cancer cells. Cell Death Differ 2013; 20: 235–247.
Jia L, Bickel JS, Wu J, Morgan MA, Li H, Yang J et al. RBX1 (RING box protein 1) E3 ubiquitin ligase is required for genomic integrity by modulating DNA replication licensing proteins. J Biol Chem 2011; 286: 3379–3386.
Adams RH, Alitalo K . Molecular regulation of angiogenesis and lymphangiogenesis. Nat Rev Mol Cell Biol 2007; 8: 464–478.
Ferrara N, Kerbel RS . Angiogenesis as a therapeutic target. Nature 2005; 438: 967–974.
Otrock ZK, Mahfouz RA, Makarem JA, Shamseddine AI . Understanding the biology of angiogenesis: review of the most important molecular mechanisms. Blood Cells Mol Dis 2007; 39: 212–220.
Tan M, Li H, Sun Y . Endothelial deletion of Sag/Rbx2/Roc2 E3 ubiquitin ligase causes embryonic lethality and blocks tumor angiogenesis. Oncogene 2013 e-pub ahead of print 11 November 2013 doi:10.1038/onc.2013.473.
Lee HW, Nam SK, Choi WJ, Kim HO, Jeong LS . Stereoselective synthesis of MLN4924, an inhibitor of NEDD8-activating enzyme. J Org Chem 2011; 76: 3557–3561.
Wang B, Sun J, Kitamoto S, Yang M, Grubb A, Chapman HA et al. Cathepsin S controls angiogenesis and tumor growth via matrix-derived angiogenic factors. J Biol Chem 2006; 281: 6020–6029.
Brooks PC, Montgomery AM, Rosenfeld M, Reisfeld RA, Hu T, Klier G et al. Integrin alpha v beta 3 antagonists promote tumor regression by inducing apoptosis of angiogenic blood vessels. Cell 1994; 79: 1157–1164.
Chen J, Wei D, Zhuang H, Qiao Y, Tang B, Zhang X et al. Proteomic screening of anaerobically regulated promoters from Salmonella and its antitumor applications. Mol Cell Proteomics 2011; 10: M111 009399.
Malinda KM . In vivo matrigel migration and angiogenesis assay. Methods Mol Biol 2009; 467: 287–294.
Hoffman RM . Orthotopic metastatic mouse models for anticancer drug discovery and evaluation: a bridge to the clinic. Invest New Drugs 1999; 17: 343–359.
Bockbrader KM, Tan M, Sun Y . A small molecule Smac-mimic compound induces apoptosis and sensitizes TRAIL- and etoposide-induced apoptosis in breast cancer cells. Oncogene 2005; 24: 7381–7388.
Shan T, Ma J, Ma Q, Guo K, Guo J, Li X et al. beta2-AR-HIF-1alpha: a novel regulatory axis for stress-induced pancreatic tumor growth and angiogenesis. Curr Mol Med 2013; 13: 1023–1034.
Wang Q, Mora-Jensen H, Weniger MA, Perez-Galan P, Wolford C, Hai T et al. ERAD inhibitors integrate ER stress with an epigenetic mechanism to activate BH3-only protein NOXA in cancer cells. Proc Natl Acad Sci USA 2009; 106: 2200–2205.
Tang QL, Xie XB, Wang J, Chen Q, Han AJ, Zou CY et al. Glycogen synthase kinase-3beta, NF-kappaB signaling, and tumorigenesis of human osteosarcoma. J Natl Cancer Inst 2012; 104: 749–763.
Acknowledgements
We thank Dr Yi Sun (University of Michigan) and Dr Avnish Kapoor (University of Texas MD Anderson Cancer Center) for their critical reading of our manuscript. This work was supported by National Basic Research Program of China (973 program, 2012CB910302), National Natural Science Foundation Grant of China (grant numbers 31071204, 81172092 and 81372196), the Program for Professor of Special Appointment (Eastern Scholar) at Shanghai Institutions of Higher Learning and Shanghai Pujiang Talent Program (12PJ1400600) to Lijun Jia.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Agostini
Supplementary Information accompanies this paper on Cell Death and Disease website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Yao, WT., Wu, JF., Yu, GY. et al. Suppression of tumor angiogenesis by targeting the protein neddylation pathway. Cell Death Dis 5, e1059 (2014). https://doi.org/10.1038/cddis.2014.21
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.21
Keywords
This article is cited by
-
Targeting NEDD8-activating enzyme for cancer therapy: developments, clinical trials, challenges and future research directions
Journal of Hematology & Oncology (2023)
-
NEDD8-Activating Enzyme Inhibitor MLN4924 Inhibits Both the Tumor Stroma and Angiogenesis in Pancreatic Cancer via Gli1 and REDD1
Digestive Diseases and Sciences (2023)
-
Mechanical stress shapes the cancer cell response to neddylation inhibition
Journal of Experimental & Clinical Cancer Research (2022)
-
Targeting neddylation E2s: a novel therapeutic strategy in cancer
Journal of Hematology & Oncology (2021)
-
Impact of posttranslational modifications in pancreatic carcinogenesis and treatments
Cancer and Metastasis Reviews (2021)