Abstract
Alterations in the functions of neuronal RNA-binding proteins (RBPs) can contribute to neurodegenerative diseases. However, neurons also express a set of widely distributed RBPs that may have developed specialized functions. Here, we show that the ubiquitous member of the otherwise neuronal Elavl/Hu family of RNA-binding proteins, Elavl1/HuR, has a neuroprotective role. Mice engineered to lack exclusively HuR in the hippocampal neurons of the central nervous system (CNS), maintain physiologic levels of neuronal Elavls and develop a partially diminished seizure response following strong glutamatergic excitation; however, they display an exacerbated neurodegenerative response subsequent to the initial excitotoxic event. This response was phenocopied in hippocampal cells devoid of ionotropic glutamate receptors in which the loss of HuR results in enhanced mitochondrial dysfunction, oxidative damage and programmed necrosis solely after glutamate challenge. The molecular dissection of HuR and nElavl mRNA targets revealed the existence of a HuR-restricted posttranscriptional regulon that failed in HuR-deficient neurons and is involved in cellular energetics and oxidation defense. Thus, HuR acts as a specialized controller of oxidative metabolism in neurons to confer protection from neurodegeneration.
Similar content being viewed by others
Main
Excitatory neurotransmitters are important mediators of synaptic connectivity and plasticity in the central nervous system (CNS); however, acute and/or chronic CNS insults can lead to their excess and support neuronal loss as suggested for neurodegenerative syndromes like amyotrophic lateral sclerosis (ALS), Parkinson’s disease, multiple sclerosis and others.1 For glutamate, these neurotoxic effects proceed via: (a) a massive influx of calcium ions induced by its surface receptors (e.g. ionotropic GluRs) leading to intracellular oxidative stress and (b) the concomitant blockade of antioxidant countermeasures (e.g. blockade of the cystine/glutamate antiporters driving the synthesis of glutathione (GSH)). As endogenous defense systems fail to detoxify, intracellular oxidants ultimately lead to the activation of cellular execution programs and neurodegeneration.1, 2, 3 The balance between neuronal death and survival relies on several regulatory modalities that include the posttranscriptional regulation (PTR) of mRNAs encoding relevant factors. Owing to their complex features, neurons use a variety of PTR machines involved in the processing, localization and translation of RNA molecules engaged in activity-dependent reactions even at distant structures (e.g. dendrites). Trans-acting factors that guide such PTR events include RNA-binding proteins (RBPs) and noncoding RNA populations, which target a range of motifs on mRNAs.4 The strong correlation between RBP dysfunction and neuropathology has been exemplified in ALS and Fragile X/ataxia syndromes. However, the large fraction of mammalian RBPs expressed in the nervous system5 remains largely unexplored.
The family Elavl/Hu of RBPs share significant homology to the Drosophila ELAV gene and appear as highly relevant to neuronal processes.4,6,7 Consistently, three members of this family (HuB/Hel-N1/Elavl2, HuC/Elavl3 and HuD/Elavl4) are mostly expressed in neurons (nElavls); the fourth member – Elavl1/HuR – is expressed in both neuronal and non-neuronal tissues. Elavl/Hu proteins share similarities in their structures, which include RNA-recognition motifs for specific U-rich RNA structures. Furthermore, they contain domains for nuclear translocation and cytoskeletal interactions allowing for their signal-dependent subcellular localization. A considerable body of literature demonstrates the importance of nElavls in neuronal differentiation and the PTR of mRNAs involved in neuronal excitation and synaptic plasticity.7, 8, 9, 10 On the contrary, HuR was considered as a redundant factor in neurons; recent evidence contradicts this supposition. Genetic ablation and molecular experiments demonstrated that HuR can promote the renewal of neuronal progenitors, whereas in differentiated cells it is silenced by nElavls via the alternative polyadenylation of its mRNA.11,12 In adult neuron dendrites exposed to dopaminergic or glutamatergic activators, HuR and nElavls recognize overlapping and also distinct RNA targets, suggesting a functional disparity in activity-dependent responses.7 In non-neuronal tissues, HuR has a proven involvement in a multitude of cellular programs (e.g. cell cycle control, inflammation, genotoxic stress, cell death) by altering the PTR of its target mRNAs – as such it could also be engaged in neurophysiologic or neurodegenerative settings. Here, we dissect HuR’s functions in adult hippocampal neurons via the genetic restriction of its ablation and reveal its involvement in neuronal protection.
Results
The loss of Elavl1/HuR in hippocampal neurons does not compromise the expression of neuronal Elavls
To examine whether HuR is involved in the activity-dependent responses of adult neurons, we focused on the brain hippocampus; this structure contains highly excitable, glutamatergic, pyramidal neurons in its cornus ammonis (CA) subregions that receive peripheral input from granule cells and interneurons of the dentate gyrus (DG). Despite the extensive regional variability in the expression of Elavl/Hu proteins6 in the CNS (Supplementary Figure 1), both HuR and nElavls are present in the hippocampus formation as indicated by the signals detected with anti-HuR and anti-HuD antisera (Figure 1a); we selected to focus primarily on HuD as a representative of nElavls given prior indications on its involvement in the activity-dependent responses of pyramidal neurons.7,13 HuR’s expression is strong in the CA1 and DG regions, but lower in the CA2/CA3 regions. In contrast, the expression of HuD is strong in the CA2/CA3 regions, but low in the CA1 and DG regions. As in most non-neuronal tissues, HuR localizes predominantly to the nuclei of the neurons’ somata, whereas HuD extents strongly to axons and dendrites (Figure 1c). These differences in the hippocampal expression of Elavls could reflect their differential requirement in neuronal subsets and/or activity-dependent responses.
The loss of HuR from hippocampal neurons does not affect neuronal Elavl expression. (a–d) Immunohistochemical detection of Elavl1/HuR and Elavl4/HuD in the hippocampi of control and CN-KO mice. (e) Detection of β-galactosidase activity in brains from CamKIICre+ROSA26fl mice. (f) Detection of Elavl1fl locus recombination via Southern blot analysis of genomic DNA isolated from different brain regions of Elavlfl/fl mice in the presence or absence CamKIICre. (g) Immunodetection of Elavls in lysates from control and CN-KO brain compartments using antisera for all Elavls (3A2 antibody), Elavl4/HuD and Elavl1/HuR. Lysates from the lung serve as a site for unique HuR expression. β-Actin serves as a loading control. Numerical marking on the left indicates protein marker positions. Note that both the 3A2 and anti-HuD antibodies react with bands higher than their predicted molecular weights (marking in right). In the 3A2 blots, bands higher than HuR may also include Elavl2 and 3, and hence are annotated as nElavls. Magnification bars: a, b and e, 100 μm; c and d, 20 μm. Cb, cerebellum; CrB, cerebrum; FL, frontal lobe; H, hematoxylin; Hc, hippocampus; Lu, lung; OlB, olfactory bulb; NFR: Nuclear Fast Red; ST, striatum
To assess this directly, we induced the selective deletion for HuR in the hippocampus formation. Mice in which an essential region of the Elavl1 gene was flanked by loxP sequences14 were crossed to mice expressing Cre recombinase under the control of an 8.5-kb promoter fragment of the calcium/calmodulin-dependent protein kinase II-α (CamKIIa) gene.15 We selected this CamKIICre+ transgenic line for two reasons: (a) Cre expression is restricted to differentiated, postmitotic neurons allowing us to avoid problems arising from HuR’s involvement in neuronal development; and (b) as elucidated by crossings to ROSA26 reporter mice16 and detection of β-galactosidase activity, Cre-mediated recombination is prevalent in the pyramidal neurons of the hippocampus, and lesser to negligible in other brain areas (Figure 1e). CamKIICre+Elavl1fl/fl (referred hereafter as CN-KO) mice were born normally with no apparent gross phenotypic abnormalities. Southern blot analysis of genomic DNA from different brain regions revealed that the recombination of Elavlfl locus in the mutant mice was near 70–80% in hippocampal areas, <20% in frontal lobe and cerebral areas and absent in the cerebellum (Figure 1f). Immunohistochemistry with anti-Elavl1/HuR and anti-Elavl4/HuD antisera revealed the consistent absence of HuR protein in ≥80% of hippocampal neurons in CN-KO mice relative to its presence in control (CamKIICre+Elavlfl/+ and CamKIICre−Elavl1fl/fl) neurons and the presence of HuD protein in the same neurons (Figures 1a–d). Similarly, immunoblots with extracts from different brain regions with anti-Elavl1/HuR antibody revealed the consistent reduction of this protein primarily in the hippocampus (Figure 1g). This reduction was also confirmed using an antibody known to react with HuR in all peripheral tissues (3A2) and also with the other Elavls in the brain (albeit with unknown affinities for the latter). Although we cannot discriminate the type of neuronal Elavl recognized by this antibody, comparative immunoblots with anti-Elavl4/HuD antisera overlapped in size with most bands and remained unperturbed in HuR-deficient neurons. Thus, CN-KO mice are characterized by a significant and specific reduction of HuR protein in the hippocampal neurons, allowing us to further investigate HuR’s functional role in neuronal excitation.
The loss of HuR sensitizes hippocampal neurons to glutamatergic toxicity
Changes in the activity-dependent responses of hippocampal neurons can be detected using strong excitatory agents that cause neuronal dysfunction and death. Kainic acid (KA) is a potent agonist of a subtype of glutamate receptors that can cause such an excitotoxic response;17,18 its systemic administration targets specifically hippocampal CA1 and CA2/3 neurons, leading to early epileptic seizures (minutes to hours) and delayed neuronal death (past 5 days). To assess HuRs involvement in KA-induced neuronal dysfunction, we treated control (Elavl1fl/fl and CamKIICre−Elavlfl/fl) and CN-KO mice with two systemic doses of KA (20 and 30 mg/kg). These doses were previously reported to induce a mild excitotoxic response in the genetic background of our mutations (C57BL/6),19,20 which is partially resistant to KA challenges. Following KA administration, mice were initially scored for the extent of epileptic seizures occurring within the first 3 h. In control mice, seizure-rating scores increased by 10 min, peaked between 20 and 30 min and gradually subsided thereafter. Following the same treatment, female CN-KO mice showed a statistically significant reduction in the average seizure intensity and convulsion-induced death (Figures 2a and b), suggesting HuR’s involvement in the excitation circuits driving seizure elicitation. However, this difference appeared to be sex-restricted; both seizure activity and mortality in male CN-KO mice were statistically similar to control groups (Figures 2a and b).
Differential involvement of HuR in kainate-induced excitation versus neurodegeneration. (a) Measurement of the extent of epileptic seizures after administration of two doses of KA. Data were derived from: male (♂) CN-KO and control mice at 30 mg/kg (n=17 and 20) and 20 mg/ml (n=9 and 7); and female (♀) CN-KO and control mice at 30 mg/kg (n=21 and 19) and 20 mg/ml (n=8 and 8). (b) Kaplan–Meier distribution of control and CN-KO male and female mice that survived from early epileptic seizures induced by KA administration. (c) Photomicrographs of hippocampi in paraffin brain sections from control and CN-KO mice killed 5 days following the administration of KA and chemically stained with hematoxylin/eosin, Nissl or immunostained for Mac3 and counterstained with hematoxylin. Magnification: upper panels, 100 μm; lower panels, 20 μm. (d) Histopathologic quantitation of the damaged area (hematoxylin/eosin) and loss of neurons (Nissl) from n=13 mixed-sex CN-KO mice and 16 control mice, and microglia activation (Mac3; CN-KO: n=7; control: n=17). *Statistical differences to control groups with a P≤0.05
Next, we assessed the degree of permanent neuronal damage in surviving mice that is manifested days after the initial KA shock.19,21 We focused specifically on CA1 and CA2/3 pyramidal neurons of the hippocampus, which are known to exhibit selective vulnerability to KA-induced neurotoxicity.17,18 Hematoxylin/eosin (cell bodies) and Nissl (neuronal perikarya) stainings were performed on brain sections from animals killed 5 days following the administration of KA. Strikingly, the microscopic examination of neuronal damage throughout the CA1 to CA3 subfields demonstrates that the extent of KA-induced hippocampal lesions observed in CN-KO mice was significantly greater than in control mice (Figures 2c and d). A partial increase in the microglial protein (Mac3) verified the activation of local microglia, which usually occurs as a response to neuronal damage (Figures 2c and d). Examination for the Elavl content in KA-challenged control mice demonstrated that both HuR and HuD displayed a biphasic pattern of expression reaching a maximum during the early stage, but decreasing during the degenerative stage (Supplementary Figure 2). In the challenged CN-KO hippocampi, the response of HuD was maintained, suggesting that the loss of HuR itself accounts for increases in neuronal death (Supplementary Figure 2). Most importantly, the neurodegenerative response of CN-KO mice was similar in both sexes, irrespective of their differences in seizures, suggesting that the exacerbation of the neuronal death following the deletion of HuR is not the result of enhanced receptor activity but rather due to changes in the death process.
The loss of HuR sensitizes hippocampal neurons to oxidative damage
Our in vivo measurements suggested the differential involvement of HuR in neuronal excitation versus neuronal death. To gain further insight at the cellular level, we isolated primary neurons from the hippocampi of prenatal E19 embryos. Cultures were established in media supplemented with trophic factors and antioxidants and under conditions that prohibit glial expansion. We anticipated that the CamKIICre transgene would become active during the terminal in vitro differentiation of CN-KO neurons. Indeed, the recombination of the loxP-flanked Elavl1 locus in mutant cultures reached 60% between the sixth and the eighth day in vitro (DIV6–8) correlating with the progressive expression of the Cre-mRNA (Supplementary Figures 3a and b). At DIV7, control and mutant cultures contained β-tubulin-expressing neurons possessing branched and overlapping neurites (Supplementary Figures 3d). Mutant cultures also showed a substantial loss in HuR protein and Elavl1 mRNA expression whereas they maintained Elavl2 and Elavl4 mRNAs at control levels; however, we did note a partial reduction in the Elavl3 mRNA (Supplementary Figures 3c and 4). Between DIV7 and 10 mutant cultures degenerated rapidly showing extensive fragmentation of neurites, nuclear condensation and cell death (Supplementary Figures 3d–f), thus prohibiting analysis in fully developed neurons possessing axons and synapses (i.e. past DIV12).22,23 However, less mature hippocampal neurons (DIV5–7) have a limited expression of functional ionotropic and metabotropic GluR, as well as functional cystine/glutamate antiporter x(c)(−) that elicit glutamate-induced toxicity.24,25 Thus, we removed antioxidants from the culturing media, and using a sensitive viability dye (PrestoBlue), we determined that DIV7 neurons exposed to KA for 3, 12 and 20 h showed a small and late toxicity response, which was enhanced in HuR-lacking cultures (Figure 3a). In contrast, glutamate induced a potent detrimental response in DIV7 neurons, which was significantly augmented in HuR-lacking cultures. Under these conditions, the mRNAs of nElavls responded in a differential manner, but the loss of HuR did not alter their response (Supplementary Figure 4). The concomitant blockade of NMDA and AMPA receptors by the dl-2-amino-5-phosphonovaleric acid (AP5) and CNQX antagonists provided only a small benefit for protection against glutamate in both cultures. A more profound prosurvival effect was observed when control and mutant neurons were pretreated with N-acetyl-cysteine (NAc), which acts as a reactive oxygen species (ROS) scavenger and potent antioxidant. Collectively, these data suggested that HuR could be required for defense against glutamate-induced oxidative damage. We do note, however, that during the window of our analysis (i.e. 0–20 h), basal survival values in HuR-lacking cultures declined significantly, thus obscuring the derivation of definitive conclusions.
The loss of HuR enhances the sensitivity of hippocampal neurons to glutamate (Glu)-induced oxidative damage. (a) Timed death response of primary hippocampal cultures from control and CN-KO embryos in response to 20 μM KA or 100 μM glutamate (Glu) (first panel), and in the absence or presence of antagonists for ionotropic Glu receptors (AP5 and CNQX) or the antioxidant NAc (second panel, control; third panel, CN-KO). Data were derived from viability assays using PrestoBlue repeated for two times with three individual cultures per experiment. *Statistical differences to 0 h. (b) Death response of HT-22 knockdown cells to increasing doses of Glu for 20 h. Data were derived from MTT assays repeated for at 4–8 times with triplicate cultures per experiment. *Statistical differences to parental and control groups. (c) Detection of ROS in cells labeled with DCFDA and treated with the indicated quantities of Glu for 10 h. Data were derived (mean fluorometric values±S.D.) from two independent experiments with four replicas per experiment. *Statistical differences to parental and control groups with a P≤0.05. (d) Detection of mitochondrial membrane potentials or intracellular superoxide release via the flow cytometric analyses of changes in the intensities (FI) of DiOC6 or MitoSox Red, respectively; cells were monitored 10 h after exposure to 3 and 6 mM Glu. Numbers indicate the percentage of cells with reduced potential (lower intensity peak in DiOC6) or increased superoxide (high intensity peak in MitoSox) in each case. Representative data were derived from two independent experiments with triplicate per group per experiment. (e) Immunohistochemical detection of mitochondrial ATPase (catalytic alpha-subunit) as a marker of intact mitochondria in the CA2/3 region of control and CN-KO hippocampi challenged with KA. Counterstain: hematoxylin. Boxes in low magnifications (left; bar, 100 μm) indicate areas magnified in right (bar, 20 μm). Note the perinuclear granular staining in control neurons and its loss in intact and condensing hippocampal neurons from CN-KO mice (arrows). (f) Effect of the antioxidants NAc (1 mM), Trolox (0.1 mM) and PD146176 (0.5 μM) to the Glu-induced death response of HT-22 sublines and at 20 h after exposure. Data in bar graphs (mean values±S.D.) were derived from MTT assays repeated for at least four times with triplicate cultures per experiment. *Statistical differences to Glu (+solvent vehicle)-treated groups with a P≤0.05
To discriminate further excitatory stress from oxidative neurotoxicity in a system devoid of the complications in neuronal differentiation, we switched to HT-22 hippocampal neurons. These cells originate from the mouse CA1 region and are a suitable model for glutamate-induced toxicity rather than excitotoxicity, as they lack ionotropic GluRs; instead, glutamate blocks the cystine/glutamate antiporter system xc−.26 Using shRNA (Supplementary Figures 5a and b), we knocked down HuR in HT-22 neurons and compared their responses with mock and scrambled shRNA transfectants. 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assays revealed that glutamate induced a dose-dependent death response on control cells with a median lethal dose (LD50) of 5 mM (Figure 3b). In transiently transfected HT-22 cells, a 90% reduction in HuR decreased the LD50 of glutamate to 3 mM. Stable shHuR clones showing a 60–90% loss in HuR protein (clones c4 and 5; named HuRmod and HuRlo hereafter) relative to control clones (selected scramble c2 and parental HT-22 cells; HuRhi and HT22p hereafter) verified that glutamate sensitivity correlates inversely to the expression of HuR (Figure 3b and Supplementary Figures 5a and b). This effect appeared specific since the death response of HuRlo cells to peroxide or tumor necrosis factor was similar to that of HuRhi controls (Supplementary Figure 6).
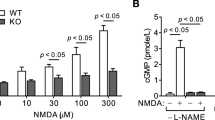
The response of HT-22 neurons to glutamate includes (a) the depletion of GSH, (b) activation of intracellular oxidative enzymes, such as 12/15-LOX lipoxygenase, (c) the depolarization of mitochondrial electron transport chain and (d) production of ROS, superoxide and hydroperoxy radicals that oxidize further lipids and proteins.27,28 The depleting response of oxidized and reduced forms of glutathione (GSH+GSSG) to glutamate was comparable between HuRhi, HuRmod and HuRlo cells, suggesting that HuR does not interfere with the xc− antiporter system (Supplementary Figure 5d). In contrast, the incorporation of 2′7′-dichlorofluorescein diacetate (DCFDA) was elevated in HuRlo cells signifying an increase in their intracellular ROS (Figure 3c and Supplementary Figure 5c). Similarly, downstream targets of ROS, such as SOD1/2 and COX2,27,28 appeared augmented in glutamate-challenged HuRlo cells (Supplementary Figure 5e). On the other hand, the increase in intracellular ROS correlated to the enhanced dysfunction of mitochondria in HuRlo neurons; this was demonstrated by significant drop in 3,3′-dihexyloxacarbocyanine iodide (DiOC6(3)) marking their membrane potential; and an enhancement in the incorporation of MitoSox marking the augmented release of superoxide from damaged organelles (Figure 3d). Similar to HuRlo HT-22 neurons, excessive mitochondrial damage was detected in the CA2 neurons of CN-KO mice challenged with KA. This was revealed via the immunohistochemical detection of the catalytic, ATP synthase αlpha-subunit, which is part of the mitochondrial oxidative phosphorylation complex V. ATP synthase-positive signals were maintained in CA2 control neurons and the CA1 neurons of CN-KO mice, indicating that the expression of this subunit is not affected by the loss of HuR (Figure 3e and Supplementary Figure 7); however, these signals were nearly undetected in the challenged CA2 neurons of CN-KO mice, indicating that the activity-dependent loss of active mitochondria is enhanced by the absence of HuR.
To prove oxidative damage as the key problem of HT-22 HuRlo cells, we tried to revert their death response to glutamate using three antioxidants: (a) NAc; (b) the vitamin E derivative, 6-hydroxy-2,5,7,8-tetramethylchroman-2-carboxylic acid (Trolox); and (c) an inhibitor of lipoxygenases (LOX; PD146176). These agents were used at concentrations that were previously reported to inhibit toxicity in HT-22 cells.27,28 Despite the increased sensitivity of HuRlo cells to glutamate, they became fully protected by antioxidants as did HuRhi cells (Figure 3f). Thus, HuR acts to protect neurons from oxidative neurotoxicity.
HuR controls the sensitivity of neurons to necrosis
The flow cytometric detection of Annexin-V (AnxV) and propidium iodide (PI) suggested that all HT-22 transfectants followed a necrotic fate (AnxV+/PI+ and AnxV−/PI+) rather than an apoptotic fate after glutamate challenge; this necrotic response was clearly enhanced following the knockdown of HuR (Figure 4a). The pancaspase inhibitor zVAD-FMK did not block the toxicity of HT-22 cells to glutamate (Figure 4b) nor did any of the transfectants display cleavage of PARP (Figure 4f); these observations confirmed that apoptosis is not the predominant form of death response in these cells28, 29, 30 nor it comes into effect because of HuR’s loss. In contrast, a beneficial effect of the RIP-kinase 1 inhibitor necrostatin-1 (Nec-1) in HT-22 cells29 was partially maintained in HuRlo cells, suggesting that necrotic/necroptotic pathways are maintained in the absence of HuR (Figure 4b).
The loss of HuR enhances BiD-mediated necrosis in glutamate-challenged neurons. (a) Flow cytometric detection of apoptotic and necrotic HT-22 cells containing or lacking HuR and in response to glutamate for 14 h and detection of AnxV and PI. Representative data were derived from two independent experiments with four replicas per group. (b) Effect of inhibitors of Nec-1 (25 μM), apoptosis (zVAD-fmk; 50 μM), BiD function (BI6C9; 10 μM) or DMSO vehicle on the death response of HT-22 sublines challenged with 6 mM glutamate for 20 h. Data in bar graphs (mean values±S.D.) were derived from MTT assays repeated for at least three times with triplicate cultures per experiment. *Statistical differences to glutamate (+solvent vehicle)-treated groups with a P≤0.05. (c) Effect of the Bcl-2/Bcl-xl inhibitor ABT-737 on the death response of HT-22 sublines challenged with 6 mM glutamate for 20 h. Data in histograms (mean values±S.D.) were derived from MTT assays repeated for two times with triplicate cultures per experiment. *Statistical differences to HuRhi controls. (d and e) Immunodetection of AIF and BiD in total cell, nuclear (N) and cytoplasmic (C) lysates from HT-22 cells containing or lacking HuR and exposed to 4 mM glutamate for the indicated time points. GAPDH, nuclear lamin A/C and actin are shown as loading and fractionation controls. Note the absence of detectable cleaved BiD (tBiD). (f) Representative immunodetection of Bcl-xl, Bcl-2, Bad, Bax, PARP and p53 in total cell lysates from HT-22 cells containing or lacking HuR and exposed to 4mM glutamate for the indicated time points. GAPDH is shown as the loading control. In all immunoblots, numerical marking on the left represent predicted molecular weights for each molecule
The death responses of HT-22 neurons require the translocation of full-length Bid to the mitochondria aiding their destruction, the release of AIF and its translocation to the nucleus where it facilitates DNA damage.28,30 To examine this possibility, we blocked Bid’s function in HuRlo cells via the BI6C9 inhibitor;28 the death of HuRlo sublines was nearly abolished by the neutralization of Bid and was similar to that of HuRhi cells (Figure 4b). Examination of Bid and AIF proteins in extracts from challenged cells demonstrated that the loss of HuR did not affect their accumulation or nuclear translocation (Figures 4d and e); similarly, the levels of other proteins involved in DNA damage-like p53− were similar to those in controls (Figure 4f). The downstream interactors of Bid, Bax and Bad were normally expressed in HuRlo cells; however, we did note an increase in the inhibitory phosphorylation of Bad. Bad’s phosphorylation is promoted by MAPK/SAPK signals that facilitate the oxidative response of HT22 cells,31 but these were not affected by HuR’s loss (Supplementary Figure 8).
Collectively, neuronal HuR appears to act for the resistance of mitochondria to Bid-mediated necrosis reciprocating the effects of prosurvival Bcl-2 family members. Indeed, HuRlo neurons expressed lower levels of basal of Bcl-2 but not Bcl-xl proteins; interestingly, glutamate led to the rapid disappearance of Bcl-2 in both control and test cultures (Figure 4f). To correlate this to neurotoxicity, we selectively blocked the functions of prosurvival Bcl-2 family members by the BH3 mimetic drug ABT-737, which then enforces Bax activation and death.32 At basal conditions, ABT-737 was sufficient to induce a lethal response to HT-22 cells; this response was enhanced in HuRlo HT-22 cells (Figure 4c). In the presence of glutamate, ABT-737 had a minimal additive effect in HuRhi cells that was diminished in HuRlo cells – demonstrating that prosurvival programs counteracting mitochondrial damage are inhibited by glutamate and HuR deficiency alike.
HuR targets and regulates a group of RNAs involved in oxidative metabolism to confer neuroprotection
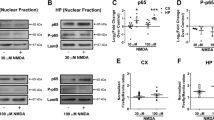
qRT-PCR analysis indicated that resting HT-22 HuRhi cells express Elavl1, Elavl2 and Elavl4 mRNAs, which were not altered in HuRlo cells (Figure 5a); on the other hand, we failed to detect the Elavl3 mRNA in either of these cultures. Interestingly, and in comparison with brain tissue, only the proteins of HuR and HuD could be detected in HT-22 cells (Figure 5b and Supplementary Figures 9a and b). Based on immunoblots with the anti-Elavl antibody (3A2) and comparisons to anti-HuR and anti-HuD immunostains, it seems that HT-22 cells may contain a higher HuR content than HuD (Figures 5b and d and Supplementary Figures 9a and b), although the affinity of the 3A2 antibody for HuD relative to HuR is unknown. Most strikingly, we could not detect any HuB protein with the specific antibody used, despite the presence of its mRNA in both HuRhi and HuRlo cells and its significant enhancement under glutamate (Figure 5a and Supplementary Figures 9a–c). Glutamate also induced small yet differential responses in Elavl1 and Elavl4 mRNAs in HuRhi cells; in HuRlo cells, however, the Elavl4 mRNA response was diminished at late time points (Figure 5a). These differences in mRNA did not directly correlate with the response of the corresponding proteins. At 10 h following exposure, glutamate induced a significant increase in both total and cytoplasmic HuR but not in HuD (Figures 5b–e); afterwards – that is, at 18 h –the levels of HuR protein declined (Figures 5b and c). 2D gel analysis of HuR-containing extracts from challenged HT-22 cells suggested that glutamate could induce posttranslational modifications to HuR protein (Figure 5f). Untreated cells contained a band of 34 kDa mass of a pI 8.9, predicted for neutral HuR. Glutamate led to the appearance of at least two more acidic versions of HuR (pI 8.6 and 8.7, respectively) compatible with Ser/Thr monophosphorylation or N-terminal acetylation as indicated by the ProMoST algorithm.33
Examination of Elavl responses in glutamate-challenged neurons. (a) Bar graphs depicting differences in Elavl1, Elavl2 and Elavl4 mRNAs in HuRhi (white bars) and HuRlo (black bars) HT22 cells after treatment with 4 mM glutamate for 10 and 20 h. Data are presented as fold change±S.D. relative to HuRhi untreated cells and derive from qRT-PCR experiments using RNA from three individual cultures. *Increases and **decreases with P<0.01. (b) Representative immunodetection of all Elavls (3A2), Elavl4/HuD and Elavl1/HuR in total cell lysates from HT-22 cells containing or lacking HuR and exposed to 4 mM glutamate for the indicated time points. GAPDH is shown as the loading control. Numerical annotations on the right represent protein marker positions; named marking on the left indicate positions for the predicted molecular weights. (c) Quantitation of immunoblots is represented in (b); data were derived from three independent experiments. (d) Representative immunodetection of Elavls (3A2 antibody) in fractionated nuclear (N) and cytoplasmic (C) lysates from HT-22 cells containing or lacking HuR and exposed to 4 mM glutamate for the indicated time points. Nuclear lamin A/C and actin are shown as loading and fractionation controls. Marking on the right indicate positions at the predicted molecular weight for each molecule. Based on (b) and Supplementary Figure 9, we predict that nElavl corresponds to HuD. (e) Quantitation of immunoblots is represented in (d); data were derived from three independent experiments. (f) Immunodetection of HuR in immunoblots from 2D gels analyzing total lysates from parental HT-22 cells as untreated or treated with 4mM glutamate for 10 h. Direction (charges) and detected pI values are indicated
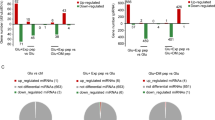
Given the known involvement of HuR in mRNA regulation, alterations of its protein should affect the posttranscriptional fates of RNA targets acting to oppose the neurotoxic response. Some of these associations could also be shared with nElavls. To discriminate HuR-restricted from Elavl combinatorial associations, we used a two-step strategy of ribonucloprotein (RNP) immunoprecipitations (IPs)34 (Supplementary Figure 10a). In the first step, we aimed to identify mRNA associations with all Elavl/Hu’s altered in an HuR-deficient setting; to do so, we immunoprecipitated Elavl-containing RNPs from HuRhi neurons using anti-Elavl antisera (3A2) and following exposure to glutamate for 10 h (Supplementary Figure 10b). The RNA content of these RNPs was analyzed via hybridizations to microarrays (RIP-Chip); bioinformatic comparisons with control IgG IPs revealed 2129 RNAs enriched in the Elavl/RIPs. To relate interactions to the effects of HuR deficiency in neurons, we performed differential microarray hybridization of total RNA from control HuRhi and HuRlo cells challenged with glutamate under the same conditions and identified 297 differentially expressed genes (DEGs). Next, DEGs were compared with interacting mRNAs to reveal 61 Elavl targets misexpressed in HuRlo cells (Supplementary Figure 10c and Supplementary Tables 1 and 2). The Elavl1/HuR mRNA was among those identified validating our approach as it is regulated both by its protein and nElavls12,35 and has been knocked down in HT-22 cells. To restrict our analyses to putative HuR targets, we examined whether the human homologs of these 61 mRNAs were identified to physically interact with HuR in PAR-CLiP36,37 assays deposited in the DoRiNA38 database; based on phenotypic relevance, we selected six such mRNAs. These included mRNAs encoding for: (a) major energy sensors involved in oxidative metabolism such as Ppargc1a (aka PGC1α), which is a master regulator of ROS-scavenging enzymes, and mitochondrial biogenesis, previously shown to counteract cerebral ischemia;39 and NQO1 (NAD(P)H dehydrogenase, quinone 1), a major antioxidant whose dysfunction associates with many disease states and cancer;40 (b) pleiotropic controllers involved in cell survival and cell stress such as Myc41 and Cirbp (cold-inducible RNA-binding protein);42 and (c) factors involved in neuronal polarity and plasticity such as semaphorin 3a (Sema3a) and Gja1 (connexin-43).43 One gene that associated with Elavls but was not identified in DoRiNA (Prkar2a) was also selected because of the relevance of protein kinase A signals to neuronal excitation.44 For validation, we extended our list to mRNAs: (a) which did not show changes in expression but could associate with Elavls in our study and were represented in DoRiNA (Bcl-2, Sods); (b) which are involved in oxidative phosphorylation and ion transport and are recognized by Elavls as indicated herein and by others (plasma membrane ATPase, Atp2b4 and mitochondrial ATP synthase subunit β Atp5b)45 or hinted by our phenotypic observations (mitochondrial ATP synthase subunit α, Atp5a1; Figure 3e); and (c) which showed networking relationships with Ppargc1a and NQO1 in antioxidant defenses such as Ppargc1β, cytochrome C (Cycs) and the transcription factors Nef2L2 (Nrf2) and FOXO1. The latter was also inferred upon the prediction of aberrant transcription factor activities by the corresponding DEGs and Elavl-interacting RNAs (Supplementary Table 3), which indicated the lack FOXO1 transcriptional activity.
In the second step, we refined our search for exclusive HuR targets by detecting the selected mRNAs in HuR/RNP versus HuD/RNP IPs from HuRhi neurons (Figure 6a and Supplementary Figure 10d) and qRT-PCR. HuR/RNPs contained exclusively the Bcl-2, Cirbp, Gja1, Elavl1, Myc, NQO1 and PGC1 mRNAs. Crossrecognition by both HuR and HuD was observed for Prkar2 and Sod2 mRNAs – the latter being in compliance also to the bioinformatic comparison of our Elavl/RNP IP data to those previously published for HuD46 (Supplementary Table 4). Interestingly, these two mRNAs were among the very few mRNAs crossrecognized in all our IPs that appeared affected in HuRlo cells. Finally, the mRNA encoding Atp2b4 was marginally interacting with HuD, whereas those of Atp5a1, Atp5b, Cycs, Nrf2, FOXO1, Ppargc1β, Sod1 and Sema3A did not associate with either RBP (Figure 6a).
Exclusive HuR targeting of mRNAs used during antioxidant defenses. (a) qRT-PCR validation of selected mRNA targets in HuR/RNP IPs (black bars) or HuD/RNP IPs (gray bars) from HuRhi HT-22 neurons and in the absence (−) or presence (+) of 4 mM glutamate for 10 h. Data were derived form three biologic replicas and are presented as fold enrichment (±S.D.) of each mRNA in the IP samples compared with its abundance in IgG IPs. (b) Bar graphs depicting differences in selected mRNAs in HuRhi (white bars) and HuRlo (black bars) HT22 cells after treatment with 4 mM glutamate for 10 and 20 h. (c) Bar graphs depicting differences in selected mRNAs in control (white bars) and CN-KO (black bars) primary hippocampal neurons at DIV7 and after treatment with 100 μM glutamate for 4 h. For both (b) and (c), data are presented as fold change±S.D. relative to untreated control cells and are derived from qRT-PCR experiments using RNA from three individual cultures. *Increases and **decreases with P<0.05
Last, we focused on exclusive targets of HuR. HuR’s associations with Bcl2, NQO1 and Ppargc1α mRNAs increased in HT-22 neurons responding to glutamate (Figure 6a); conversely, HT-22 HuRlo neurons displayed reductions in the basal and inducible accumulation of Bcl2, NQO1 and Ppargc1a mRNAs (Figure 6b). CN-KO primary hippocampal neurons also displayed a significant reduction in the basal accumulation of these mRNAs, which were less evident in the presence of glutamate probably because of their enhanced execution (Figure 6c). These results demonstrate that HuR activates a central network for mitochondrial maintenance and resistance to oxidative damage. This also connects to the reported activities of PGC1 in the coactivation of the nuclear factor erythroid-2-related factor 2 (Nrf2)-mediated transcription of the NQO1 gene and its feedback for further PGC1 stabilization, and the coactivation of FOXO1 transcription that is reduced in HT-22 HuRlo cells. Interestingly, the expression of FOXO1 mRNA was also reduced in HT-22 HuRlo neurons – yet by indirect means. The estimation of mRNA’s half-lives in HT-22 HuRhi and HuRlo neurons pulsed with a transcriptional inhibitor (actinomycin D) (Figure 7a and Supplementary Figure 11) revealed that the loss of HuR enhanced the basal and/or inducible decay of Bcl2, NQO1 and Ppargc1a mRNAs providing explanations for the detected changes in mRNA accumulation. Moreover, the fractionation of these mRNAs as free, monosomal or polysomal in the same cells (Figure 7b and Supplementary Figure 12) revealed that glutamate increases the translation of NQO1 and Ppargc1a mRNAs but not of Bcl2. In that setting, the loss of HuR led to marked decrease in NQO1 mRNA translation and a partial impediment in the Bcl2 mRNA translation, indicating that HuR acts as their translational coactivator.
Effects of HuR’s loss on mRNA turnover and translation. (a) Estimation of mRNA decay in actinomycin D-treated HuRhi (white bars) and HuRlo (black bars) HT22 neurons before and after glutamate (4 mM) challenge. Data are half-lives±S.D. estimated from qRT-PCR measurements and decay plots from three independent experiments (see also Supplementary Figure 11). (b) Detection of changes in mRNA translation by means of qRT-PCR in free, monosomal and polysomal fractions from HuRhi (white bars) and HuRlo (black bars) before and after glutamate treatment. Data from measurements±S.D. in individual fractions normalized to GAPDH mRNA and presented as total free, monosomal or polysomal percentages of cytoplasmic RNA (see also Supplementary Figure 12). *Statistically different to control values; **statistically different to unstimulated fractions. In all cases P≤0.01
HuR could also repress the mRNAs encoding Myc, Cirbp and Gja1; their associations with HuR decreased under glutamate in HuRhi cells. The abundance of Myc and Cirbp mRNAs increased in stimulated HT-22 HuRlo neurons and unstimulated CN-KO primary hippocampal neurons, whereas Gja1 showed enhancements primarily in HT-22 HuRlo neurons (Figures 6b and c). These changes may relate to the effects of these mRNAs in the enhancement of stress and death responses when augmented. At the molecular level, and in HT-22 neurons, the loss of HuR inhibited the inducible decay of Cirbp and Gja1 mRNAs but not their translation; in contrast, the decay and translation of Myc mRNA remained unaltered, suggesting that its increase is because of changes in nuclear events (Figures 7a and b and Supplementary Figures 11 and 12).
Collectively, our data demonstrate that HuR activates a defense network of RNAs (i.e. a posttranscriptional regulon) to counteract oxidative metabolism, while limiting potentially hazardous gene expression, to protect neurons from degeneration and necrosis.
Discussion
In this report, we provide evidence for a specialized and indispensable role for the widely expressed HuR in the adult CNS. Our findings disprove original suppositions on its redundancy to nElavls and suggest that the members of the Elavl family possess distinct functions even among the pyramidal neurons of the hippocampus. The graded expression of HuR from the CA1 to CA3 regions of hippocampus is of particular interest. Both regions contain glutamatergic neurons of similar morphology; still their sensitivity to kainate varies with CA3 neurons (i.e. HuRlo) being more sensitive than CA1 neurons (HuRhi).47 Distinctions in kainate receptor subtype usage, as well as differences in gene expression, have been considered to contribute to this difference in sensitivity.48,49 In our mouse system, the induced genetic mutation rendered all neurons both as HuRlo and as sensitive to the degenerative effects of kainite, suggesting that HuR constitutes a determinant for sensitivity to neurodegeneration. This phenotype appears as exclusive to HuR as we did not observe any compensatory changes in nElavls. Further evidence supports a dichotomy of functions between HuR and nElavls in the adult CNS. For example, the loss of Elavl3/HuC in the mouse causes asymptomatic seizures because of a reduction in enzymes required for glutamate biosynthesis.10 As we demonstrate herein, HuR is involved downstream of this pathway to oppose a stress response to glutamate excess. Moreover, the physical comparison of HuR/mRNA targets (herein) to that of neuronal HuD46 suggest that cognitive targets in hippocampal neurons do not overlap. This is also apparent by their distinct nucleocytoplasmic response to glutamate where the localization of HuR is strongly altered but that of HuD is maintained. We do note, however, that our analysis is restricted to non-ionotropic signals; the fact that HuR and HuD comigrate strongly during status epilepticus and can recognize some common targets7 suggests that there may be a role for a HuR/HuD RNPs during ionotropic excitation. Indeed, our studies demonstrate a sex-restricted pathologic role for HuR in kainite-induced excitation. This effect could also relate to the regulation of steroid signals (e.g. estrogen and estradiol) or – as demonstrated herein – of protein kinase A signals by Elavls, which can alter neuronal excitability.50,51
Irrespective of the differences in excitation, the loss of HuR led to a profound exacerbation in neurodegeneration. A plethora of evidence has linked KA-induced neuronal death to glutathione reduction, lipid peroxidation, production of ROS, mitochondrial damage and oxidative stress.18,52, 53, 54, 55 In our study, these effects could not be fully linked to HuR functions in primary hippocampal cultures as they manifested a basal degenerative response – both in the presence or absence of antioxidants. Still, the response of immature primary cultures to kainate and glutamate and the partial rescue of the latter by antioxidants suggests that this may be the case. Clear evidence is herein provided in HT-22 neurons where glutamate incapacitates the built-up of a proper antioxidant defense. In these cells, HuR opposes the engagement of oxidation-induced ‘death cascades’. The connection between HuR and cell death has been obscured by controversy. A large body of work renders HuR as an ‘antiapoptotic’ factor, based on its cognition of pre-mRNAs/mRNAs encoding apoptotic regulators and the catastrophic effects of its inducible whole-body deletion in adult mice.56,57 In these cases, however, the specific type of death induced was not fully analyzed. Conversely, HuR can also act as proapoptotic molecule. Fas-induced apoptosis and caspase-3 activation is perturbed in HuR-knockout T cells.58 In response to lethal stress, caspase-3 and -7 cleave HuR to release interacting apoptosome effectors and amplify proapoptotic loops.59,60 Our current findings suggest for the first time that these differences may be because of the differential need for HuR in apoptosis versus necrosis, which includes several modes of programmed cell death (e.g. necroptosis, oxytosis, parthanatos). HT-22 cells are known to respond as caspase-3-deficient cells and thus die by programmed necrosis.28,30 The loss of HuR enhanced the necrotic program mediated by the Bid/Bax/Bad axis, mitochondrial dysfunction, oxidation and the release of DNA-damaging agents. This was in part because of a lack of prosurvival Bcl-2 both in HT-22 cells and primary neurons. Previous reports established the neuroprotective effect of Bcl-2 in neurons.61, 62, 63 HuR has also been linked to the activation of prosurvival Bcl-2 family members56,64,65 through the regulation of their turnover and translation,56,66, 67, 68 In HT-22 cells, HuR maintains Bcl2 mRNA stability and translation at a constant rate to buffer its requirement during assault and preserve mitochondrial homeostasis.
Our diagnostic platform revealed a HuR-regulated network connecting energy regulation to neurodegeneration through HuR’s activities upon PGC1a and NQO1 mRNAs in both HT-22 cells and primary neurons. PGC-1α is an inducible coactivator of several transcriptional programs relating to a energy expenditure, sensitivity to low temperature and – at the subcellular level – mitochondrial biogenesis. Fluctuations in the hypothalamic expression of PGC-1α relate to changes in body energy content, a fact supported also by the ‘lean’ condition of PGC1α-deficient mice. These mice also develop lesions in the striatum, substantia nigra and the hippocampus,69,70 and are sensitive to glutamatergic death induced by KA phenocopying the response of HuR-deficient neurons. PGC1α is activated by ROS and its augmented expression can protect from the oxidative neurodegeneration.69,71,72 The current mechanistic proposition is that PGC-1α interacts with transcription factors to cotranscribe neuronal genes involved in the regulation of mitochondrial density, oxidative metabolism and detoxification.73,74 Known interactions include Nrf2,73 which targets genes with antioxidant response elements in their promoters and counteracts KA neurotoxicity.75 One such gene is NQO1 that detoxifies quinones and protects cells against redox cycling and oxidative stress; its mutation or misexpression in humans correlate with cancer and numerous degenerative syndromes such as tardive dyskinesia (TD) and Alzeihemer’s disease.40 Based on the effects of HuR upon the NQO1 mRNA, we postulate that HuR ensures the proper activation of Nrf2 programs by targeting both its coactivation by PGC1α and the proper control of its transcribed genes. In that sense, it was surprising that HuRlo neurons did not display alterations to additional Nrf2 target genes; one explanation could be that the hyperactivity of Myc compensates for the loss of activation of such programs by PGC1α. On the other hand, the underrepresentation of genes transcribed by the metabolic transcription factor FOXO1 point towards a connection to PGC1 activity as it can physically interact with PGC1α76 and can also be involved in several facets of oxidative damage control.77
HuR affects also the mRNAs of pleiotropic factors whose excess in HT-22 cells and primary neurons may cause further damage or alter differentiation programs. For example, Myc can promote cell survival but its excess can lead to transformation or death.41 Similarly, Cirbp counteracts cold shock and hypoxia, and also can also enhance stress reactions as in the case of inflammation.42 Connexin-43 is required for cell polarity, migration and gap-junction formation, but its overexpression enhances gap-junctional communication and arrests developmental programs.78
The bimodal effects of HuR toward mRNA activation and suppression are suggestive of its involvement in different, tissue-restricted RNP configuration that may include other neuronal RBPs and regulatory RNAs. Exploiting the involvement of such HuR/RNPs in neurodegenerative circuitries can provide novel means for the diagnosis and therapeutic management of related neurological syndromes.
Materials and Methods
Animal models of neurodegeneration
The generation of Elavl1fl/fl, CamKIICre and ROSA26 mice has been described previously.14, 15, 16 Mice were bred in a C57BL/6J background and maintained in the animal facilities of the Biomedical Sciences Research Center ‘Alexander Fleming’ under specific pathogen-free conditions. For the detection of Cre-mediated recombination of the Elavl1 locus, HindIII-digested genomic DNA from tissues or biopsies was blotted onto membranes and analyzed via specific probes.14 The recombination of ROSA26 by CamKIICre was analyzed via the detection of the β-galactosidase activity.15 Experiments involving the administration of KA (intraperitoneal; K0250; Sigma, St. Louis, MO, USA) were supervised by Fleming’s Veterinarian. Seizure severity was quantified via a 5-pointed rating scale20 that included the frequency, latency and duration of shakes, convulsions, tonic–clonic seizures and death by two observers (OP and DLK). Surviving mice returned to their cages 3 h after the cessation of seizure activity or killed for histology. All protocols were approved by Fleming’s Ethical Committee and licensed by the Prefecture of Eastern Attica, Directorates General of Veterinary Services and Animal Health (Licenses Nos 3962B/26012009 and 1423/23062010).
Histologic analysis
For histology, paraffin-embedded brain sections were stained via standard procedures with hematoxylin and eosin (detection of hippocampal degeneration), and Nissl (detection of neuronal perikarya). Pathologic score was derived from H/E staining with the following scoring system: 0, no pathology; 0.5, low number of shrunk (dark neurons); 1, many shrunk (dark) neurons; 2, shrunk neurons and loss. Neuronal damage was derived from Nissl stains with the following scoring system: 0=no shrinkage; 1=5–15 neurons with shrinkage; 2=16–40 neurons with shrinkage; 3=>40 neurons with shrinkage; 4=light loss of neurons (5–10% neuronal loss); 5= moderate (10–40% neuronal loss); 6=severe (>40% neuronal loss).
Cell culture and transfections
For primary hippocampal cultures, CamKIICre+Elavl1fl/fl male mice were mated to female Elavl1fl/fl mice. On the 19th day of gestation, embryos were removed and sampled for the genotypic identification of the Cre gene via PCR. In parallel, individual embryos were processed for harvesting of hippocampi23 and preservation in warm Hibernate-E Solution (Invitrogen, Life Technologies, Carlsbad, CA, USA; A12476-01). Following genotyping, hippocampi from three embryos per culture were pooled, dissociated via trypsinization and trituration.23 Triturated cells were resuspended in Neurobasal medium (Invitrogen; 21103-049) supplemented with 0.5mM glutamine (Glutamax; 35050-061; Invitrogen), B27 with antioxidants (17504-044; Invitrogen) and penicillin/streptomycin (15140-148; Invitrogen). Cells were counted and plated onto 24-well (8 × 104 cells/well; with or without coverslips) or 6-well plates (5 × 105/well) coated with a solution of three parts rat collagen I (354236; BD Biosciences, San Jose, CA, USA) to one part poly-DL-lysine (P6407; Sigma). During DIV2–4, the medium was supplemented with 5 μM cytosine arabinoside (AraC; S1648; Selleckchem, Munich, Germany) to block glial differentiation. The medium was half changed at DIV4, DIV6 and DIV9 and cultures were terminated at DIV10 because of excessive degeneration in test cultures. For toxicity experiments, the medium was changed in DIV6 neurons to Neurobasal medium supplemented with B27 minus antioxidants (35050-061; Invitrogen) and experiments were performed 24 h later. HT-22 murine hippocampal neuronal cells were maintained in complete DMEM (Biochrom, Berlin, Germany) supplemented with 10% FBS, 2 mmol/ml L-glutamine and 100 μg/ml penicillin/streptomycin (Gibco-Invitrogen, Life Technologies) in 5% CO2 at 37 °C. ShRNA plasmids were constructed upon a pSilencer-U6 carrier containing a neomycin resistance selection cassette. For HuR, hairpins showing maximal efficacy targeted a sequence located 158 bases after the ATG of the murine Elavl1 transcript (ENSMUST00000098950: 5′-GGGATAAAGTAGCAGGACACA-3′); the scramble sequence (5′-GAAGGACCAGGAACGGTAATA-3′) was designed using the siRNAWizard algorithm. Plasmids were transfected with Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. For stable transfectants, cells were selected and maintained in medium supplemented with G418 (Invitrogen).
Cell viability assays
For assessment of cell viability in primary hippocampal neurons, cultures were supplemented at the indicated time points with PrestoBlue Cell Viability Reagent (Invitrogen), incubated for 1 h and then fluorescence was measured according to the manufacturer’s instructions on an InfiniteM200 fluorometer (Tecan, Männedorf, Switzerland). For the viability of HT-22 neurons, cells were seeded in 96-well plates at a density of 5000 cells/well and treated with different concentrations of glutamate for 24 h. Antioxidants or signaling modifiers were added 2 h before glutamate challenge and included NAc and Nec-1 from Sigma; AP5 (NMDA antagonist), CNQX disodium salt (AMPA antagonist), BI6C9, Trolox and PD146176 from Santa Cruz (Dallas, TX, USA); zVAD from R&D (Abingdon, UK); and ABT-737 from Selleckchem. Following treatment, cells were washed with serum-free culture medium and incubated with MTT solution (500 μg/ml) for 2 h at 37 °C; subsequently, the solution was removed and plates were frozen for 2–2 h. Plates were developed with dimethyl sulfoxide (DMSO) and read at 550 nm using a Optimax plate reader (Molecular Devices, Sunnyvale, CA, USA). Unless otherwise indicated, raw values were converted to percentages of untreated control values.
Immunohistochemistry and immunoblotting
For immunohistochemistry, slides were and deparaffinized, microwave treated for 20 min in epitope-unmasking buffer (10 mM sodium citrate, 0.05% Tween-20, pH 6.0) and used for primary antibody staining. Detection was performed with horseradish peroxidase- or alkaline phosphatase-conjugated secondary IgG’s (Vector, Burlingame, CA, USA; Southern Biotechnologies, Birmingham, AL, USA) and visualized with DAB or NBT/BCIP, respectively (Vector). For western blots, whole-cell lysates from dissected brain regions or cells were prepared in RIPA buffer (25 mM Tris, 150 mM NaCl, 0.1% SDS, 1% NP-40, pH 7.4); nuclear and cytoplasmic extracts were isolated with NE-PER reagents according to the manufacturer's protocols (Pierce, Thermo Scientific, Rockford, IL, USA). Lysates where quantified with a BCA Protein Assay Kit (Thermo Scientific). Equimolar amounts of protein (10–50 μg) were analyzed on sodium dodecyl sulfate-containing polyacrylamide gels (SDS-PAGE; 10–12%) along with protein molecular weight markers (SM0431; Nippon MWP03; Fermentas, Thermo Scientific) and blotted onto nitrocellulose membranes (GE Healthcare, Freiburg, Germany). After blocking with 5% milk membranes, they were sequentially probed with primary antibodies; detection was performed via horseradish peroxidase-conjugated secondary antibodies (Southern Biotechnologies) and visualized by enhanced chemiluminescence (ECL+ or ECL prime; Amersham, GE Healthcare) using films or a STORM Phosphor-/FluorImager device (Molecular Dynamics, GE Healthcare). Primary antibodies included those against: all Elavls (3A2), HuD (E-1), PGC-1 (H-300), Bcl-xl (S18), actin (C11) and laminin A/C (N-18) from Santa Cruz; AIF (4642), BID (2003), Bax (2772), Bad (9292), phospho-Bad (9296), Bcl-2 (2876), p53 (1C12) and PARP (9542) from Cell Signaling Technology (Danvers, MA, USA); HuR (19F12) from Upstate Biotech (Merck-Millipore, Billerica, MA, USA); anti-F1F0-ATP synthase subunit α (7H10) from Molecular Probes (Life Technologies); and GAPDH (glyceraldehyde-3-phosphate dehydrogenase) (6C5) from Ambion (Life Technologies).
Mitochondrial activity assays and flow cytometry
For the detection of cell death, cells were stained with PI (Sigma) and AnxV FITC (eBioscience, San Diego, CA, USA). For the detection of changes in the mitochondrial membrane potential, cells were stained with 25 nM of DiOC6(3) (Molecular Probes) for 15 min at 37 °C. For the detection of superoxide formation, cells were stained with MitoSOX red (Molecular Probes) according to the manufacturer’s instructions. In all cases, signals were detected with a FACS Canto II flow cytometer (BD Biosciences), acquired with FACSDiva software (V6; BD Biosciences) and analyzed using FlowJo (Tree Star, Ashland, OR, USA). Intracellular ROS was measured using the DCFDA Cellular ROS Detection Assay Kit (Abcam, Cambridge, UK) and protocols provided therein; microplate measurements were performed on an InfiniteM200 fluorometer (Tecan).
RNA analysis and RNP IPs
Total RNA was extracted from HT-22 neurons using Trizol reagent (Invitrogen); for microarrays, RNAs were further purified by affinity columns (Qiagen). For mRNA decay, cells were pulsed with actinomycin D (5 μg/ml; Sigma) and RNAs were collected at the indicated time points. For polysome analysis, cytoplasmic fractions containing monosomes and polysomes were isolated as described previously.14 For cDNA synthesis, 0.5–5 μg of total RNA was reverse transcribed with MMLV-RT (Promega, Madison, WI, USA). qRT-PCR was performed using EvaGreen SsoFast mix (Bio-Rad, Hercules, CA, USA) on a RotorGene 6000 machine (Corbett Research, Qiagen, Venlo, Netherlands). Expression was normalized to β2-microglobulin and/or GAPDH mRNAs. The relative mRNA expression in the test samples was calculated as the difference from the control values that were assigned an arbitrary expression value of 1, using Bio-Rad RelQuant (Bio-Rad). mRNA half-lives were calculated by setting the normalized amount of target mRNA in unstimulated samples at 100 and extrapolating from the corresponding semilogarithmic plots of percentages versus time. Ribonucleoprotein IP experiments were performed as described previously.34 Briefly cells were lysed in buffer containing 100 mM KCl, 25 mM EDTA, 5 mM MgCl2, 10 mM HEPES, 0.5% Nonidet P40, 2 mM DTT, 0.2% vanadyl ribonucleoside complex (Invitrogen) and 100 U/ml RNAse OUT (Invitrogen). Antibody-coated beads (agarose or protein A sepharose) were washed and maintained in 750 μl of NT-2 buffer (50 mM Tris, pH 7.4, 150 mM NaCl, 1 mM MgCl2 and 0.05% Nonidet P40). For IP, 200 μl of lysate was loaded onto the beads and incubated for 4 h overnight on a rotary at 4 °C. Subsequently, beads were washed five times in NT-2 buffer plus one time in NT-2+1M urea (for αElavl, and αHuR) or two times in NT-2+1M urea (for α-HuD) and finally resuspended in NT-2. Ten microliters of samples was removed for immunoblot verification and the remaining were used for RNA isolation via proteinase K treatment, phenol/chloroform extraction and precipitation. Antibodies included agarose-conjugated anti-Elavl (3A2; Santa Cruz); unmodified anti-HuR (4C8; Abcam), anti-HuD (E-1; Santa Cruz); and mouse immunoglobulin antibodies (Santa Cruz and R&D). RNA was extracted from the IP material to perform the qRT-PCR. Target mRNAs were identified by the >5-fold enrichment of a transcript in test IPs relative to control IPs, normalizing to the low-level contaminating transcript of GAPDH. Methods relating to microarray hybridizations, bioinformatic analyses and the list of PCR primers used in this study are presented in Supplementary Methods.
Statistics
Unless otherwise indicated, Student’s t-test was used for statistical analysis. Results with a P-value <0.05 considered to be statistically significant
Abbreviations
- ALS:
-
amyotrophic lateral sclerosis
- AMPA:
-
α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid
- AP5:
-
DL-2-amino-5-phosphonovaleric acid
- CA:
-
cornus ammonis
- CamKIIa:
-
calcium/calmodulin-dependent protein kinase II-α
- CNQX:
-
6-cyano-7-nitroquinoxaline-2,3-dione
- CNS:
-
central nervous system
- DCFDA:
-
2'7'-dichlorofluorescein diacetate
- DG:
-
dentate gyrus
- DIV:
-
day in vitro
- ELAVL/Hu:
-
embryonic lethal abnormal vision-like human antigen
- GluR:
-
glutamate receptor
- HuR:
-
human antigen R
- KA:
-
kainic acid
- MTT:
-
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide
- NAc:
-
N-acetylcysteine
- NMDA:
-
N-methyl-D-aspartate
- PAR-CliP:
-
photoactivatable-ribonucleoside-enhanced crosslinking and immunoprecipitation
- PARP:
-
poly ADP ribose polymerase
- PGC1α:
-
peroxisome proliferator-activated receptor gamma coactivator 1-alpha
- PTR:
-
posttranscriptional regulation
- RBPs:
-
RNA-binding proteins
- ROS:
-
reactive oxygen species
References
Wang Y, Qin ZH . Molecular and cellular mechanisms of excitotoxic neuronal death. Apoptosis 2010; 15: 1382–1402.
Danbolt N . Glutamate uptake. Progr Neurobiol 2001; 65: 1–105.
Michaelis E . Molecular biology of glutamate receptors in the central nervous system and their role in excitotoxicity, oxidative stress and aging. Progr Neurobiol 1998; 54: 369–415.
Darnell R . RNA protein interaction in neurons. Annu Rev Neurosci 2013; 36: 243–270.
McKee A, Minet E, Stern C, Riahi S, Stiles CD, Silver PA et al. A genome-wide in situ hybridization map of RNA-binding proteins reveals anatomically restricted expression in the developing mouse brain. BMC Dev Biol 2005; 5: 1–9.
Okano H, Darnell R . A hierarchy of Hu RNA binding proteins in developing and adult neurons. J Neurosci 1997; 17: 3024–3037.
Tiruchinapalli D, Caron M, Keene J . Activity-dependent expression of ELAV/Hu RBPs and neuronal mRNAs in seizure and cocaine brain. J Neurochem 2008; 107: 1529–1543.
Akamatsu W, Okano HJ, Osumi N, Inoue T, Nakamura S, Sakakibara S, Miura M et al. Mammalian ELAV-like neuronal RNA-binding proteins HuB and HuC promote neuronal development in both the central and the peripheral nervous systems. Proc Natl Acad Sci USA 1999; 96: 9885–9890.
Akamatsu W, Fujihara H, Mitsuhashi T, Yano M, Shibata S, Hayakawa Y et al. The RNA-binding protein HuD regulates neuronal cell identity and maturation. Proc Natl Acad Sci USA 2005; 102: 4625–4630.
Ince-Dunn G, Okano HJ, Jensen KB, Park WY, Zhong R, Ule J et al. Neuronal Elav-like (Hu) proteins regulate RNA splicing and abundance to control glutamate levels and neuronal excitability. Neuron 2012; 75: 1–14.
Garcia-Dominguez D, Morello D, Cisneros E, Kontoyiannis D, Frade J . Stabilization of Dll1 mRNA by Elavl1/HuR in neuroepithelial cells undergoing mitosis. Mol Biol Cell 2011; 22: 1227–1239.
Mansfield K, Keene J . Neuron-specific ELAV/Hu proteins suppress HuR mRNA during neuronal differentiation by alternative polyadenylation. Nucleic Acids Res 2012; 40: 2734–2746.
Tiruchinapalli DM, Ehlers MD, Keene JD . Activity-dependent expression of RNA binding protein HuD and its association with mRNAs in neurons. RNA Biol 2008; 5: 157–168.
Katsanou V, Milatos S, Yiakouvaki A, Sgantzis N, Kotsoni A, Alexiou M et al. The RNA-binding protein Elavl1/HuR is essential for placental branching morphogenesis and embryonic development. Mol Cell Biol 2009; 29: 2762–2776.
Mantamadiotis T, Lemberger T, Bleckmann SC, Kern H, Kretz O, Martin Villalba A et al. Disruption of CREB function in brain leads to neurodegeneration. Nat Genet 2002; 31: 47–54.
Soriano P . Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat Genet 1999; 21: 70–71.
Ben-Ari Y, Cossart R . Kainate, a double agent that generates seizures: two decades of progress. Trends Neurosci 2000; 23: 580–587.
Wang Q, Yu S, Simonyi A, Sun G, Sun A . Kainic acid-mediated excitotoxicity as a model for neurodegeneration. Mol Neurobiol 2005; 31: 3–16.
McLin J, Steward O . Comparison of seizure phenotype and neurodegeneration induced by systemic kainic acid in inbred, outbred, and hybrid mouse strains. Eur J Neurosci 2006; 24: 2191–2202.
Royle S, Collins F, Rupniak H, Barnes J, Anderson R . Behavioural analysis and susceptibility to CNS injury of four inbred strains of mice. Brain Res 1999; 816: 337–349.
Mckhann G . Mouse strain differences in kainic acid sensitivity, seizure behavior, mortality, and hippocampal pathology. Neuroscience 2003; 122: 551–561.
Kaech S, Banker G . Culturing hippocampal neurons. Nat Protoc 2006; 1: 2406–2415.
Seibenhener ML, Wooten MW . Isolation and culture of hippocampal neurons from prenatal mice. J Vis Exp 2012; 3634.
Schubert D, Piasecki D . Oxidative glutamate toxicity can be a component of the excitotoxicity cascade. J Neurosci 2001; 21: 7455–7462.
Hilton GD, Nunez JL, Bambrick L, Thompson SM, McCarthy MM . Glutamate-mediated excitotoxicity in neonatal hippocampal neurons is mediated by mGluR-induced release of Ca++ from intracellular stores and is prevented by estradiol. Eur J Neurosci 2006; 24: 3008–3016.
Albrecht P, Lewerenz J, Dittmer S, Noack R, Maher P, Methner A et al. Mechanisms of oxidative glutamate toxicity: the glutamate/cystine antiporter system xc− as a neuroprotective drug target. CNS Neurol Disord Drug Targets 2010; 9: 373–382.
Fukui M, Song JH, Choi J, Choi H, Zhu B . Mechanism of glutamate-induced neurotoxicity in HT22 mouse hippocampal cells. Eur J Pharmacol 2009; 617: 1–11.
Tobaben S, Grohm J, Seiler A, Conrad M, Plesnila N, Culmsee C et al. Bid-mediated mitochondrial damage is a key mechanism in glutamate-induced oxidative stress and AIF-dependent cell death in immortalized HT-22 hippocampal neurons. Cell Death Differ 2010; 18: 282–292.
Xu X, Chua CC, Kong J, Kostrzewa RM, Kumaraguru U, Hamdy RC et al. Necrostatin-1 protects against glutamate-induced glutathione depletion and caspase-independent cell death in HT-22 cells. J Neurochem 2007; 103: 2004–2014.
Zhang Y, Bhavnani B . Glutamate-induced apoptosis in neuronal cells is mediated via caspase-dependent and independent mechanisms involving calpain and caspase-3 proteases as well as apoptosis inducing factor (AIF) and this process is inhibited by equine estrogens. BMC Neurosci 2006; 7: 49.
Luo Y, DeFranco D . Opposing roles for ERK1/2 in neuronal oxidative toxicity - Distinct mechanisms of ERK1/2 action at early versus late phases of oxidative stress. J Biol Chem 2006; 281: 16436–16442.
Young K, Piñón LG, Dhiraj D, Twiddy D, Macfarlane M, Hickman J et al. Mitochondrial fragmentation and neuronal cell death in response to the Bcl-2/Bcl-x(L)/Bcl-w antagonist ABT-737. Neuropharmacology 2010; 58: 1258–1267.
Halligan B, Ruotti V, Jin W, Laffoon S, Twigger SN, Dratz EA et al. ProMoST (Protein Modification Screening Tool): a web-based tool for mapping protein modifications on two-dimensional gels. Nucleic Acids Res 2004; 32: W638–W644.
Tenenbaum SA, Lager PJ, Carson CC, Keene JD . Ribonomics: identifying mRNA subsets in mRNP complexes using antibodies to RNA-binding proteins and genomic arrays. Methods 2002; 26: 191–198.
Dai W, Zhang G, Makeyev E . RNA-binding protein HuR autoregulates its expression by promoting alternative polyadenylation site usage. Nucleic Acids Res 2012; 40: 787–800.
Mukherjee N, Corcoran DL, Nusbaum JD, Reid DW, Georgiev S, Hafner M et al. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol Cell 2011; 43: 327–339.
Lebedeva S, Jens M, Theil K, Schwanhäusser B, Selbach M, Landthaler M et al. Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol Cell 2011; 43: 340–352.
Anders G, Mackowiak SD, Jens M, Maaskola J, Kuntzagk A, Rajewsky N et al. doRiNA: a database of RNA interactions in post-transcriptional regulation. Nucleic Acids Res 2012; 40: D180–D186.
Chen SD, Yang DI, Lin TK, Shaw FZ, Liou CW, Chuang YC et al. Roles of Oxidative Stress, Apoptosis, PGC-1+- and mitochondrial biogenesis in cerebral ischemia. Int J Mol Sci 2011; 12: 7199–7215.
Dinkova-Kostova A, Talalay P . NAD(P)H:quinone acceptor oxidoreductase 1 (NQO1), a multifunctional antioxidant enzyme and exceptionally versatile cytoprotector. Arch Biochem Biophys. 2010; 501: 116–123.
Phesse TJ, Myant KB, Cole AM, Ridgway RA, Pearson H, Muncan V et al. Endogenous c-Myc is essential for p53-induced apoptosis in response to DNA damage in vivo. Cell Death Differ 2014; 21: 956–966.
Qiang X, Yang WL, Wu R, Zhou M, Jacob A, Dong W et al. Cold-inducible RNA-binding protein (CIRP) triggers inflammatory responses in hemorrhagic shock and sepsis. Nat Med 2013; 19: 1489–1495.
Liu X, Sun L, Torii M, Rakic P . Connexin 43 controls the multipolar phase of neuronal migration to the cerebral cortex. Proc Natl Acad Sci USA 2012; 109: 8280–8285.
Waltereit R, Weller M . Signaling from cAMP/PKA to MAPK and synaptic plasticity. Mol Neurobiol 2003; 27: 99–106.
Izquierdo JM . Control of the ATP synthase beta subunit expression by RNA-binding proteins TIA-1, TIAR, and HuR. Biochem Biophys Res Commun 2006; 348: 703–711.
Bolognani F, Perrone-Bizzozero N . RNA−protein interactions and control of mRNA stability in neurons. J Neurosci Res 2008; 86: 481–489.
Westbrook G, Lothman E . Cellular and synaptic basis of kainic acid-induced hippocampal epileptiform activity. Brain Res 1983; 273: 97–109.
Bureau I, Bischoff S, Heinemann S, Mulle C . Kainate receptor-mediated responses in the CA1 field of wild-type and GluR6-deficient mice. J Neurosci 1999; 19: 653–663.
Newrzella D, Pahlavan PS, Krüger C, Boehm C, Sorgenfrei O, Schröck H et al. The functional genome of CA1 and CA3 neurons under native conditions and in response to ischemia. BMC Genom 2007; 8: 370.
Pryzbylkowski P, Obajimi O, Keen J . Trichostatin A and 5 aza-2' deoxycytidine decrease estrogen receptor mRNA stability in ER positive MCF7 cells through modulation of HuR. Breast Cancer Res Treat 2008; 111: 15–25.
Rudick C, Woolley C . Estrogen regulates functional inhibition of hippocampal CA1 pyramidal cells in the adult female rat. J Neurosci 2001; 21: 6532–6543.
Liang L, Ho Y, Patel M . Mitochondrial superoxide production in kainate-induced hippocampal damage. Neuroscience 2000; 101: 563–570.
Patel M . Mitochondrial dysfunction and oxidative stress: cause and consequence of epileptic seizures. Free Rad Biol Med 2004; 37: 1951–1962.
Shin EJ, Jeong JH, Bing G, Park ES, Chae JS, Yen TP et al. Kainate-induced mitochondrial oxidative stress contributes to hippocampal degeneration in senescence-accelerated mice. Cell Signal 2008; 20: 645–658.
Shin EJ, Ko KH, Kim WK, Chae JS, Yen TP, Kim HJ et al. Role of glutathione peroxidase in the ontogeny of hippocampal oxidative stress and kainate seizure sensitivity in the genetically epilepsy-prone rats. Neurochem Int 2008; 52: 1134–1147.
Abdelmohsen K, Lal A, Kim H, Gorospe M . Posttranscriptional orchestration of an anti-apoptotic program by HuR. Cell Cycle (Georgetown, TX) 2007; 6: 1288–1292.
Ghosh M, Aguila HL, Michaud J, Ai Y, Wu MT, Hemmes A et al. Essential role of the RNA-binding protein HuR in progenitor cell survival in mice. J Clin Invest 2009; 119: 3530–3543.
Papadaki O, Milatos S, Grammenoudi S, Mukherjee N, Keene JD, Kontoyiannis DL et al. Control of thymic T cell maturation, deletion and egress by the RNA-binding protein HuR. J Immunol 2009; 182: 6779–6788.
Roretz C, Lian XJ, Macri AM, Punjani N, Clair E, Drouin O et al. Apoptotic-induced cleavage shifts HuR from being a promoter of survival to an activator of caspase-mediated apoptosis. Cell Death Differ 2013; 20: 154–168.
Roretz C, Gallouzi IE . Protein kinase RNA/FADD/caspase-8 pathway mediates the proapoptotic activity of the RNA-binding protein human antigen R (HuR). J Biol Chem 2010; 285: 16806–16813.
Farlie P, Dringen R, Rees S, Kannourakis G, Bernard O . Bcl-2 transgene expression can protect neurons against developmental and induced cell death. Proc Natl Acad Sci USA 1995; 92: 4397–4401.
Kitagawa K, Matsumoto M, Tsujimoto Y, Ohtsuki T, Kuwabara K, Matsushita K et al. Amelioration of hippocampal neuronal damage after global ischemia by neuronal overexpression of BCL-2 in transgenic mice. Stroke 1998; 29: 2616–2621.
Sahin M, Saxena A, Joost P, Lewerenz J, Methner A . Induction of Bcl-2 by functional regulation of G-protein coupled receptors protects from oxidative glutamate toxicity by increasing glutathione. Free radical research 2006; 40: 1113–1123.
Durie D, Hatzoglou M, Chakraborty P, Holcik M . HuR controls mitochondrial morphology through the regulation of BclxL translation. Translation 2013; 1: e23980.
Filippova N, Yang X, Wang Y, Gillespie GY, Langford C, King PH et al. The RNA-binding protein HuR promotes glioma growth and treatment resistance. Mol Cancer Res 2011; 9: 648–659.
Donnini M, Lapucci A, Papucci L, Witort E, Jacquier A, Brewer G et al. Identification of TINO: a new evolutionarily conserved BCL-2 AU-rich element RNA-binding protein. J Biol Chem 2004; 279: 20154–20166.
Ishimaru D, Ramalingam S, Sengupta TK, Bandyopadhyay S, Dellis S, Tholanikunnel BG et al. Regulation of Bcl-2 expression by HuR in HL60 leukemia cells and A431 carcinoma cells. Mol Cancer Res 2009; 7: 1354–1366.
Marash L, Liberman N, Henis-Korenblit S, Sivan G, Reem E, Elroy-Stein O et al. DAP5 promotes cap-independent translation of Bcl-2 and CDK1 to facilitate cell survival during mitosis. Mol Cell 2008; 30: 447–459.
St-Pierre J, Drori S, Uldry M, Silvaggi JM, Rhee J, Jäger S et al. Suppression of reactive oxygen species and neurodegeneration by the PGC-1 transcriptional coactivators. Cell 2006; 127: 397–408.
Lin J, Wu PH, Tarr PT, Lindenberg KS, St-Pierre J, Zhang CY et al. Defects in adaptive energy metabolism with CNS-linked hyperactivity in PGC-1alpha null mice. Cell 2004; 119: 121–135.
Shin JH, Ko HS, Kang H, Lee Y, Lee YI, Pletinkova O et al. PARIS (ZNF746) repression of PGC-1+- contributes to neurodegeneration in Parkinson's disease. Cell 2011; 144: 689–702.
Thau N, Knippenberg S, Körner S, Rath KJ, Dengler R, Petri S et al. Decreased mRNA expression of PGC-1+- and PGC-1+-regulated factors in the SOD1G93A ALS mouse model and in human sporadic ALS. J Neuropathol Exp Neurol 2012; 71: 1064–1074.
Ventura-Clapier R, Garnier A, Veksler V . Transcriptional control of mitochondrial biogenesis: the central role of PGC-1alpha. Cardiovasc Res 2008; 79: 208–217.
Wareski P, Vaarmann A, Choubey V, Safiulina D, Liiv J, Kuum M et al. PGC-1{alpha} and PGC-1{beta} regulate mitochondrial density in neurons. J Biol Chem 2009; 284: 21379–21385.
Kraft AD, Lee JM, Johnson DA, Kan YW, Johnson JA . Neuronal sensitivity to kainic acid is dependent on the Nrf2-mediated actions of the antioxidant response element. J Neurochem 2006; 98: 1852–1865.
Puigserver P, Rhee J, Donovan J, Walkey CJ, Yoon JC, Oriente F et al. Insulin-regulated hepatic gluconeogenesis through FOXO1–PGC-1alpha interaction. Nature 2003; 423: 550–555.
Wang K, Zhang T, Dong Q, Nice EC, Huang C, Wei Y et al. Redox homeostasis: the linchpin in stem cell self-renewal and differentiation. Cell Death Dis 2013; 4: e537.
Ewart JL, Cohen MF, Meyer RA, Huang GY, Wessels A, Gourdie RG et al. Heart and neural tube defects in transgenic mice overexpressing the Cx43 gap junction gene. Development 1997; 124: 1281–1292.
Acknowledgements
In vivo experiments were performed by OP and DLK; histologic evaluation by JB; experiments with HT-22 cells were performed by AS, assisted by JK, SG and DLK; genomics experiments performed by AS and YK; bioinformatics by OH and MR; and manuscript written by AS and DLK. We thank Gunther Schutz for CamKIICre mice; Vaggelis Harokopos for microarray hybridizations; Anastasia Kotsoni, Margarita Andreadou, Nikos Natsaridis, Niki Lourou and Vasso Kostourou for technical assistance; and the InfrafrontierGR Infrastructure for providing animal housing, transgenic and genomic services. InfrafrontierGR is co-financed by the ERDF and NSRF 2007-2013 (MIS 423575). This work was supported in part by the Hellenic General Secreteriat of Research and Technology Grants ARISTEIA-I No. 1096 PREcISE and PENED-03EΔ394; and the European Commission Grant FP7-PEOPLE-2010-IEF, No. 274837.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest
Additional information
Edited by N Bazan
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Skliris, A., Papadaki, O., Kafasla, P. et al. Neuroprotection requires the functions of the RNA-binding protein HuR. Cell Death Differ 22, 703–718 (2015). https://doi.org/10.1038/cdd.2014.158
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2014.158
This article is cited by
-
LncRNA NKILA Exacerbates Alzheimer’s Disease Progression by Regulating the FOXA1-Mediated Transcription of TNFAIP1
Neurochemical Research (2023)
-
ELAVL1 is transcriptionally activated by FOXC1 and promotes ferroptosis in myocardial ischemia/reperfusion injury by regulating autophagy
Molecular Medicine (2021)
-
The RNA-binding protein HuR is a negative regulator in adipogenesis
Nature Communications (2020)
-
Integrated identification of key genes and pathways in Alzheimer’s disease via comprehensive bioinformatical analyses
Hereditas (2019)
-
Depletion of HuR in murine skeletal muscle enhances exercise endurance and prevents cancer-induced muscle atrophy
Nature Communications (2019)