Abstract
Caspase-dependent apoptotic cell death has been extensively studied in cultured cells and during embryonic development, but the existence of analogous molecular pathways in single-cell species is uncertain. This has reduced enthusiasm for applying the advanced genetic tools available for yeast to study cell death regulation. However, partial characterization in mammals of additional genetically encoded cell death mechanisms, which lead to a range of dying cell morphologies and necrosis, suggests potential applications for yeast genetics. In this light, we revisited the topic of gene-dependent cell death in yeast to determine the prevalence of yeast genes with the capacity to contribute to cell-autonomous death. We developed a rigorous strategy by allowing sufficient time for gene-dependent events to occur, but insufficient time to evolve new populations, and applied this strategy to the Saccharomyces cerevisiae gene knockout collection. Unlike sudden heat shock, a ramped heat stimulus delivered over several minutes with a thermocycler, coupled with assessment of viability by automated counting of microscopic colonies revealed highly reproducible gene-specific survival phenotypes, which typically persist under alternative conditions. Unexpectedly, we identified over 800 yeast knockout strains that exhibit significantly increased survival following insult, implying that these genes can contribute to cell death. Although these death mechanisms are yet uncharacterized, this study facilitates further exploration.
Similar content being viewed by others
Main
Studies in yeast models have provided key insight into normal cell physiology and human disease processes including neurological disorders.1 In contrast, yeast model systems have not yet advanced our understanding of mammalian cell death mechanisms. Disputes over the existence of apoptosis-like mechanisms in yeast have further thwarted efforts in this direction. However, an emerging list of alternative non-apoptotic cell death mechanisms in mammalian cells helps to allay these concerns.2 Because less is known about these alternate mammalian mechanisms and the responsible genes, it is difficult to predict which, if any, of these processes will be evolutionarily conserved in yeast. Several lines of evidence support the existence of genetically encoded cell death functions in single-cell species,3, 4, 5, 6 but even less is known about these molecular mechanisms.
Before pursuing a strategy to uncover potential cell death genes in yeast, it is useful to articulate a generally acceptable definition of a cell death gene that applies to both single- and multi-cellular species. In this study we will adopt the term programmed cell death in the broadest sense, referring to any genetically encoded function of the dying cell that contributes importantly to its own demise. This definition can be justified by the following arguments. Conceptually, the genes responsible for cell death can be grouped into two main categories, those selected during evolution to cause cell suicide and those that did not evolve for this purpose (accidental cell death). The first category includes genes that delete selected subsets of plant and animal cells required to construct complex networks during development, to maintain healthy tissues in maturity and to respond to infectious pathogens and environmental stresses.7, 8 These cell death genes must not be constitutively active, but are selectively activated by specific signals to promote cell autonomous death. The severe structural malformations of the head and brain of knockout mice lacking caspase-9 or caspase-3 provide experimental evidence that these caspases evolved at least in part to eliminate the vast excess number of neurons produced during development.9 New evidence strongly argues that at least some necrotic cell death mechanisms were also selected during evolution, for example, in the hematopoietic system.10 It could also be argued that evolved cell death genes include those genes that promote the death of one cell to sustain the survival of another cell by providing nutrients derived from the dead cell.11, 12
The second category of cell death genes includes those that hasten cell death accidentally and, therefore by definition, did not evolve for this purpose. Originally it was assumed that essentially all forms of accidental cell death were non-programmed (not gene-dependent) and were described in morphological terms as necrosis. However, an important distinction was overlooked. Indeed it is unlikely that gene products encoded by the dying cell will have any influence on cell fate if the assault is sufficiently strong to kill by sheer force; this is gene-independent (non-programmed) cell death. In contrast, examples of gene-dependent accidental cell death may include the detrimental programmed necrotic cell deaths observed with non-apoptotic ischemic injury, lysosome leakage and the exaggerated inflammatory responses characteristic of virus-induced encephalitis.13, 14
Although evolved and accidental cell death genes can be distinguished conceptually, they are more difficult to distinguish experimentally except where developmental processes (e.g., embryogenesis and lymphoid system) are available for study. For example, it is not currently possible to rigorously determine if the pathological cell death during ischemic injury in adults is evolved, accidental or a combination thereof, because knockout animals could be expected to have the same phenotype in any case. This effort is further complicated if the pro-death gene also has essential pro-survival functions (e.g., calpain). Regardless of whether the pro-death functions of cathepsins, calpain and RIPK are the result of evolution or can occur accidentally, their importance as therapeutic targets is not dependent on this distinction. Discerning evolved versus accidental cell death of protists is equally problematic, and both categories of cell death genes may be informative about human disease mechanisms. The genes responsible for an array of necrosis-like morphologies are currently being identified and may outnumber the list of genes that facilitate caspase-mediated apoptosis. Therefore, we sought to determine the prevalence of death-promoting genes in yeast, regardless of their evolutionary origins and regardless of the cell morphologies that occur during the dying process. However, because we found that the available yeast cell death assays were not amenable to this task, we first had to develop a strategy to reliably quantify gene-dependent cell death of yeast.
Results
Experimental definition of gene-dependent yeast cell death
While performing the hundreds of trials required for developing treatment conditions that reliably detect gene-dependent yeast cell death, we routinely compared three test strains with distinct susceptibilities to cell death (wild type, Δdnm1 and Δfis1*). The dynamin-related GTPase Drp1/Dnm1 is a conserved mitochondrial fission factor that can promote cell death in mammals,15 flies,8 worms16 and yeast,3 though the detailed mechanisms are not delineated. Deletion of the yeast DNM1 gene renders yeast resistant to a range of insults, including H2O2, acetic acid, viral toxins and aging.3, 17, 18, 19, 20 DNM1 deletion also protects yeast from heat treatment in a dose-dependent manner measured by colony formation (Figures 1a and b). To mark the lower bound of survival during assay development, we used the death-sensitive FIS1 knockout strain.3 Because Fis1 is also a conserved mitochondrial fission factor, it was not anticipated that FIS1 deletion would cause sensitivity to cell death stimuli. This paradox is explained by a secondary WHI2 gene mutation that arose independently in several FIS1 knockouts (Δfis1*).21 Spontaneous WHI2 mutations apparently compensate for a fitness defect unrelated to mitochondrial fission caused by deletion of FIS1.21
Genetic definition of programmed cell death for assay development. (a) Log-phase wild type (BY4741) and yeast MATa deletion strains were analyzed using a heat ramp assay (30–51 °C over 30 min and held at 51 °C for the indicated times). Undiluted and fivefold serial dilutions were plated on YPD agar; Δfis1* (Δfis1/whi2-d1). (b) Cell viabilities for experiments in panel (a) were also determined by vital dye staining with FUN1 (Molecular Probes), presented as mean±S.D. for three independent experiments each in duplicate, counting double blinded samples, ∼300 cells per sample. P<0.00002 (t-test) for wild type versus each mutant under all conditions except untreated
Importantly, the cell death phenotypes of knockout strains Δdnm1 and Δfis1* in this assay were detectable only after an insult, as these strains were indistinguishable from wild type under our conditions before treatment (Figure 1a). Vital dye staining yields similar results, demonstrating that reduced colony formation following treatment is not caused by failure of living cells to grow into colonies (Figure 1b).
Gene-specific cell death phenotypes revealed by heat ramp treatment
Using these test strains, we sought to establish a cell death assay that invariably distinguishes the survival of these test strains from wild type under any given condition. Key concepts for success were to establish treatment conditions that are sufficiently mild to allow time for gene-dependent cell suicide to occur, and to deliver this stimulus uniformly and simultaneously to all cells in the sample. It is well recognized in mammalian systems that excessive doses of an otherwise apoptotic stimulus (e.g., heat) can kill cells by non-programmed assault that cannot be influenced by any genetically encoded intracellular factor.22 This phenomenon is illustrated for yeast cultures (mid log-phase) that are quickly heated within seconds using a thermocycler. Although these conditions can occasionally reveal differences in survival between wild type and the test strains, including the death-resistant metacaspase knockout Δyca1/Δmca1 (Figure 2a), small variations in temperature, treatment duration or growth conditions before treatment, abolish these differences (Figure 2b and data not shown). This problem is overcome by using a thermocycler (PCR machine) programmed to ramp from ambient to 51 °C over 12–30 min, rather than seconds. Essentially without fail, these conditions simultaneously distinguish the survival of test strains (Figures 2c and d). In contrast, ramping the temperature over the course of 1 h allows sufficient time for test strains to adapt, again concealing genetic differences in cell death susceptibility (Figure 2e) as previously described.23
Thermocycler conditions to induce gene-dependent cell death. (a and b) Log-phase yeast cultures were untreated or treated with heat ramp conditions as graphed, ramping from 30–51/52 °C in ∼6–20 s, holding at temperature for 5 min before plating on YPD. (c) In 2 min 30–40 °C, then from 40–51 °C in 10 min and held at 51 °C for 10 min. (d) In 30 min 30–51 °C and held at 51 °C for 30 min. (e) In 60 min 30–51 °C and held at 51 °C for 30 min
Using this general approach, several parameters (e.g. ramp rate, maximum temperature and hold time) can be adjusted with relative latitude to accommodate changes in metabolic state of the cells, media types, background strains and strains bearing exogenous plasmids, each with high reproducibility. For example, while post-log-phase yeast cultures are known to be more resistant to death than log-phase cells treated with the same heat ramp conditions (Figure 2c versus Figure 3a-left), adjustments in assay conditions easily accommodate both 16 h and 48 h post-log cultures (Figure 3a). Even a single condition can distinguish test strains that become increasingly sensitive to death with increasing time spent in continuous log phase before treatment (Figure 3b).
Heat ramp assays accommodate different metabolic states. (a) Cells from dense 16 and 48 h cultures were diluted to OD600 0.5 in YPD and immediately treated with the indicated heat ramp conditions before plating on YPD. (b) Cultures grown in continuous log phase for 3, 4 or 7 h (starting at OD600 of 0.25, 0.20 and 0.05, respectively) were treated (all at OD600 0.5) as shown in panel a (left) and colony counts are presented as mean±S.D. colony-forming units (CFU) for at least four independent experiments per condition. t-test, P<0.00001 comparing WT to all others at 3–4 h, P<0.04 at 7 h where counts are low (not significant for DNM1 when corrected for multiple comparisons). Untreated samples were indistinguishable between strains (not shown)
Importantly, a thermocycler eliminates confounding edge effects encountered when multi-well plates or large tubes are heated in other devices (e.g. air or liquid). This strategy also overcomes any problems with delivery of liquid death stimuli (e.g. acetic acid, H2O2) that may not reach all cells equally during mixing or that are inactivated rapidly upon aqueous dilution. Importantly, survival differences between strains prevail under a range of heat ramp conditions and are highly reproducible, facilitating presentation of statistical analyses and avoiding the need to report representative experiments.
Confirmation of established phenotypes
Although the existence of evolved (or accidental) cell death genes in yeast is currently debated, yeast do encode many orthologues of genes with established roles in metazoan cell death, some of which have been implicated in promoting death in yeast. Therefore, to further validate our heat ramp assay, we analyzed the eight knockout strains identified in the early yeast apoptosis literature. The corresponding deletion strains from the MATa YKO collection (BY4741) were tested in both log and post-log phase. All strains exhibited phenotypes that matched published results where various cell death stimuli had been employed (Figure 4a). Two strains are more prone to cell death, the knockout of SOK2, a transcription factor that regulates production of ammonia required for cell-cell signaling and long-term survival in colonies of wild strains,5 and the knockout of NUC1, encoding a conserved mitochondrial nuclease (EndoG).24 The remaining six knockout strains exhibited resistance to cell death suggesting pro-death functions for these genes: Nma111 (nuclear mediator of apoptosis), orthologue of four mammalian mitochondrial serine proteases including Htra2/Omi;25 Aif1 (apoptosis-inducing factor), an NADH oxidoreductase named after its mammalian orthologue AIF;26 the Aif1-related yeast protein Ndi1, an NADH dehydrogenase;27 Mmi1, the highly conserved homolog of human TCTP (translationally controlled tumor protein);28 Tat-D, a pro-death DNase in yeast, worms and mammals;29 and Stm1, which lacks an obvious mammalian orthologue but is reported to induce yeast cell death.30
Confirmation of yeast cell death phenotypes. (a) Log-phase and 16-h post-diauxic phase cultures were treated with the heat ramp graphed in Figure 3a left and middle, respectively. Log-phase cultures were prepared from the same 16-h cultures. Cell viability was determined by colony-forming units (CFU) counted with the BioSpot instrument and data are presented as mean±S.D. of 3–6 independent experiments. All data generated are graphed on both linear and log scales to reveal all differences. t-test, comparing wild type to all others, P<0.00001 log phase, P<0.0006 post log. Untreated samples were indistinguishable between strains (not shown). (b) Results for the same strains are presented as percentile rank survival based on full genomic screen (Supplementary Table S1)
Many gene deletion strains are resistant to cell death
Using plate-1 of the MATa deletion collection, which contains the fission factor Dnm1/Drp1 knockout among 94 knockout strains from chromosomes 1 and 12, we adapted the heat ramp assay for high throughput screening (see Figure 6a). Colony formation is a standard measure of yeast survival, equivalent to mammalian clonogenic assays. Surprisingly, there was no commercially available laboratory equipment capable of accurately enumerating yeast colonies in high throughput formats. Instruments that measure only the density of yeast growth on solid medium or in liquid cultures do not provide the desired information. Only colony number regardless of colony size provides a measure of cell survival that is not confounded by cell growth rates (e.g., 100 tiny colonies derived from 100 live cells have far less mass than a single large colony). Unless viability is measured with vital dyes (confounded by the time delay required to sequentially sample 96 wells by flow cytometry), it is necessary to allow surviving cells to outgrow dead cell numbers following treatment.
To help overcome these problems, we performed automated colony counting of 2-day plates using an EliSpot Reader (Cellular Technology Limited/CTL, Shaker Heights, OH, USA) designed to read immunological ELISA assays, which proved more accurate than hand counting (Figure 5a and Supplementary Figure S1). To minimize differences in starting cell numbers between strains, we used saturated cultures (48 h). Cell densities measured before treatment had no systematic correlation with colony counts following heat ramp, suggesting that small differences in cell density are not a confounding factor (Figure 5b). Despite using media and other consumables from the same suppliers and lot numbers, there are inevitable day-to-day differences in the total number of colonies combining all strains on plate-1, although replicate plates generated on the same day are highly similar (Figure 5c). Despite this issue, the relative survival of individual knockout strains was highly reproducible on different days (Figure 5d and Supplementary Figure S2).
Variables affecting colony counts in heat ramp assays. (a) Following heat ramp treatment of YKO strains on MATa plate-1 from the MATa YKO collection, the 96 samples were plated in 96 spots and incubated at 30 °C for 2 days for colony visualization. Colony counts were determined for the same plate using two methods, hand counting and automated counting using an EliSpot Reader (CTL, Inc.), expected correlation (dashed line); hand method systematically undercounts in spots with >20 colonies. (b) Absorbance (595 nm) of each of four replicates for all 96 wells/samples in plate-1 of the MATa YKO collection after 48 h incubation (from panel c) are plotted against corresponding colony counts determined with a BioSpot instrument after heat ramp treatment using the assay in panel a. Correlation coefficient is not significant (0.238). Arrow marks results for wells A1 and H2 lacking yeast. (c) Colony counts for all 94 knockout strains in plate-1 were determined on four different days, each starting with an independently pinned plate prepared from the same original frozen stock. After heat ramp treatment, samples were plated on three or four replicate YPD plates, incubated ∼18 h at 30 °C and counted with a prototype BioSpot instrument. Median colony counts combining all strains (heavy line) on each replicate plate are shown with 25 and 75% quartiles (boxes), and the range of 95% counts (dash lines). (d) Rank order survival of individual deletion strains on plate-1 (mean±S.E. of data shown in panel c); shaded quantiles; for gene names, see Supplementary Figure S2
High throughput enumeration of single and merged colonies ranging widely in size was ultimately achieved by developing the new BioSpot platform (Cellular Technology Limited/CTL) with automated object recognition capability and a specialized image capture and analysis system. This system is capable of accurately counting at least 300 microscopic colonies per spot at only 18 h after plating (Figure 6a), overcoming the 2-day growth period required to generate visible colonies, which exacerbates the problem of overlapping colonies, reduces the countable range of discernable colonies per spot (≤30) and biases counts downward.
Scheme for high throughput heat ramp cell death assay. (a) See Materials and Methods for details. Bright-field image of one 18 h spotted sample from the BioSpot instrument is shown. (b) Viability measured as colony number for 4–8 replicates of all strains in the MATa YKO collection are plotted (each point representing the mean for 10 knockout strains. The results for control strains HO (mean of 108) and Δdnm1 (mean of 202) are indicated (Supplementary Table S1). Mean count for all strains=122; the quantile for wild-type and HO knockout is boxed, with 2000 YKOs surviving better than WT
The heat ramp cell death assay (Figure 6a, heat ramp conditions from Figure 3a right) was applied to the MATa yeast knockout collection of ∼4800 single gene knockout strains, and the results of 4–8 replicates were counted using the BioSpot platform. Because the wild-type parental strain is not present in the YKO collection, wild-type strains from different sources were processed equivalently in parallel or inserted into an empty well on a subset of plates for direct comparisons. In addition, the heat ramp survival result for wild type was estimated using the HO gene (YDL227C) knockout strain in the YKO collection (average of 108 colonies), which was a non-functional gene at the time of deletion, yielding results similar to wild-type control strains (Figure 6b). Unexpectedly, ∼800 different knockout strains in the YKO collection were more resistant to heat ramp-induced death compared with the death-resistant strain Δdnm1, implying that at least one function of each of these deleted genes contributes in some way to the death of the cell upon receiving a death stimulus. The output data from the genome-wide cell death screen were curated by further optimizing the automated counting parameters, and by removing any miscounted replicates and other artifacts identified upon inspection. After the curated genomic screen data were assembled and the ∼4800 knockout strains were ordered by survival rank (Supplementary Table S1), we then retrieved the results from this screen for the original test strains and for the eight strains with previously reported cell death phenotypes. These results from the high throughput screen (Figure 4b) are strikingly similar to the hand screen (Figure 4a), further validating the high throughput heat ramp assay.
Shared death phenotypes despite different metabolic states and death stimuli
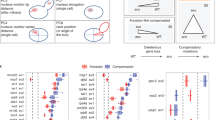
Dnm1/Drp1 has a conserved pro-death activity following multiple death stimuli and across several eukaryotic species.3, 8, 15, 16, 31 Therefore, we predicted that many other cell death genes will behave similarly regardless of the death stimulus type or metabolic conditions imposed. To test this possibility, plate-1 of the MATa knockout collect was treated in log phase with either the heat ramp assay (as in Figure 2c) or with a range of acetic acid concentrations and compared with the heat ramp results in Figure 6 for the same strains in 48-h post-log phase (Figure 7). Most strains (71%) exhibit a similar phenotype under all three conditions, 11% exhibited a distinct phenotype under one of the three conditions, while 18% exhibited distinct phenotypes under all three conditions. Interestingly, heat ramp sensitivity did not correlate well with reported heat shock screens on the SGD database (red dots). Thus, the heat ramp cell death assays appear to detect novel phenotypes that can be further delineated using yeast model systems.
High throughput cell death assays reveal novel phenotypes. Yeast knockout strains on plate-1 MATa YKO collection were assessed for survival by colony formation following a death stimulus. Gene names are ordered by mean survival rank in three different assay types. Results are shown for acetic acid treatment40 in log phase (AA-log, three independent experiments each with 140 mM, 145 mM and 150 mM acetic acid as previously described), heat ramp treatment graphed in Figure 2c in log phase (HR-log, four independent experiments), and for heat ramp treatment graphed in Figure 3a right for 48-h post-diauxic phase (HR-post, four independent experiments)
Discussion
Contrary to expectation, we found that ∼2000 yeast knockout strains are more resistant to cell death than wild type, and ∼800 different gene knockouts (representing ∼15% of the genes in the genome) are more resistant to death than the DNM1 (DRP1) knockout. The implication is that the activities of a large number of gene products are detrimental to cell survival following a stress stimulus. These gene functions could inadvertently contribute to cell death when inappropriately deregulated during stress, analogous to damage caused by a dislodged mechanical machine part. Alternatively, if unchecked cell survival is detrimental to the long-term survival of a given species, then these genes could have evolved in part to serve as checks and balances built into most cellular processes to avoid inappropriate cell survival. Many conserved cellular processes, including mitochondrial dynamics, bioenergetics, autophagy and the beneficial effects of calorie restriction have been linked to the control of programmed cell death in mammals and other model organisms and are well studied in yeast. Thus, yeast genetic tools represent a powerful yet largely untapped resource poised for exploitation of cell death mechanisms. The stringent assays reported in this study increase the feasibility of exploring yet unknown cell death pathways in yeast.
Identification of genes that inhibit cell death is more complicated because deletion of many important genes individually could be expected to impair cell survival during stress. These genes simply could be required for normal cellular functions, rather than encoding biochemical anti-death activities analogous to mammalian Bcl-2 or any other active inhibitors of known cell death mechanisms.32 Thus, we refer to these yeast genes as pro-survival (rather than anti-death) until they are proven to actively inhibit a cell death program. A key component of our assay is that survival assessment is irrespective of growth rate as long as they grew into a colony of ∼100 microns after plating.
Despite ongoing debate, evolutionarily selected death pathways have been purported to exist in single-cell species including yeast, parasites and bacteria. For example, dying cells near the center of a colony of wild yeast strains appear to be required for nourishment and survival of younger cells.5 Yeast viruses can trigger gene-dependent death, a phenomenon assumed to be a host defense mechanism in mammals unless the plan goes awry and excessive cell death causes disease pathology.17 Experimental models of evolution suggest that ultimate survival of a species relies on programmed death of a portion of the population in aging yeast cultures as nutrients become diminished.33 In this scenario, long-term survival of all cellular species could require programmed cell death to eliminate infected, damaged or defective cells, to match population sizes with resources, and to facilitate survival of better-adapted genomes to arise in the population. Yet little is known genetically or molecularly about these death processes in single-cell species. Yeast and plants encode metacaspases that are distantly related to animal caspases.7, 34 However, there is contention about whether the yeast metacaspase is involved in a process resembling mammalian apoptotic self-destruction and subsequent engulfment by healthy mammalian cells.35, 36 Nevertheless, several labs have found that deletion of the yeast metacaspase gene YCA1/MCA1 can protect yeast from death after specific insults.31, 34 Thus, it is reasonably clear that YCA1/MCA1 can promote yeast cell death, although the detailed mechanisms by which any yeast factor kills cell autonomously are essentially unknown.
A major obstacle to studying gene-dependent cell death in yeast is the limited array of assays suitable for evaluating gene-dependent cell death. Thus, the study of programmed cell death in yeast is arguably in a catch-up mode compared with other well-established model organisms.8 The application of mammalian apoptosis assays to yeast has been a standard in the field. However, there are controversies regarding these assays because it is unclear if the biochemical events detected by apoptosis assays directly reflect yeast cell death mechanisms.5, 37, 38 For example, DNA fragmentation detected in dying yeast by the TUNEL assay (labels DNA ends) could be interpreted at least three different ways. TUNEL reactivity could detect an evolutionarily selected pro-death nuclease (analogous to mammalian CAD), or could detect an unintentionally (accidentally) activated nuclease that significantly contributes to the death. More prosaically, yeast TUNEL reactivity could simply reflect unleashed nucleases that have no part in the death process (i.e., cells die at the same rate with or without the nuclease directly responsible for TUNEL reactivity). A similar argument holds for Annexin V staining of surface-exposed phosphatidylserine, an ‘eat me’ signal for phagocytosis of dying cells in metazoans.39 Detection of caspase-like activity and reactive oxygen species with fluorescent reporters in dying yeast has also been challenged.35, 37 Presently, it is prudent to avoid conclusions about underlying yeast cell death mechanisms based on mammalian apoptosis assays. Apoptosis assays applied to yeast could still be useful as a live-dead assay analogous to Trypan blue staining of dead mammalian cells. However, in our hands these assays can introduce new variables (e.g., cell wall removal easily compromises the plasma membrane, inadvertently labeling the inner leaflet of the plasma membrane with annexin V). Therefore, improved tools are urgently needed to take advantage of the unsurpassed tools available for yeast.
The possible existence of multiple phylogenetically conserved death pathways and of genes that accidentally contribute to cell death further increase the prospects of devising interventions against necrotic cell death previously not considered druggable. These discoveries coupled with the opportunity to gain new insights into triggering death of pathogenic yeast, extend the potential utility of this tractable genetic model.
Materials and Methods
Yeast strains
Saccharomyces cerevisiae knockout (YKO) strains are from the MATa haploid yeast deletion collection (MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 yfg::KanMX4; Research Genetics) and wild-type BY4741 (MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0) was provided by Jef Boeke. Δfis1* refers to the YKO Δfis1, which has a premature stop codon at amino acid 152 in WHI2 (whi2-d1).21 Yeast were grown in YPD medium.
FUN1 viability assay
Log-phase yeast were stained with FUN1 (Live/Dead Yeast Viability Kit, Molecular Probes, Invitrogen, Carlsbad, CA, USA) by adding 0.5 μl. FUN1 (10 mM in DMSO) and 5 μl calcofluor white M2R (5 mM in H2O) to 1 ml cultures (OD600 adjusted), and 100 μl aliquots were subsequently treated with heat ramp. Immediately following, 1.5 μl of concentrated yeast suspensions were trapped between slide and cover glass and counted using fluorescence microscopy (Nikon Eclipse E800, Nikon Inc., Melville, NY, USA; × 100 objective lens, multipass filter set for DAPI, fluorescein and rhodamine).
Heat ramp cell death assays
For low throughput assays, individual yeast knockout strains were streaked onto YPD agar plates from −80 °C stocks (without thawing) and incubated at 30 °C over two nights. Single colonies were inoculated into 2 ml YPD and incubated at 30 °C on a roller drum overnight. For low throughput assays of log-phase cultures, 0.2 OD600 of yeast from 16 h yeast cultures (∼150 μl) were diluted to 3 ml with fresh YPD and incubated at 30 °C on a roller drum for ∼4 h (longer for dropout media) until OD600 reaches 0.5. About 30 min before reaching OD600 0.5, cell densities of all strains were adjusted to match, so that they arrive at OD600 0.5 simultaneously. Minor differences in cell density were corrected again before heat treatment; no strain-specific differences in growth rates were detected under these conditions. Cell suspensions (100 μl) were transferred to 0.2-ml PCR tubes for heat ramp treatment.
For high throughput analyses of the YKO knockout collection, a 96-well plate was pinned onto YPD agar plates from −80 °C stocks (without thawing), incubated at 30 °C over two nights and transferred with a pin tool into 200 μl YPD for 48 h at 30 °C. Absorbance (Abs595) of cultures was determined with a plate reader. A total of 100 μl of OD600 ∼0.5 yeast cultures (e.g., 10 μl 48-h cultures of OD600 ∼5 into 90 μl YPD) were transferred into 0.2-ml PCR tubes for heat ramp treatment. For high throughput log-phase assays, 48-h cultures in 96-well plates were diluted 30-fold into YPD (∼0.2 OD600 of yeast), incubated for 4 h to reach mid-log phase and 100 μl of each well-mixed sample was transferred into 0.2-ml 96-well PCR tubes for heat ramp treatment.
Yeast were treated with a heat ramp at OD600 0.5 regardless of their metabolic state. Samples (100 μl) were heat-treated using an Eppendorf Mastercycler EP gradient cycler (Eppendorf AG, Hammburg, Germany) (e.g., ramp temperature from RT to 30 °C immediately, hold 1 min at 30 °C, ramp temperature from 30 °C to 62 °C over 20 min, return to RT). Immediately after heat ramp treatment, samples were serially diluted in 96-well microtiter plates (e.g., 20 μl plus 80 μl ddH2O) and 5 μl was plated on YPD agar plates (starting with undiluted). For untreated controls, another 100 μl of the same samples were diluted and plated without heat treatment.
Unlike screens in which treatments are applied for several days, during which new mutations could potentially be selected, our 20-min heat ramp stimulus is too short to allow selection of suppressors. By counting colony numbers regardless of colony sizes at ∼18 h after treatment, we further reduce the influence of any newly arising mutations.
Automated colony counting
Images of ∼18-h yeast colonies on all replicate YPD agar test plates were captured with a BioSpot prototype machine at 1024 by 768 pixels and archived. Automated colony counting was performed on imaged plates using the BioSpot counting software (BioSpot Academic Software Version 5.0, Cellular Technology Limited/CTL) set to parameters in Supplementary Table S2.
Abbreviations
- YKO:
-
yeast knockout
- MAT:
-
mating type
- YPD:
-
yeast-extract peptone dextrose
References
Khurana V, Lindquist S . Modelling neurodegeneration in Saccharomyces cerevisiae: why cook with baker's yeast? Nat Rev Neurosci 2010; 11: 436–449.
Vandenabeele P, Galluzzi L, Vanden Berghe T, Kroemer G . Molecular mechanisms of necroptosis: an ordered cellular explosion. Nat Rev Mol Cell Biol 2010; 11: 700–714.
Fannjiang Y, Cheng WC, Lee SJ, Qi B, Pevsner J, McCaffery JM et al. Mitochondrial fission proteins regulate programmed cell death in yeast. Genes Dev 2004; 18: 2785–2797.
Ellermeier CD, Hobbs EC, Gonzalez-Pastor JE, Losick R . A three-protein signaling pathway governing immunity to a bacterial cannibalism toxin. Cell 2006; 124: 549–559.
Vachova L, Palkova Z . Physiological regulation of yeast cell death in multicellular colonies is triggered by ammonia. J Cell Biol 2005; 169: 711–717.
Engelberg-Kulka H, Amitai S, Kolodkin-Gal I, Hazan R . Bacterial programmed cell death and multicellular behavior in bacteria. PLoS Genet 2006; 2: e135.
Coll NS, Vercammen D, Smidler A, Clover C, Van Breusegem F, Dangl JL et al. Arabidopsis type I metacaspases control cell death. Science 2010; 330: 1393–1397.
Abdelwahid E, Rolland S, Teng X, Conradt B, Hardwick JM, White K . Mitochondrial involvement in cell death of non-mammalian eukaryotes. Biochim Biophys Acta 2010.
Zheng TS, Hunot S, Kuida K, Flavell RA . Caspase knockouts: matters of life and death. Cell Death Differ 1999; 6: 1043–1053.
Peter ME . Programmed cell death: apoptosis meets necrosis. Nature 2011; 471: 310–312.
McPhee CK, Logan MA, Freeman MR, Baehrecke EH . Activation of autophagy during cell death requires the engulfment receptor Draper. Nature 2010; 465: 1093–1096.
Cap M, Vachova L, Palkova Z . How to survive within a yeast colony? Change metabolism or cope with stress? Commun Integr Biol 2010; 3: 198–200.
Griffin DE, Levine B, Ubol S, Hardwick JM . The effects of alphavirus infection on neurons. Ann Neurol 1994; 35 (Suppl): S23–S27.
Kirkegaard T, Jaattela M . Lysosomal involvement in cell death and cancer. Biochim Biophys Acta 2009; 1793: 746–754.
Frank S, Gaume B, Bergmann-Leitner ES, Leitner WW, Robert EG, Catez F et al. The role of dynamin-related protein 1, a mediator of mitochondrial fission, in apoptosis. Dev Cell 2001; 1: 515–525.
Jagasia R, Grote P, Westermann B, Conradt B . DRP-1-mediated mitochondrial fragmentation during EGL-1-induced cell death in C. elegans. Nature 2005; 433: 754–760.
Ivanovska I, Hardwick JM . Viruses activate a genetically conserved cell death pathway in a unicellular organism. J Cell Biol 2005; 170: 391–399.
Palermo V, Falcone C, Mazzoni C . Apoptosis and aging in mitochondrial morphology mutants of S. cerevisiae. Folia Microbiol (Praha) 2007; 52: 479–483.
Scheckhuber CQ, Erjavec N, Tinazli A, Hamann A, Nystrom T, Osiewacz HD . Reducing mitochondrial fission results in increased life span and fitness of two fungal ageing models. Nat Cell Biol 2007; 9: 99–105.
Weinberger M, Mesquita A, Caroll T, Marks L, Yang H, Zhang Z et al. Growth signaling promotes chronological aging in budding yeast by inducing superoxide anions that inhibit quiescence. Aging (Albany NY) 2010; 2: 709–726.
Cheng WC, Teng X, Park HK, Tucker CM, Dunham MJ, Hardwick JM . Fis1 deficiency selects for compensatory mutations responsible for cell death and growth control defects. Cell Death Differ 2008; 15: 1838–1846.
Milleron RS, Bratton SB . ‘Heated’ debates in apoptosis. Cell Mol Life Sci 2007; 64: 2329–2333.
De Virgilio C, Piper P, Boller T, Wiemken A . Acquisition of thermotolerance in Saccharomyces cerevisiae without heat shock protein hsp 104 and in the absence of protein synthesis. FEBS Lett 1991; 288: 86–90.
Buttner S, Eisenberg T, Carmona-Gutierrez D, Ruli D, Knauer H, Ruckenstuhl C et al. Endonuclease G regulates budding yeast life and death. Mol Cell 2007; 25: 233–246.
Fahrenkrog B, Sauder U, Aebi U . The S. cerevisiae HtrA-like protein Nma111p is a nuclear serine protease that mediates yeast apoptosis. J Cell Sci 2004; 117: 115–126.
Wissing S, Ludovico P, Herker E, Buttner S, Engelhardt SM, Decker T et al. An AIF orthologue regulates apoptosis in yeast. J Cell Biol 2004; 166: 969–974.
Li W, Sun L, Liang Q, Wang J, Mo W, Zhou B . Yeast AMID homologue Ndi1p displays respiration-restricted apoptotic activity and is involved in chronological aging. Mol Biol Cell 2006; 17: 1802–1811.
Rinnerthaler M, Jarolim S, Heeren G, Palle E, Perju S, Klinger H et al. MMI1 (YKL056c, TMA19), the yeast orthologue of the translationally controlled tumor protein (TCTP) has apoptotic functions and interacts with both microtubules and mitochondria. Biochim Biophys Acta 2006; 1757: 631–638.
Qiu J, Yoon JH, Shen B . Search for apoptotic nucleases in yeast: role of Tat-D nuclease in apoptotic DNA degradation. J Biol Chem 2005; 280: 15370–15379.
Ligr M, Velten I, Frohlich E, Madeo F, Ledig M, Frohlich KU et al. The proteasomal substrate Stm1 participates in apoptosis-like cell death in yeast. Mol Biol Cell 2001; 12: 2422–2432.
Cheng WC, Leach KM, Hardwick JM . Mitochondrial death pathways in yeast and mammalian cells. Biochim Biophys Acta 2008; 1783: 1272–1279.
Cheng EH, Levine B, Boise LH, Thompson CB, Hardwick JM . Bax-independent inhibition of apoptosis by Bcl-XL. Nature 1996; 379: 554–556.
Longo VD, Mitteldorf J, Skulachev VP . Programmed and altruistic ageing. Nat Rev Genet 2005; 6: 866–872.
Madeo F, Herker E, Maldener C, Wissing S, Lachelt S, Herlan M et al. A caspase-related protease regulates apoptosis in yeast. Mol Cell 2002; 9: 911–917.
Dudgeon DD, Zhang N, Ositelu OO, Kim H, Cunningham KW . Nonapoptotic death of Saccharomyces cerevisiae cells that is stimulated by Hsp90 and inhibited by calcineurin and Cmk2 in response to endoplasmic reticulum stresses. Eukaryot Cell 2008; 7: 2037–2051.
Enoksson M, Salvesen GS . Metacaspases are not caspases – always doubt. Cell Death Differ 2010; 17: 1221.
Wysocki R, Kron SJ . Yeast cell death during DNA damage arrest is independent of caspase or reactive oxygen species. J Cell Biol 2004; 166: 311–316.
Hardwick JM, Cheng WC . Mitochondrial programmed cell death pathways in yeast. Dev Cell 2004; 7: 630–632.
Fadeel B, Xue D, Kagan V . Programmed cell clearance: molecular regulation of the elimination of apoptotic cell corpses and its role in the resolution of inflammation. Biochem Biophys Res Commun 2010; 396: 7–10.
Teng X, Hardwick JM . Reliable method for detection of programmed cell death in yeast. Methods Mol Biol 2009; 559: 335–342.
Acknowledgements
We thank Jef Boeke for MATa yeast strains and valuable advice. This project was supported by the National Institutes of Health R01 GM077875 (JMH).
Author Contributions
XT conceived and performed the heat ramp experiments presented; WCC contributed to heat ramp assay development and performed the acetic acid assays; BQ contributed to assay development; TY and KR participated in data collection; FJP and MDB contributed to data analyses; PVL and TH developed and optimized the BioSpot instrument for yeast colony counting; JMH conceived the project, oversaw data analyses, and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Supplementary Information accompanies the paper on Cell Death and Disease website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Teng, X., Cheng, WC., Qi, B. et al. Gene-dependent cell death in yeast. Cell Death Dis 2, e188 (2011). https://doi.org/10.1038/cddis.2011.72
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2011.72
Keywords
This article is cited by
-
ATP-citrate lyase B (ACLB) negatively affects cell death and resistance to Verticillium wilt
BMC Plant Biology (2022)
-
Genetic interaction between RLM1 and F-box motif encoding gene SAF1 contributes to stress response in Saccharomyces cerevisiae
Genes and Environment (2021)
-
Autophagy, programmed cell death and reactive oxygen species in sexual reproduction in plants
Journal of Plant Research (2017)
-
Genome-wide identification of genes involved in the positive and negative regulation of acetic acid-induced programmed cell death in Saccharomyces cerevisiae
BMC Genomics (2013)