Abstract
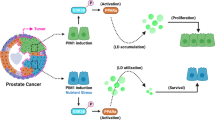
Recent observations indicate prostatic diseases are comorbidities of systemic metabolic dysfunction. These discoveries revealed fundamental questions regarding the nature of prostate metabolism. We previously showed that prostate-specific ablation of PPARγ in mice resulted in tumorigenesis and active autophagy. Here, we demonstrate control of overlapping and distinct aspects of prostate epithelial metabolism by ectopic expression of individual PPARγ isoforms in PPARγ knockout prostate epithelial cells. Expression and activation of either PPARγ 1 or 2 reduced de novo lipogenesis and oxidative stress and mediated a switch from glucose to fatty acid oxidation through regulation of genes including Pdk4, Fabp4, Lpl, Acot1 and Cd36. Differential effects of PPARγ isoforms included decreased basal cell differentiation, Scd1 expression and triglyceride fatty acid desaturation and increased tumorigenicity by PPARγ1. In contrast, PPARγ2 expression significantly increased basal cell differentiation, Scd1 expression and AR expression and responsiveness. Finally, in confirmation of in vitro data, a PPARγ agonist versus high-fat diet (HFD) regimen in vivo confirmed that PPARγ agonization increased prostatic differentiation markers, whereas HFD downregulated PPARγ-regulated genes and decreased prostate differentiation. These data provide a rationale for pursuing a fundamental metabolic understanding of changes to glucose and fatty acid metabolism in benign and malignant prostatic diseases associated with systemic metabolic stress.
Similar content being viewed by others
Main
Benign prostatic hyperplasia (BPH) and prostate cancer (PCa) are age-related diseases associated with complications of metabolic syndrome (MetS).1 However, the molecular underpinnings of prostatic susceptibility to systemic metabolic dysfunction are poorly understood, in part because dietary and transgenic animal models display a limited recapitulation of human benign growth and stromal expansion or adenocarcinoma. Furthermore, unlike adipose, muscle and liver, understanding of the effects of systemic metabolic stressors on prostate growth and/or transformation are hampered by a limited understanding of the prostate’s normal nutritional metabolism.
Epidemiological links between BPH and diabetes have been recognized for many years2 and recent studies have demonstrated that the incidence and severity of BPH are correlated with obesity, atherosclerosis, diabetes mellitus, hyperinsulinemia, hyperglycemia and hypercholesterolemia.3, 4, 5, 6, 7 Although diabetes mellitus has a negative correlation with the incidence of multiple cancers including prostate, diabetic patients exhibit increased mortality.8 Moreover, MetS as a set of comorbidities (obesity, insulin resistance, dyslipidemia and hypertension) is correlated with PCa incidence.9 Such associations have prompted the investigation of metabolic genes and potential metabolic therapies in benign and malignant prostatic diseases.3, 10
The peroxisome proliferator-activated receptors (PPARs) are a family of nuclear fatty acid receptors that regulate tissue-specific cellular metabolism and differentiation and have been widely sought after therapeutic targets for a number of obesity-related metabolic diseases owing to their ability to regulate glucose and fatty acid metabolism.11, 12 A class of PPARgamma (PPARγ) agonists called thiazolidinediones (TZDs) are used in the treatment of insulin resistance and regulate a wide range of genes with tissue-specific effects.13
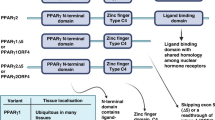
Historically, PPARγ has been associated with pre-adipocyte expansion and differentiation,14 but other tissues also show a functional role for PPARγ, including liver15 and muscle.16 We showed previously that PPARγ ablation in mouse prostate causes tumorigenesis and active autophagy,17, 18 suggesting PPARγ may provide a molecular link between systemic metabolism and prostate differentiation and growth.19 There are two isoforms of PPARγ with the longer PPARγ2 isoform using an alternate transcription start site containing a 30-amino acid N-terminal extension.
Our goal in this study was to garner a fundamental molecular and cellular understanding of the role of PPARγ in mediating metabolic control of prostatic differentiation. Because of the importance of individual PPARγ isoforms in systemic metabolism and our previous work implicating PPARγ in prostate epithelial growth and differentiation, we chose to examine the potential roles of individual PPARγ isoforms in mediating nutrient metabolism in the prostate, which has not been performed in any tissue. A prostatic epithelial cell line (mPrE-PPARγKO) restored with either PPARγ1 or PPARγ2 isoform was used to determine how each isoform might contribute to prostatic metabolism, differentiation and disease. We show, using in vitro analysis, lipidomics and in vivo animal models that PPARγ isoforms control overlapping and distinct metabolic programs in prostate epithelia that lead to functional changes in glucose and lipid metabolism and that these changes are coordinate with reduced lipogenesis, increased β-oxidation and markers of basal and luminal epithelial differentiation. Furthermore, we show in animals that prostate differentiation is oppositely affected after chronic treatment with a TZD versus high-fat diet (HFD) through disparate regulation of PPARγ-and its downstream genes. These data suggest, as in other tissues, that PPARγ agonization may directly or indirectly modulate the nutritional supply of glucose and lipids for prostate metabolism and differentiation.
Results
Restoration of PPARγ2, but not PPARγ1, reverses PPARγKO-induced mouse prostatic carcinogenesis
Alternative transcription start sites and splicing produce two PPARγ isoforms, so only the longer PPARγ2 isoform can be knocked out individually.20 In order to study the independent functions of PPARγ1 and -γ2 isoforms on prostate metabolism and differentiation, we developed a prostate epithelial cell line (mPrE-PPARγKO) with genetic knockout (KO) of both PPARγ1 and -γ2 isoforms from an adult PB-Cre4 tg/0/ PPARγflox/flox double-transgenic male mouse.17 We then restored mPrE-γKO cells with mouse PPARγ1 cDNA (+PPARγ1), PPARγ2 cDNA (+PPARγ2) or an empty vector (+EV) as control, respectively, to create an isogenic series of cell lines for genetic and functional comparisons (see Materials and Methods).
In order to determine the effects of PPARγ isoforms on tissue morphogenesis in vivo, mPrE-γKO and restored cell lines were each recombined with inductive 18-day fetal rat urogenital mesenchyme (UGM) and grafted into the kidney capsule for 2 months (Figures 1a–c). Control mPrE-γKO+EV cells (empty vector-transfected) regenerated high-grade mouse prostatic intraepithelial neoplasia (HGPIN) (Jiang et al.17; Figure 1a) with predominantly Ck8/Ck18+ luminal epithelial glands and few Ck14+ basal cells (Figure 1d). Upon restoration with PPARγ1, large areas of Ck8/18++/Ck14− middle or highly differentiated adenocarcinoma were observed in +PPARγ1 tissue recombinants (Figure 1b, black star, Figure 1e), but large fluid-filled cysts were also formed (Figure 1b, white star). PPARγ2 restoration resulted in the regeneration of Ck8/18+/Ck14++ acini that resembled developing prostate glands without evidence of tumor formation (Figures 1c and f). Furthermore, androgen receptor (AR) expression was demonstrated in regenerated tissues by immunohistochemical staining using +EV or +PPARγ2 cells, but not with +PPARγ1 cells (Figures 1g–i). These data indicate that restoration of PPARγ2 isoform, but not PPARγ1 isoform, reverses PPARγ-deficient mouse prostatic carcinogenesis through an increase in Ck14+ basal cells.
Restoration of PPARγ2, but not PPARγ1, reverses PPARγ KO-induced mouse prostate carcinogenesis. mPrE-PPARγKO+EV, +PPARγ1 or +PPARγ2 cell lines were recombined with inductive rat UGM and grafted under the kidney capsule for 2 months (N=3 each). Histological analysis revealed regeneration of Ck14+/Ck18+/AR+ HGPIN in mPrE-PPARγ PPARγKO+EV grafts (a, d and g). Restoration of PPARγ1 resulted in a mixture of fluid-filled cysts (black star) and CK14−/Ck18+/AR− middle to highly differentiated adenocarcinoma (white star) (b, e and h). Restoration of PPARγ2 resulted in regeneration of Ck14++/Ck18+/AR+ benign acinus formation without any tumors (c, f and i)
PPARγ isoforms 1 and 2 differentially regulate mouse prostate benign epithelial cell differentiation as well as luminal AR expression and function
In order to confirm the in vivo restoration of basal and luminal differentiation by PPARγ2 expression shown in Figure 1, protein expression in mPrE-γKO +EV, +PPARγ1 and +PPARγ2 cells was examined by western blot, which revealed increases in both Ck14 and AR upon expression of PPARγ2 (Figure 2a). To determine whether PPARγ isoform expression increased the differentiation of basal cells in vitro, each mPrE-γKO cell line was double stained in culture for PPARγ and Ck14 (Figure 2b(i–iii)). Results demonstrated that although PPARγ expression in +PPARγ1 cells was increased 14% versus +EV cells, this resulted in an insignificant change in Ck14 expression (Figure 2b (ii versus iii)). Alternatively, and consistent with regeneration experiments in vivo (Figure 1f), PPARγ2 restoration resulted in a 15% increase in PPARγ+ cells as well as a 15% increase in Ck14+ cells (Figure 2b(iii), quantified in Figure 2c). Interestingly, only 6–9% of the PPARγ+ cells had overlapping Ck14 expression, suggesting a potential paracrine regulation of basal cell differentiation. To determine whether the increase in basal cells also resulted in increase luminal differentiation, cells were treated with dihydrotestosterone (DHT) and co-stained for AR and CK14 (Figure 2b(iv-vi)). In PPARγ2-rescued cells, cells adjacent to PPARγ+ cells were observed to have nuclear AR localization (Figure 2b(vi), arrowhead). In contrast, fewer cells were found with nuclear AR localization in +EV or +PPARγ1 cells treated with DHT (Figure 2b(iv versus v)). To confirm increased AR responsiveness, an androgen response element (ARE)-luciferase construct was transfected in each cell line, which demonstrated that +PPARγ2 cells significantly increased AR responsiveness 3-fold, while +PPARγ1 cells had no response and mPrE-γKO+EV cells had a 1.8-fold increase (Figure 2d). The results showed restoration of PPARγ2 rescues and drives mouse prostate benign epithelial cell differentiation associated with AR activation.
PPARγ isoforms 1 and 2 differentially regulate prostate basal differentiation as well as luminal AR expression and function. (a) Western blot analysis of mPrE PPARγKO +EV, +PPARγ1 or +PPARγ2 cells shows dual increase in Ck14 and AR in mPrE PPARγKO +PPARγ2 cells compared with +EV or +PPARγ1 cells. (b) ICC of PPARγ (red) and Ck14 (green) in mPrE-PPARγKO+EV (i), +PPARγ1 (ii) or +PPARγ2 (iii) cells in culture shows an increase in Ck14+ cells in +PPARγ2, but not +PPARγ1 cells. ICC for AR and Ck14 in mPrE-PPARγKO+EV (iv), +PPARγ1 (v) or +PPARγ2 (vi) cells treated with DHT shows cytoplasmic AR in most mPrE-PPARγKO+EV or +PPARγ1 cells, whereas +PPARγ2 cells displayed increased nuclear AR immunoreactivity in cells adjacent to Ck14+ cells. (c) Quantitation of PPARγ immunoreactivity shows a significant increase of 15% for PPARγ-restored cells compared with +EV cells, whereas only PPARγ2-restored cells showed a significant 15% increase in Ck14 immunoreactivity compared with +EV cells. (d) Androgen responsiveness was increased three-fold in +PPARγ2 cells compared with +EV cells as measured by ARE-luciferase (N=3). *P value<0.05
PPARγ isoforms 1 and 2 regulate both overlapping and distinct metabolic networks
In order to determine the potential molecular disparity between PPARγ1- and PPARγ2-driven epithelial differentiation, microarrays were performed on mPrE-PPARγKO+EV versus +PPARγ1 or +PPARγ2 cells. As outlined in Figure 3a, +EV cells minus/plus Rosiglitazone (Rosi) were examined to eliminate PPARγ-independent effects of Rosi (Supplementary Figure 1a). These independent effects were subtracted from results generated from comparison of +EV versus +PPARγ1 and +EV versus +PPARγ2 to identify PPARγ-specific effects of Rosi. Using INGENUITY software (Redwood City, CA, USA) and the significance analysis of microarray (SAM) test for significance, Figure 3b displays the top networks regulated by PPARγ isoforms in prostate epithelial cells. Individual genes regulated by PPARγ1 and PPARγ2 (top 10 up and down-regulated genes shown in Supplementary Figure 1a) included numerous focus molecules with functions related to amino acid, carbohydrate and lipid metabolism, drug metabolism and cellular detoxification as well as inflammation and immunity.
Microarray and network analyses of cell lines. (a) Schematic of microarray analysis. In order to distinguish PPARγ isoform effects downstream of TZD treatment, PPARγ KO cells (mPrE-PPARγKO+EV) were treated with Rosiglitazone (Rosi) and these Rosi-independent results (representing 82 genes, see Supplementary Figure 1a) were subtracted from the downstream regulation demonstrated in Rosi-treated PPARγ1- (358 differentially regulated genes, see Supplementary Figure 1a) or PPARγ2- (400 differentially regulated genes, see Supplementary Figure 1a) rescued cells. Further analysis showed that 230 genes were differentially regulated in PPARγ1 versus PPARγ2-restored PPARγ2-rescued cells. N=3 for each of the four samples. (b) Top networks differentially regulated by PPARγ isoforms using INGENUITY
Restoration of PPARγ1 or -γ2 isoforms reduces lipogenesis/oxidative stress
Microarray data analysis of PPARγ isoform-regulated genes showed a strong upregulation of genes involved in fatty acid metabolism, which has been shown to reduce de novo lipogenesis in some tissues;21 however, the influence of fatty acid metabolism on prostate differentiation has not been examined. Accordingly, western blotting revealed that PPARγ1 or PPARγ2 expression resulted in a decrease in lipogenic pathways (Akt, mTOR, Fasn, Acc) and oxidative stress (Cox-2) (Figure 4a). Flow cytometry of dihydroethidium-stained cell lines confirmed a significant reduction in reactive oxygen species (ROS) upon expression of PPARγ isoforms (Figure 4b). These data suggest that paracrine PPARγ expression satisfies the endogenous lipid needs, thereby negating the need for de novo lipogenesis.
PPARγ isoforms equally decrease de novo lipogenesis and oxidative stress, but differentially modulate triglyceride saturation and glucose metabolism. (a) Ectopic expression of PPARγ1 or PPARγ2 in mPrE-PPARγKO cells resulted in decreased activation of lipogenic pathways including Akt, mTOR, Fasn and Acc. In addition, Cox-2 levels were decreased indicative of lower levels of ROS. (b) Dihydroethidium staining followed by flow cytometry (N=3) confirmed a decrease in ROS. (c) Fatty acid analysis of triglycerides by TLC/MS revealed increased levels and saturation of stearic acid (increased 18 : 0, decreased 18 : 1n9) in PPARγ1-restored cells compared to +EV cells (N=3), consistent with the decreased expression of Scd1 (see Table 1). (d) Glucose/lactate flux analysis in mPrE-PPARγKO+EV, +PPARγ1 or +PPARγ2 cells over 4 days (2 time points/day) demonstrated significantly decreased glucose uptake in mPrE-PPARγKO+PPARγ1 cells and significantly increased lactate secretion in mPrE-PPARγKO+PPARγ2 cells compared to +EV cells. (e) IC50 analysis of the glucose oxidation inhibitor 2DG suggested a reliance of mPrE-PPARγKO+EV cells on glucose in the absence of +PPARγ1 or +PPARγ2 cells, which highly regulate fatty acid transport and metabolism (see Table 1). *P value <0.05
To confirm candidate genes identified by INGENUITY software analysis, qRT-PCR plates were custom-designed for analysis of PPARγ-restored cell lines. Results showed that both PPARγ isoforms regulated genes involved in metabolism (Table 1, Section I), including the modification (Elovl4, Scd1), transport (Cd36, Lpl, Fabp4) and β-oxidation (Acsf2, Lipa, Acot1) of fatty acids. In addition, multiple genes involved in detoxification were upregulated (Table 1, Section II), notably including Cox-2, further confirming the decrease in ROS shown in Figure 4b. Finally, multiple markers of differentiation (Table 1, Section III) were upregulated, notably including regulation of basal cell (Trp63, Ck14) and luminal cell (Pbsn, AR, PTEN) markers by PPARγ2, confirming the changes shown in Figure 1. These data suggest that increased fatty acid import results in a reduction in lipogenesis and oxidative stress.
PPARγ isoforms differentially regulate glucose and fatty acid metabolism
One of the most interesting examples of isoform-specific changes in fatty acid modification genes was the differential regulation of Scd1, which was upregulated by PPARγ2 but downregulated by PPARγ1 (Table 1, Section I). Scd1 is an ER-resident fatty acid desaturase strongly induced by dietary saturated fat and responsible for the production of monounsaturated fatty acids (MUFAs) from 12 to 19 carbon saturated fatty acids, and has been implicated in numerous metabolic diseases. MUFAs are the preferred substrates in the synthesis of major lipid classes including phospholipids, cholesterol esters, wax esters and triglycerides.22, 23
When various lipid classes were analyzed (phospholipids, diglycerides, triglycerides and cholesterol esters) in PPARγ1 and -γ2-restored mPrE-γKO cells, we found that not only were total triglyceride stearic acid levels increased, but also that the PPARγ1-mediated Scd1 decrease resulted in a significantly increased abundance of stearic acid and decreased abundance of oleic acid (Figure 4c). As shown above, PPARγ1 expression also failed to induce prostatic differentiation (Figures 1 and 2), indicating that an increased availability of MUFAs through PPARγ2-mediated Scd1 expression may be beneficial for normal prostate epithelial differentiation.
As for other PPARs,21 we found using qRT-PCR that PPARγ1 and -γ2 also regulated glucose metabolism genes, notably increasing Pdk4 expression (Table 1). Pdk4 phosphorylates and inactivates pyruvate dehydrogenase, resulting in the shunting of pyruvate toward lactate production rather than entry in the mitochondrial tricarboxylic acid (TCA) cycle. In tissues such as muscle, metabolic switching from glucose to fatty acid oxidation is mediated by increased Pdk4 expression.24 To determine whether increased Pdk4 expression resulted in altered glucose/lactate flux, we collected conditioned media from mPrE-PPARγKO and the isoform-restored cells over a 4-day period. Results demonstrated a decrease in glucose flux in PPARγ-restored cells (Figure 4d), coordinate with the level of Pdk4 expression (Table 1), with PPARγ1 mediating the strongest upregulation of Pdk4 and decrease in glucose consumption. A significant increase in lactate production was observed in PPARγ2-restored cells, indicating increased glycolytic metabolism. Furthermore, the increased glucose/lactate ratios observed in both PPARγ-restored cells points to a clear shift away from glucose oxidation in the TCA cycle in favor of lactate production. Measurement of the IC50 of the glucose analog and oxidation inhibitor 2-deoxy-D-glucose (2DG) showed that mPrE-PPARγKO+EV cells were significantly more sensitive to inhibition of glucose metabolism than their PPARγ-restored counterparts (Figure 4e), which are more likely to rely on fatty acid oxidation according to genes shown in Table 1.
These data demonstrate that although restoration of either PPARγ isoform in PPARγ KO HGPIN cells decreases de novo lipogenesis and oxidative stress, only PPARγ2-regulated genes induce a metabolic switch for induction of a prostatic differentiation program, potentially through disparate regulation of Scd1.
TZD or HFD treatment drives opposing effects on mouse prostate metabolism and differentiation in vivo
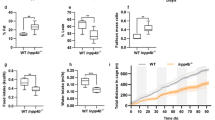
PPARγ has been hypothesized to provide a metabolic link between obesity and tissue dysfunction.13 In order to determine whether TZDs or obesity affect PPARγ-mediated prostatic differentiation in vivo, we fed male mice a Western diet or Rosiglitazone chow for 6 months and examined their prostates for changes in morphology (Figure 5) and PPARγ-regulated genes (Table 2). Significant increases in overall animal weight were detected following HFD treatment only (Figure 5a). Notable increases in smooth muscle density were observed with TZD treatment (Figure 5d), with little epithelial hyperplasia. In addition, intra-muscular adipocytes were also highly enriched in TZD-treated animals (Figure 5d, star), which made clean dissection for RNA analysis of prostate-specific genes extremely difficult. This is consistent with the deposition of intramuscular adipocytes in skeletal muscle of TZD-treated human subjects.25 Although PPARγ2 is thought of as an adipose-specific gene, we were able to demonstrate using a PPARγ2-specific antibody that it is also expressed sporadically in mouse prostate luminal epithelia (Figure 5f) as well as throughout the smooth muscle (Figure 5b).
Administration of TZD or HFD drives opposing effects on mouse prostate metabolism and differentiation in vivo. (a) Western diet-, but not TZD-fed animals significantly increased total weights by 20 g (N=3 each). (b) PPARγ2 is coexpressed in prostate smooth muscle. (c–n) C57B male mice were fed control (a, f, i and l), Rosiglitazone (b, g, j and m) or Western diet chow (e, h, k and n) for 6 months. PPARγ2 levels were similar in prostate epithelia after TZD treatment (f versus g) but decreased in some acini of HFD-treated animals (h, star). Intramuscular adipocytes were increased in TZD-treated animals (g, star), which also resulted in increased AR expression increased in smooth muscle (j, arrow). AR expression was decreased in HFD-treated animals (* in k) similar to qRT-PCR analysis (see Table 2). PDK4 expression was similar in prostate tissue of TZD-treated animals (m, * indicates adipose) and decreased in HFD-treated animals (n) (see Table 2)
As shown in Table 2, large increases in PPARγ-regulated genes were observed in TZD-treated animals. Given the increase in intraprostatic adipocytes after TZD treatment, it was unclear whether these changes were regulated in adipocytes or prostate. Therefore, immunohistochemistry was performed on metabolic proteins including Scd1, Lpl, Cd36, Fabp4 and Pdk4. Although ectopic PPARγ expression was able to regulate some of the fatty acid metabolism genes in vitro (Table 1), immunoreactivity for Fabp4 (Supplementary Figure 1c), Lpl and Scd1 was low in mouse prostate and high in adjacent adipocytes, likely reflecting the strong increases in RNA expression shown in Table 2, Section II. However, immunoreactivity for Cd36 (expressed in epithelia and stroma, Supplementary Figure 1b) and Pdk4 (expressed in epithelia, Figure 5l–n) were strong, suggesting that these genes might be important in directly mediating the metabolism and differentiation of prostate, versus Fabp4 and Lpl, which may indirectly mediate prostate differentiation through adjacent adipocyte fatty acid metabolism.
As PPARγ-regulated genes were so drastically affected in TZD-treated animals, epididymal white adipose tissue (eWAT) gene expression was also examined by qRT-PCR under normal conditions or TZD treatment and was directly compared with that of prostate (Table 2, row 1 versus row 2). Results are compartmentalized into genes predominantly regulated by TZD in either eWAT (I) or prostate (II) or both (III). Contrary to common perceptions of both adipose and prostate, total PPARγ levels (isoforms were not discriminated by qRT-PCR) were higher in prostate than eWAT under normal feeding conditions and AR levels were equivalently expressed (contamination by adipocytes in prostates of animals fed regular chow was minimal). However, under TZD treatment, AR (and probasin) levels increased in prostate and not eWAT. Notably, PPARγ2 is allostatically induced in adipose under HFD,26 which could explain the lower levels of total PPARγ in lean mouse eWAT versus prostate. Recent evidence of the molecular signature of various adipose depots could also explain the difference in PPARγ-regulated genes in eWAT versus the intramuscular adipose of TZD-treated prostate,27 which may indicate the expansion of highly metabolic brown adipose shown previously to be regulated by TZDs.28
Other notable differences between prostate and eWAT gene expression patterns included numerous oxidative stress genes (Cat, Gsta2, Sirt5, Nox1, Nfkb2) and the triglyceride-synthesizing enzyme Dgat2, which were highly enriched in adipose versus prostate. However, a number of fatty acid metabolism (Acsf2, Adipor1, Cd36, Lpl) and differentiation (AR, probasin) genes were equally high in prostate, including the glucose oxidation inhibitor, Pdk4, which suggests that the differentiation induced by PPARγ expression shown here may represent a metabolically regulated program of prostatic differentiation in vivo.
Given the regulation of prostate differentiation by PPARγ2 in mouse, we also wanted to confirm the expression of PPARγ2 and some of its downstream-regulated genes in human prostate tissue. Figure 6a shows that PPARγ2 is highly enriched in prostate smooth muscle and that Scd1 (upregulated by PPARγ2 expression in vitro) is highly enriched in prostate basal cells (Figure 6b). Furthermore, CD36 was expressed in both epithelia and smooth muscle (Figure 6c), whereas PDK4 was expressed predominantly in epithelia (Figure 6d). The compartmentalization of cellular glucose and fatty acid metabolism (Figure 6e) suggests a model of stromal–epithelial, as well as basal–luminal interactions whose disruption by systemic metabolic disease may adversely affect the health and differentiation of prostate (Figure 6f). These results suggest that PPARγ is a major metabolic regulator in the control of mouse and human prostate differentiation.
PPARγ-regulated genes in human prostate tissues. (a) Double staining for PPARγ2 and alpha smooth muscle actin (α-SMA) by immunofluorescence revealed colocalization in a subset of smooth muscle. (b) SCD1 expression is enriched in prostate basal epithelia by immunohistochemical staining (IHC). (c) CD36 is expressed in smooth muscle and epithelia. (d) PDK4 is expressed predominantly in the epithelium. (e) Cellular model of metabolic genes and functions regulated by PPARγ. (f) Tissue interaction model of the potential role of paracrine fatty acid metabolism in regulating prostate differentiation and diseases as comorbidities of diabetes and obesity
Discussion
A recent study of the global prevalence of glycemia and diabetes demonstrated an increase from 153 million affected individuals in 1980 to 347 million in 2008,29 which, according to epidemiological correlations, will likely have a major direct impact on prostate disease incidence. Upon maximal lipid storage capacity of white adipose tissue (WAT), peripheral tissues begin to store lipid in excess of their natural oxidative or storage capacity resulting in lipotoxicity, inflammation and eventually insulin resistance.30 Recent evidence squarely positions prostatic diseases as sequelae of systemic metabolic dysfunction, including hyperinsulinemia, hyperglycemia and hypercholesterolemia31; however, the underlying etiologies of such susceptibilities remain unknown largely because of the absence of a molecular understanding of the basic metabolic machinery governing prostatic function.
Here, we demonstrate that expression of PPARγ2 drives benign prostate epithelial cell differentiation. In mouse prostate PPARγ2 was expressed in both smooth muscle and epithelium (Figures 5b and f), whereas in human prostate PPARγ2 expression seems to be restricted to smooth muscle (Figure 6a). Stromal–epithelial interactions have long been recognized to have a role in prostate differentiation,32 but the underlying mechanisms remain elusive. These data suggest that paracrine fatty acid metabolism may drive epithelial differentiation, resulting in decreased glucose metabolism, oxidative stress and lipogenesis (Figure 4).
PPARγ has been shown to regulate the balance between glucose and lipid oxidation in a tissue-specific manner.33 Here, we show that TZD treatment upregulated markers of prostatic differentiation in correlation with an increase in highly metabolic smooth muscle and intramuscular adipose (Figure 5, Table 2), which also correlated with the decreased glucose flux and lipogenesis shown in PPARγ-restored cells in vitro (Figure 2).
The isoform-specific effects of PPARγ have not been directly compared in any tissue. One of the most interesting genes differentially regulated by PPARγ isoforms in prostate was the fatty acid desaturase Scd1. Systemic deletion of Scd1 provides protection against obesity due to reduced fatty acid and triglyceride synthesis as well as increased oxidation.22, 23 In contrast to other tissues, SCD1 expression is high in human prostate epithelia, but its expression and role in prostate cancer has been debated.34, 35 Moreover, PPARγ2 upregulated Scd1 expression and drove basal cell differentiation (Figure 2), and SCD1 was shown to be predominantly restricted to the basal cell compartment of normal human prostate (Figure 6b) whereas PPARγ2 was predominantly expressed in smooth muscle (Figure 6a). As modeled in Figure 6f, these data led us to hypothesize that PPARγ2-mediated import and oxidation of fatty acids may dictate prostate basal cell differentiation through increased SCD1.
Tissue-specific effects of PPARγ agonization have demonstrated that the upregulation of liver fatty acid and sterol synthesis in HFD-fed rats could be reversed by PPARγ agonists, whereas the same HFD stimulated PPARγ downregulation and lipogenesis in muscle.13 Concordant with studies demonstrating positive effects of TZDs on HFD mouse prostates,36, 37 we demonstrated here that chronic HFD treatment resulted in decreased androgen signaling and low-grade PIN, coordinate with decreased PPARγ signaling (Figure 5, Table 2). These data suggest that an allostatic response to downregulate fatty acid import and metabolism may negatively affect prostate differentiation through metabolic switching.
Cells access and metabolize fatty acids through the activities of lipases (e.g., Lpl, Lipa), transporters (e.g., Fabp4, Cd36) and enzymes (acyl-coa synthetases, thioesterases), which supply the cell with acetyl coenzyme A (acetyl-CoA) for eventual entry into mitochondria for energy production. Alternatively, acetyl-CoA can be used as a building block for MUFA production (Scd1) and subsequently converted into triglycerides, cholesterol esters and phospholipids (Figure 6e). The therapeutic efficacy of targeting fatty acid metabolism (synthesis, modification, transport and oxidation) has had some success, but off-target effects have limited their broad usage.10 Similarly, more selective drugs targeting PPARγ and Scd1 promising fewer side effects are being pursued.22, 38 Future studies must be able to link changes in systemic metabolism to local metabolic changes in prostate, which mandates a deeper understanding of the fundamental metabolic infrastructure regulating prostatic differentiation and what allostatic changes may occur in response to systemic metabolic stress. The data presented here suggest that a microenvironment of PPARγ2-mediated fatty acid metabolism by stroma or adipose may drive prostatic epithelial differentiation; however, under conditions of diabetes and obesity fatty acid supply may become saturated, leading to inflammation and hyperplasia in prostate disease (Figure 6f).
Materials and Methods
Generation of cell lines, microarray studies and qRT-PCR
mPrE-PPARγKO cells were spontaneously immortalized from an adult PBCre4tg/0/PPARγflox/flox double-transgenic male mouse.17 The pQCXIP-empty vector, mouse PPARγ1 or PPARγ2 wild-type full-length cDNA (gifts from Drs. Y Eugene Chen and Jifeng Zhang, University of Michigan Medical Center) were stably transfected into the mPrE-PPARγγKO cells to generate mPrE-PPARγγKO+EV, mPrE-PPARγKO+PPARγ1 or mPrE-PPARγKO+PPARγ2 cell line, respectively.17
Microarray and network analyses and qRT-PCR validation
RNA was isolated with Trizol (Ambion, Austin, TX, USA) and reverse transcribed by RNeasy columns (Qiagen, Valencia, CA, USA) followed by reverse transcription (Qiagen) for hybridization on mouse Genechip St. 1.0 microarrays using triplicate samples from each cell line treated with 5 μM Rosiglitazone (Rosi) (Cayman Chemical, Ann Arbor, MI, USA) as well as mPrE-PPARγKO+EV without Rosi. PPARγ-independent effects of Rosi generated in mPrE-PPARγKO+EV cells were subtracted from results comparing mPrE-PPARγKO+EV versus PPARγ-restored cells. SAM was used to stringently select (FDR<0.05) statistically significant genes. Both SAM and unsupervised hierarchical clustering analysis were carried out using TIGR MeV program. Their possible networks and canonic pathways were identified using INGENUITY software (https://apps.ingenuity.com). Custom qRT-PCR plates manufactured by SABiosciences (Valencia, CA, USA) were designed to analyze selected PPARγ-regulated genes in Rosi-treated mPrE-PPARγKO+EV, mPrE-PPARγKO+PPARγ1 and mPrE-PPARγKO+PPARγ2 cell lines in triplicate. Results were analyzed using SABiosciences qRT-PCR analysis software (SABiosciences.com) and represented fold changes based on ΔΔCt analysis.
Detection of ROS
Each cell line was grown to confluence, trypsinized, washed and incubated with 1 μM Dihydroethidium (Life Technologies, Carslbad, CA, USA) for 15 min followed by three washes in PBS. Flow cytometry was then performed and results represent the average mean intensity (N=3). Statistical analysis was performed by GraphPad Prism software (La Jolla, CA, USA) using Student’s unpaired t test.
Fatty acid profile (TLC/MS)
Lipid class separation and fatty acid identification of cell lines were performed by the Vanderbilt Hormone Assay and Analytical Services Core (http://hormone.mc.vanderbilt.edu/). Briefly, the Folch method of lipid extraction39 was followed by thin layer chromatography (TLC) was used to isolate triglycerides. Fatty acid analysis was performed by mass spectrometry (MS) with internal standards. Total triglycerides were normalized to protein concentration as determined by the Lowry method. Statistical analysis was performed on triplicate samples by GraphPad Prism software using one way ANOVA test.
Luciferase assay
Cells were grown to 70% confluency in 12-well culture plates and cotransfected with pRL-null (0.16 μg/well) and ARR2PB-Luciferase (1.44 μg/well) using 4 μl Lipofectamine 2000 in OptiMEM (Invitrogen, Grand Island, NY, USA). After 24 h transfection, media was replaced with 10% charcoal-stripped FBS in DMEM plus or minus 10−8 M DHT and incubated for 24 h. At this time, cells were lysed and dual luciferase activity measured using the Dual Luciferase Reporter Assay System (Promega, Madison, WI, USA) on a Turner Biosystems 20/20n luminometer (Promega). Results were statistically analyzed by Student’s unpaired t test (N=3) using GraphPad Prism.
Western blot
Western blotting was performed as described previously.17 Briefly, 30 μg protein was loaded on 10% SDS acrylamide gels (Life Technologies) and transferred onto PVDF membranes (Millipore, Temecula, CA, USA). The following antibodies were used for detection: mTOR, phospho-mTOR, PTEN, ACC and AKT, phospho-AKT (S473) were from Cell Signaling (1 : 1000, Danvers, MA, USA), Cox-2 (1 : 500, Millipore), AR and PPARγ (1 : 250, Santa Cruz Biotechnology, Santa Cruz, CA, USA), FASN (1 : 1000, GeneTex, Irvine, CA, USA), Gapdh (1 : 1000, Abcam, Cambridge, MA, USA) and Ck14 (1 : 1000, Vector Labs, Burlingame, CA, USA). Blots were then incubated with secondary antibodies (1 : 1000, anti-mouse, anti-rabbit from GE Healthcare, Buckinghamshire, UK) for 45 min in 5% milk in TBST, washed and developed with Western Lightning Plus-ECL (Perkin Elmer, Waltham, MA, USA).
ICC/IHC
ICC/IHC was performed as described previously.17 Briefly, cells were cultured on glass chamber slides (LabTek II, Naperville, IL, USA), washed with PBS, fixed in 4% PFA for 15 min, blocked in 12% BSA for 1 h and incubated with primary antibodies (1 : 25 PPARγ, Santa Cruz; 1 : 50 CK14, Vector Labs; 1 : 100 AR, Santa Cruz; 1 : 500 Ck8/18, Fitzgerald, Acton, MA, USA) in 12% BSA at 4°C overnight followed by incubation with a fluorescent secondary antibody (1 : 1000, Molecular Probes, Eugene, OR, USA) for 45 min at 37°C. Cells were counterstained and mounted with Dapi Vectashield (Vector Labs) and images were taken on a Zeiss Axioplan microscope. Tissues harvested from grafted (kidney capsule) or TZD- or HFD-treated mice (epididymal WAT, prostate) were fixed in 10% formalin at 4°C overnight, paraffin embedded and then 5 μM sections cut for immunohistochemistry. Slides were deparaffinized, rehydrated and endogenous peroxidases blocked with 2% H2O2 in methanol. Following citrate retrieval and blocking with 5% goat serum, slides were incubated at 4°C overnight with the following antibodies: AR (1 : 200, Santa Cruz), PPARγ2 (1 : 200, Abcam), smooth muscle actin (1 : 2000 Invitrogen), Ck14 (1 : 100), CK18 (1 : 500), PDK4 (1 : 500, ProteinTech, Chicago, IL, USA). Secondary antibodies for IHC were from Dako (Carpinteria, CA, USA) and used at 1 : 200 dilution. Secondary antibodies for IF (1 : 1000, Molecular Probes) were incubated for 45 min followed by mounting with DAPI Vectashield.
Tissue recombination
Tissue recombination was performed as described previously.17 Briefly, 400 K of each mPrE-PPARγKO-related cell lines were recombined with 18-day fetal rat UGM and grafted in collagen into the kidney capsule of male SCID mice for 2 months. Grafts were harvested and fixed in formalin for morphologic and immunohistochemical analysis.
Glucose/lactate measurements
Extracellular uptake and excretion rates were determined in triplicate growth experiments. Eight separate 48-well tissue culture plates were seeded at a density of 20 K cells. One plate was sampled every 10–14 h, whereby the conditioned medium was removed and frozen at −80°C. The remaining cells on the plate were stained with crystal violet for assessment of cell number. Concentrations of medium glucose and lactate were determined using a YSI 2300 Stat Plus Glucose and Lactate Analyzer (YSI, Yellow Springs, OH, USA). Cell-specific rates of glucose consumption and lactate production were determined by regression analysis using the method of Glacken et al.40
Animal experiments
To examine the effects of TZD or HFD regimen on prostate gene expression in vivo, control, Rosiglitazone chow (0.188% Avandia) or Western diet chow (16% protein, 40% carbohydrate, 40% fat, 0.15% cholesterol) (Test Diets, Richmond, IN, USA) were fed to male C57B mice for 6 months at which time the animals were weighed (Student’s unpaired t test for statistical analysis) and tissues removed for formalin fixation and storage at −80°C for later RNA extraction. Because of the extreme density of adipose directly surrounding the prostate of both TZD- and HFD-fed animals, care was taken to dissect away as much as possible. WAT was taken from the epididymal fat pad. RNA was extracted and cDNA synthesized for qRT-PCR on custom-designed plates (Qiagen) using an ABI 7900HT real-time PCR machine with a standard block according to the manufacturer’s instructions. Results represent triplicate experiments and fold changes were calculated as described above. P values were mostly insignificant, given the contamination by intramuscular adipocytes and were therefore not shown.
Abbreviations
- acetyl-CoA:
-
acetyl coenzyme A
- AR:
-
androgen receptor
- ARE:
-
androgen response element
- BPH:
-
benign prostatic hyperplasia
- DHT:
-
dihydrotestosterone
- eWAT:
-
epididymal white adipose tissue
- HFD:
-
high-fat diet
- MetS:
-
metabolic syndrome
- mPrE-PPARγ KO (mPrE-γKO):
-
a PPARγ knockout mouse prostate epithelial cell line spontaneously immortalized from an adult PBCre4 tg/0/PPARγ flox/flox mouse prostate
- mPrE-PPARγKO+PPARγ1 (+PPARγ1 or +γ1):
-
mPrE-γKO overexpressing mouse PPARγ1 WT full-length cDNA
- mPrE-PPARγKO+PPARγ2 (+PPARγ2 or +γ2):
-
mPrE-γKO overexpressing mouse PPARγ2 WT full-length cDNA
- mPrE-PPARγKO+empty vector (+EV):
-
mPrE-γKO retrovirally transduced using a control empty vector
- MUFAs:
-
monounsaturated fatty acids
- KO:
-
knockout
- PCa:
-
prostate cancer or prostate carcinoma
- PIN:
-
prostatic intraepithelial neoplasia
- PPARγ:
-
peroxisome proliferator-activated receptor gamma
- ROS:
-
reactive oxygen species
- TCA:
-
tricarboxylic acid
- TZDs:
-
thiazolidinediones
- UGM:
-
urogenital mesenchyme
References
Hammarsten J, Peeker R . Urological aspects of the metabolic syndrome. Nat Rev Urol 2011; 8: 483–494.
Bourke JB, Griffin JP . Diabetes mellitus in patients with benign prostatic hyperplasia. Br Med J 1968; 4: 492–493.
Parsons JK, Carter HB, Partin AW, Windham BG, Metter EJ, Ferrucci L et al. Metabolic factors associated with benign prostatic hyperplasia. J Clin Endocrinol Metab 2006; 91: 2562–2568.
Berger AP, Bartsch G, Deibl M, Alber H, Pachinger O, Fritsche G et al. Atherosclerosis as a risk factor for benign prostatic hyperplasia. BJU Int 2006; 98: 1038–1042.
Burke JP, Rhodes T, Jacobson DJ, McGree ME, Roberts RO, Girman CJ et al. Association of anthropometric measures with the presence and progression of benign prostatic hyperplasia. Am J Epidemiol 2006; 164: 41–46.
Michel MC, Mehlburger L, Schumacher H, Bressel HU, Goepel M . Effect of diabetes on lower urinary tract symptoms in patients with benign prostatic hyperplasia. J Urol 2000; 163: 1725–1729.
Freeman MR, Solomon KR . Cholesterol and benign prostate disease. Diff Res Biol Div 2011; 82: 244–252.
Yeh HC, Platz EA, Wang NY, Visvanathan K, Helzlsouer KJ, Brancati FL . A prospective study of the associations between treated diabetes and cancer outcomes. Diab Care 2012; 35: 113–118.
Grundmark B, Garmo H, Loda M, Busch C, Holmberg L, Zethelius B . The metabolic syndrome and the risk of prostate cancer under competing risks of death from other causes. Cancer Epidemiol Biomarkers Prev 2010; 19: 2088–2096.
Flavin R, Zadra G, Loda M . Metabolic alterations and targeted therapies in prostate cancer. J Pathol 2011; 223: 283–294.
Lee CH, Olson P, Evans RM . Minireview: lipid metabolism, metabolic diseases, and peroxisome proliferator-activated receptors. Endocrinology 2003; 144: 2201–2207.
Cheatham WW . Peroxisome proliferator-activated receptor translational research and clinical experience. Am J Clin Nutr 2010; 91: 262S–266S.
Hsiao G, Chapman J, Ofrecio JM, Wilkes J, Resnik JL, Thapar D et al. Multi-tissue, selective PPARgamma modulation of insulin sensitivity and metabolic pathways in obese rats. Am J Physiol Endocrinol Metab 2011; 300: E164–E174.
He W, Barak Y, Hevener A, Olson P, Liao D, Le J et al. Adipose-specific peroxisome proliferator-activated receptor gamma knockout causes insulin resistance in fat and liver but not in muscle. Proc Natl Acad Sci USA 2003; 100: 15712–15717.
Vidal-Puig A, Jimenez-Linan M, Lowell BB, Hamann A, Hu E, Spiegelman B et al. Regulation of PPAR gamma gene expression by nutrition and obesity in rodents. J Clin Invest 1996; 97: 2553–2561.
Norris AW, Chen L, Fisher SJ, Szanto I, Ristow M, Jozsi AC et al. Muscle-specific PPARgamma-deficient mice develop increased adiposity and insulin resistance but respond to thiazolidinediones. J Clin Invest 2003; 112: 608–618.
Jiang M, Fernandez S, Jerome WG, He Y, Yu X, Cai H et al. Disruption of PPARgamma signaling results in mouse prostatic intraepithelial neoplasia involving active autophagy. Cell Death Differ 2010; 17: 469–481.
Jiang M, Jerome WG, Hayward SW . Autophagy in nuclear receptor PPARgamma-deficient mouse prostatic carcinogenesis. Autophagy 2010; 6: 175–176.
Jiang M, Strand DW, Franco OE, Clark PE, Hayward SW . PPARgamma: A molecular link between systemic metabolic disease and benign prostate hyperplasia. Differentiation 2011; 82: 220–236.
Fajas L, Auboeuf D, Raspe E, Schoonjans K, Lefebvre AM, Saladin R et al. The organization, promoter analysis, and expression of the human PPARgamma gene. J Biol Chem 1997; 272: 18779–18789.
Clarke SD . Polyunsaturated fatty acid regulation of gene transcription: a molecular mechanism to improve the metabolic syndrome. J Nutr 2001; 131: 1129–1132.
Sampath H, Ntambi JM . The role of stearoyl-CoA desaturase in obesity, insulin resistance, and inflammation. Ann N Y Acad Sci 2011; 1243: 47–53.
Miyazaki M, Sampath H, Liu X, Flowers MT, Chu K, Dobrzyn A et al. Stearoyl-CoA desaturase-1 deficiency attenuates obesity and insulin resistance in leptin-resistant obese mice. Biochem Biophys Res Commun 2009; 380: 818–822.
Kelley DE . Skeletal muscle fat oxidation: timing and flexibility are everything. J Clin Invest 2005; 115: 1699–1702.
Sears DD, Hsiao G, Hsiao A, Yu JG, Courtney CH, Ofrecio JM et al. Mechanisms of human insulin resistance and thiazolidinedione-mediated insulin sensitization. Proc Natl Acad Sci USA 2009; 106: 18745–18750.
Medina-Gomez G, Gray SL, Yetukuri L, Shimomura K, Virtue S, Campbell M et al. PPAR gamma 2 prevents lipotoxicity by controlling adipose tissue expandability and peripheral lipid metabolism. PLoS Genet 2007; 3: e64.
Walden TB, Hansen IR, Timmons JA, Cannon B, Nedergaard J . Recruited vs. nonrecruited molecular signatures of brown, "brite," and white adipose tissues. Am J Physiol Endocrinol Metab 2012; 302: E19–E31.
Kelly LJ, Vicario PP, Thompson GM, Candelore MR, Doebber TW, Ventre J et al. Peroxisome proliferator-activated receptors gamma and alpha mediate in vivo regulation of uncoupling protein (UCP-1, UCP-2, UCP-3) gene expression. Endocrinology 1998; 139: 4920–4927.
Danaei G, Finucane MM, Lu Y, Singh GM, Cowan MJ, Paciorek CJ et al. National, regional, and global trends in fasting plasma glucose and diabetes prevalence since 1980: systematic analysis of health examination surveys and epidemiological studies with 370 country-years and 2.7 million participants. Lancet 2011; 378 (9785): 31–40.
Chavez JA, Summers SA . Lipid oversupply, selective insulin resistance, and lipotoxicity: molecular mechanisms. Biochim Biophys Acta. 2010; 1801: 252–265.
De Nunzio C, Aronson W, Freedland SJ, Giovannucci E, Parsons JK . The correlation between metabolic syndrome and prostatic diseases. Eur Urol 2012; 61: 560–570.
Strand DW, Franco OE, Basanta D, Anderson AR, Hayward SW . Perspectives on tissue interactions in development and disease. Curr Mol Med 2010; 10: 95–112.
Way JM, Harrington WW, Brown KK, Gottschalk WK, Sundseth SS, Mansfield TA et al. Comprehensive messenger ribonucleic acid profiling reveals that peroxisome proliferator-activated receptor gamma activation has coordinate effects on gene expression in multiple insulin-sensitive tissues. Endocrinology 2001; 142: 1269–1277.
Moore S, Knudsen B, True LD, Hawley S, Etzioni R, Wade C et al. Loss of stearoyl-CoA desaturase expression is a frequent event in prostate carcinoma. Int J cancer 2005; 114: 563–571.
Fritz V, Benfodda Z, Rodier G, Henriquet C, Iborra F, Avances C et al. Abrogation of de novo lipogenesis by stearoyl-CoA desaturase 1 inhibition interferes with oncogenic signaling and blocks prostate cancer progression in mice. Mol Cancer Ther 2010; 9: 1740–1754.
Vikram A, Jena G, Ramarao P . Pioglitazone attenuates prostatic enlargement in diet-induced insulin-resistant rats by altering lipid distribution and hyperinsulinaemia. Br J Pharmacol 2010; 161: 1708–1721.
Vikram A, Jena GB, Ramarao P . Increased cell proliferation and contractility of prostate in insulin resistant rats: linking hyperinsulinemia with benign prostate hyperplasia. Prostate 2010; 70: 79–89.
Rikimaru K, Wakabayashi T, Abe H, Imoto H, Maekawa T, Ujikawa O et al. A new class of non-thiazolidinedione, non-carboxylic-acid-based higshly selective peroxisome proliferator-activated receptor (PPAR) gamma agonists: design and synthesis of benzylpyrazole acylsulfonamides. Bioorg Med Chem 2012; 20: 714–733.
Folch J, Lees M, Sloane Stanley GH . A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem 1957; 226: 497–509.
Glacken MW, Adema E, Sinskey AJ . Mathematical descriptions of hybridoma culture kinetics: I. Initial metabolic rates. Biotechnol Bioeng 1988; 32: 491–506.
Acknowledgements
All microarray experiments were performed in the Vanderbilt Genome Sciences Resource. The Vanderbilt Genome Sciences Resource is supported by the Vanderbilt Ingram Cancer Center (P30 CA68485), the Vanderbilt Digestive Disease Center (P30 DK58404) and the Vanderbilt Vision Center (P30 EY08126). This work was supported by a Department of Defense Prostate Cancer Training Award W81XWH-07-1-0479 to DWS, P20 DK090874 and 2R01 DK067049 to SWH, and R21 CA155964 to JDY. All fatty acid analyses were performed by the Vanderbilt Hormone Assay and Analytical Services Core. We thank Drs. Robert Matusik and David Degraff for helpful discussions and manuscript editing.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by C Munoz-Pinedo
Supplementary Information accompanies the paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Strand, D., Jiang, M., Murphy, T. et al. PPARγ isoforms differentially regulate metabolic networks to mediate mouse prostatic epithelial differentiation. Cell Death Dis 3, e361 (2012). https://doi.org/10.1038/cddis.2012.99
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2012.99
Keywords
This article is cited by
-
The role of PPARγ in prostate cancer development and progression
British Journal of Cancer (2023)
-
Transcriptional networks controlling stromal cell differentiation
Nature Reviews Molecular Cell Biology (2021)
-
Competitive glucose metabolism as a target to boost bladder cancer immunotherapy
Nature Reviews Urology (2020)
-
Exploiting vulnerabilities of cancer by targeting nuclear receptors of stromal cells in tumor microenvironment
Molecular Cancer (2019)
-
Oxidative stress in prostate hyperplasia and carcinogenesis
Journal of Experimental & Clinical Cancer Research (2016)