Abstract
Neural crest cells (NCCs) migrate from different regions along the anterior–posterior axis of the neural tube (NT) to form different structures. Defective NCC development causes congenital neurocristopathies affecting multiple NCC-derived tissues in human. Perturbed Hoxb5 signaling in vagal NCC causes enteric nervous system (ENS) defects. This study aims to further investigate if perturbed Hoxb5 signaling in trunk NCC contributes to defects of other NCC-derived tissues besides the ENS. We perturbed Hoxb5 signaling in NCC from the entire NT, and investigated its impact in the development of tissues derived from these cells in mice. Perturbation of Hoxb5 signaling in these NCC resulted in Sox9 downregulation, NCC apoptosis, hypoplastic sympathetic and dorsal root ganglia, hypopigmentation and ENS defects. Mutant mice with NCC-specific Sox9 deletion also displayed some of these phenotypes. In vitro and in vivo assays indicated that the Sox9 promoter was bound and trans-activated by Hoxb5. In ovo studies further revealed that Sox9 alleviated apoptosis induced by perturbed Hoxb5 signaling, and Hoxb5 induced ectopic Sox9 expression in chick NT. This study demonstrates that Hoxb5 regulates Sox9 expression in NCC and disruption of this signaling causes Sox9 downregulation, NCC apoptosis and multiple NCC-developmental defects. Phenotypes such as ENS deficiency, hypopigmentation and some of the neurological defects are reported in patients with Hirschsprung disease (HSCR). Whether dysregulation of Hoxb5 signaling and early depletion of NCC contribute to ENS defect and other neurocristopathies in HSCR patients deserves further investigation.
Similar content being viewed by others
Main
Neural crest cells (NCCs) are generated from the dorsal neural tube (NT), migrate to the periphery and give rise to diverse cell lineages in many tissues according to the anterior–posterior (A–P) level of the NT from which they originate.1, 2, 3 Cranial NCCs differentiate into teeth, bone, cartilage and connective tissue in the head; vagal NCCs contribute to the enteric nervous system (ENS) and cardiac outflow tracts; trunk NCCs give rise to sympathetic ganglia and norepinephrine-producing cells in the adrenal gland; sacral NCC contributes to the ENS of distal gut.4, 5, 6, 7, 8, 9 Skin pigment cells are derived from NCC from all A–P levels. Defective NCC development causes neurocristopathies.
Hox genes encode transcription factors with a DNA-binding homeodomain that recognizes a specific sequence and thereby mediates transcriptional regulation of target genes in numerous developmental processes. In mammals, 39 Hox genes are separated into four clusters Hoxa, Hoxb, Hoxc and Hoxd, on four different chromosomes. Hox genes are subdivided into 13 paralogous groups, and these Hox genes are expressed in overlapping domains with spatially staggered anterior expression boundaries along the NT.10, 11, 12, 13 The different combinations of Hox genes expressed in these domains are mirrored in the NCC derived from those domains.14, 15, 16 So far, our knowledge regarding the role of Hox genes during NCC development comes mostly from loss-of-function experiments, however, the overlapping expression and extensive functional redundancy among Hox genes preclude detailed investigations of the developmental functions of individual Hox genes in this way.
Among the Hox genes expressed in the developing gut, Hoxb5 expression pattern is intimately associated with vagal NCC and ENS development.17, 18, 19, 20, 21, 22, 23, 24 Deletion of Hoxb5 caused a rostral shift of the shoulder girdle in mice, implying a patterning role of Hoxb5 in specifying the position of limbs on the body axis.25 However, no abnormal development was observed in NCC or other tissues that express Hoxb5, probably due to functional redundancy among paralogous Hox members with overlapping expression domains.
To circumvent the problem of functional redundancy and investigate the function of Hoxb5 in NCC, we generated transgenic mice that express a dominant-negative chimeric protein, engrailed-Hoxb5 (enb5), upon Cre-induction. This enb5 repressor competes with Hoxb5 for binding to target genes, thereby disrupting the developmental pathways that require Hoxb5. Using these mice, we previously showed that blocking Hoxb5 signaling in vagal NCC causes reduced Ret expression, retarded NCC migration and ENS defects,22 indicating that Hoxb5 regulates vagal NCC and ENS development.
Hoxb5 is also expressed in trunk NCC. To investigate whether Hoxb5 regulates trunk NCC development, in this study we crossed enb5 mice with Wnt1-Cre mice to induce enb5 expression in NCC from the full length of the NT, and looked for NCC abnormalities. We showed that Hoxb5 regulates the expression of Sox9 in trunk NCC, and that perturbation of Hoxb5 signaling in NCC causes downregulation of Sox9, apoptosis of NCC and neurocristopathies in mice.
Results
Hoxb5 perturbation in NCC causes developmental defects
To investigate the function of Hoxb5 in NCC derived from the entire A–P axis of the NT, we crossed enb5 mice with Wnt1-Cre mice.26 This induced the expression of enb5 protein, perturbing Hoxb5 function in NCC derived from all along the NT. We then studied the effects on the development of NCC and NCC-derived structures.
To trace NCC in Wnt1-Cre/enb5 mice, we used Rosa26R (R26R) Cre activity-reporter mice in our crosses. At E9.0, beta-galactosidase (X-gal)-positive cells were located in the midbrain, frontal-nasal region, first and second branchial arches in both Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 (Figure 1a). By E9.5, X-gal-positive cells were also detected in the hindbrain, third and fourth branchial arches, circumpharyngeal ridge, frontal-nasal process and the entire NT. As the embryos developed further, more X-gal-positive cells were found populating the midbrain, hindbrain, NT and NCC-derived structures including the branchial arches, dorsal root ganglion (DRG), sympathetic ganglion and the frontal-nasal region at E10.5. X-gal-positive NCC populated similar regions in Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 embryos; however, the intensity of X-gal staining in the NT, DRG and sympathetic ganglion was generally weaker in the Wnt1-Cre/R26R/enb5 embryos, indicating that fewer X-gal-positive cells were present.
Reduction of LacZ-expressing cells and shrinkage of DRG in Wnt1-Cre/enb5 mice. (a) LacZ-expressing cells (stained blue) in Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 embryos (E9.0–E10.5) were localized by whole-mount X-gal staining for β-galactosidase. (b) X-gal-stained E12.5 embryos of Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 embryos were sectioned to reveal the spatial distribution of LacZ-expressing cells. Boxed regions were magnified and shown on the right. Arrowheads indicated the LacZ-expressing cells at the dorsal NT. DRG was demarcated with broken line. Expression of Sox10 (purple; c) and Islet1/2 (purple; d) in enb5 and Wnt1-Cre/enb5 embryos was analyzed by whole-mount in situ hybridization. (e) Expression of Islet1/2 (green) on sections of E11.5 enb5 and Wnt1-Cre/enb5 embryos was analyzed by immunofluorescence. Dorsal root ganglion was demarcated with broken line. (f) Average total volume of DRG (mean±S.D.) from the trunk level of E11.5 enb5 and Wnt1-Cre/enb5 embryos was determined and compared. Volume of DRGs in enb5 embryo was taken arbitrarily as 100%. Average % of Islet1/2 immuno-positive neurons versus total number of cells (mean±S.D.) in DRG of E11.5 enb5 and Wnt1-Cre/enb5 embryos was determined and compared. Number of embryos analyzed was indicated by ‘n’. ba, branchial arch; cr, circumpharyngeal ridge; drg, dorsal root ganglion; fa, facio-acoustic ganglion; fgt, foregut; gg, glossopharyngeal ganglion; li, liver; lmc, lateral motor column; lu, lung; mb, midbrain; nt, neural tube; ov, otic vesicle; sg, sympathetic ganglion; si, small intestine; sn, segmental nerve; t, tail; tg, trigeminal ganglion; vg, vagus ganglion
At the trunk level (just distal to forelimb) of E12.5 Wnt1-Cre/R26R embryos, X-gal-positive cells were localized in the dorsal NT, DRG, in the myenteric region of the small intestine and the segmental nerves of the tail (Figure 1b). At the same A–P level of the E12.5 Wnt1-Cre/R26R/enb5 embryos, only very faint X-gal staining was detected in the dorsal NT and there was no X-gal staining in the DRG (Figure 1b). Furthermore, the DRG in Wnt1-Cre/R26R/enb5 was smaller than that in Wnt1-Cre/R26R. Reduced X-gal staining was also observed in the myenteric region of the small intestine in Wnt1-Cre/R26R/enb5.
NCC-derived melanoblasts were present between the skin epidermis and the dermis in Wnt1-Cre/R26R at E12.5 (Figure 2b; arrowheads), but not in Wnt1-Cre/R26R/enb5.
Skin hypopigmentation and defective ENS development in Wnt1-Cre/enb5 mice. (a) Melanoblasts in enb5 and Wnt1-Cre/enb5 embryos were localized by in situ hybridization for Dct (purple). Regions highlighted with dotted line were magnified and shown on the right. (b) NCC-derived melanoblasts (stained blue; arrowheads) in Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 embryos were localized by X-gal staining. (c) At postnatal day 3 (P3), areas of fur without pigment were observed in Wnt1-Cre/enb5 (mut) mice. Fur was properly pigmented in enb5 (ctrl) littermates. At postnatal day 21 (P21), Wnt1-Cre/enb5 mice developed skin hypopigmentation. (d) NCC-derived melanocytes (stained blue) in adult Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 mouse skin were localized by X-gal staining. Hair follicles were demarcated with broken line. (e) Enteric neuroblasts (stained blue) in the developing gut of E16.5 Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 embryos were localized by X-gal staining. Dotted regions were magnified and shown as insets. The number of gastrointestinal tract showing different extents of colonization of enteric neuroblasts was indicated by ‘n’. (f) Guts of P28 enb5 and Wnt1-Cre/enb5 mice were dissected for morphological examination. The proximal colon of Wnt1-Cre/enb5 mice was swollen but the distal colon was constricted. Sections of the proximal and distal colons of enb5 and Wnt1-Cre/enb5 mice were immunostained (green) for Tuj1 to localize enteric neurons. The levels from which the transverse sections were analyzed were indicated by ‘a’ and ‘b’. Number of gut tissues, embryos or mice analyzed was indicated by ‘n’. ce, cecum; cm, circular muscle; co, colon; der, dermis; e, eye; epi, epidermis; mu, mucosa; lm, longitudinal muscle; ov, otic vesicle; st, stomach; si, small intestine
Defects in cranial, sympathetic and DRG in Wnt1-Cre/enb5
Glia and neuronal differentiation in the cranial, sympathetic and DRG in enb5 and Wnt1-Cre/enb5 embryos were analyzed using Sox10 (NCC and glia marker) and Islet1/2 (neuron marker). Both transcripts were detected in the trigeminal ganglion, facio-acoustic ganglion, glossopharyngeal ganglion, vagus ganglion, otic vesicle, sympathetic ganglion and DRG in both enb5 and Wnt1-Cre/enb5 at E9.5 and E10.5 (Figures 1c and d). The temporal and spatial expression patterns of Sox10 and Islet1/2 in the cranial, sympathetic and DRG were comparable, but the intensity was lower in Wnt1-Cre/enb5, in which Hoxb5 signaling was perturbed in all NCC.
We determined the total volume of the DRG from 300 consecutive transverse sections from the trunk level (between the forelimb bud and the hindlimb bud) of E11.5 enb5 and Wnt1-Cre/enb5 embryos. The average volume of DRG in Wnt1-Cre/enb5 was only one-quarter of that in enb5 (Figure 1f). Immunostaining showed Islet1/2 protein expression in neurons in the DRG and the lateral motor column of enb5 and Wnt1-Cre/enb5 mice (Figure 1e); however, the DRG was markedly smaller in Wnt1-Cre/enb5. In both genotypes, the percentages of neurons in DRG were comparable (Wnt1-Cre/enb5 68.5±5.2% (mean±S.D.), enb5 73.3±2.9% (mean±S.D.; Figure 1f), suggesting that the shrinkage of DRG was not solely attributable to loss of neurons.
Defects in skin melanoblasts in Wnt1-Cre/enb5
Trunk NCCs migrate dorsal-laterally between the surface ectoderm and dorsal surface of the somites, then differentiate into melanoblasts expressing melanin-producing enzyme dopachrome tautomerase (Dct). These melanoblasts are precursors of pigment cells in skin and iris of the eye. In enb5 embryos, at E9.5, a cluster of Dct-positive melanoblasts appeared at the cervical region posterior to the otic vesicle (Figure 2a). By E10.5, this cervical cluster dispersed and migrated ventrally, populating the branchial arches and anterior trunk (Figure 2a). Strong Dct expression was also detected in the developing eye and the telecephalon. In Wnt1-Cre/enb5 embryos, there was a drastic reduction in the number of Dct-expressing melanoblasts in the cervical region at E9.5 (Figure 2a). By E10.5, fewer melanoblasts had populated the branchial arches, and there were no detectable melanoblasts in the anterior trunk region (Figure 2a). In contrast, Dct expression in the developing eye and in the telecephalon in Wnt1-Cre/enb5 embryos was comparable to that in enb5 embryos. This absence of melanoblasts during early development led to skin hypopigmentation in Wnt1-Cre/enb5 mice at postnatal day 3 when fur started to form (Figure 2c). By postnatal day 21, patches of white fur were obvious (Figure 2c). In adults, melanocytes were detected in the hair follicles in Wnt1-Cre/R26R and in the hair follicles of the skin with normal pigmentation in Wnt1-Cre/enb5, but not in the hair follicles in the white patches (Figure 2d).
Defects in ENS in Wnt1-Cre/enb5
We investigated the colonization of the intestine by NCC using X-gal. By E12.5, X-gal-positive neuroblasts colonized the small intestine of both Wnt1-Cre/R26R and Wnt1-Cre/R26R/enb5 embryos (Figure 1b). However, the intensity in the small intestine was lower in Wnt1-Cre/R26R/enb5, which indicated that there were fewer neuroblasts. By E16.5, neuroblasts had completed populating the entire gut, reaching the end of the colon in Wnt1-Cre/R26R embryos (Figure 2e). In contrast, in five out of six Wnt1-Cre/R26R/enb5 embryos of the same litter, the neuroblasts had colonized only as far as the cecum, or up to half of the length or anterior two-thirds of the colon (Figure 2e). This defective colonization of the intestine by neuroblasts resulted in ENS anomaly in postnatal mice. At P28, a phenotype of megacolon, which resembled HSCR in humans, was observed in Wnt1-Cre/enb5 (Figure 2f). Immunostaining for Tuj1 (a pan-neuronal marker) detected no enteric neurons in the distal part of the affected colon in Wnt1-Cre/enb5, whereas the proximal part was normally populated (Figure 2f).
In summary, disruption of Hoxb5 signaling by enb5 in NCC from the entire A–P axis of the NT resulted in multiple NCC-developmental defects including hypoplasia of cranial, sympathetic and DRG, hypopigmentation and ENS defects in mice.
Hoxb5 function is required for the survival of vagal and trunk NCC
The developmental defects observed in multiple NCC-derived lineages in Wnt1-Cre/enb5 mice prompted us to examine whether the NCC population was depleted in these embryos. Using TUNEL assay, we detected apoptotic cells in the dorsal NT and the regions between the cardinal vein and dorsal aorta in the levels of both vagal and trunk regions in Wnt1-Cre/enb5 at E9.5 and E10.5 (Figures 3b and d), whereas apoptotic cells were rarely detected in enb5 embryos (Figures 3a and c). The majority of the apoptotic cells were immunostained for p75NTR (Figure 3e), a marker of NCC. Thus, blocking the activity of Hoxb5 in NCC along the whole A–P axis of the NT leads to apoptosis of NCC. Apoptosis of NCC was also reported in mice nullified for Sox9.27
Apoptosis of NCCs in Wnt1-Cre/enb5 mice. (a–d) TUNEL staining was performed to localize apoptotic cells (green) on Wnt1-Cre or enb5 embryos and Wnt1-Cre/enb5 sections of the trunk level. Boxed regions were magnified and shown as insets. (e) Section of E9.5 Wnt1-Cre/enb5 embryo was examined by TUNEL staining (green) and immunofluorescence (red) using anti-p75NTR serum. Number of embryos analyzed was indicated by ‘n’. cv, cardinal vein; da, dorsal aorta; ht, heart; pe, pharyngeal ectoderm
Deletion of Sox9 in Wnt1-Cre expressing cells resulted in vagal and trunk NCC apoptosis and developmental defects
Sox9 is a NCC marker and deletion of Sox9 specifically in NCC by Wnt1-Cre system resulted in cell apoptosis in the ventral and dorsal NT, along the NCC migration pathways in E9.5 Wnt1-Cre/Sox9flox/flox embryos (Figure 4a). Apoptotic cells were not detected in Sox9flox/flox control littermates (Figure 4a).
Apoptosis of NCCs and defective development of NCC-derived structures in Wnt1-Cre/Sox9flox/flox mice. (a) TUNEL staining on E9.5 Sox9flox/flox and Wnt1-Cre/Sox9flox/flox embryo sections of the trunk level was performed to localize apoptotic cells (green). (b and c) NCC-derived cells (stained blue) in embryos (b) and gut (c) of Wnt1-Cre/R26R and Wnt1-Cre/R26R/Sox9flox/flox mice were localized by X-gal staining. (d) Melanoblasts (purple) were localized by in situ hybridization for Dct in E10.5 Sox9flox/flox and Wnt1-Cre/Sox9flox/flox embryos. Regions highlighted were magnified and shown on the right. (e) NCC-derived melanoblasts (stained blue; arrowheads) on section of Wnt1-Cre/Sox9flox/flox embryos were localized by X-gal staining. Number of embryos analyzed was indicated by ‘n’. ba, branchial arch; ce, cecum; co, colon; cv, cardinal vein; da, dorsal aorta; der, dermis; drg, dorsal root ganglion; e, eye; epi, epidermis; ht, heart; nt, neural tube; se, surface ectoderm; sg, sympathetic ganglion; st, stomach; si, small intestine; te, telecephalon
In Wnt1-Cre/R26R embryos, X-gal-positive NCCs were found populating the trigeminal and facio-acoustic ganglion (arrows; Figure 4b), NT and DRG at E9.5; and sympathetic ganglia and DRG at E10.5 (Figure 4b). However, the blue staining in the corresponding structures at both stages of Wnt1-Cre/R26R/Sox9flox/flox embryos was relatively less extensive and less intense (Figure 4b). We determined the total volume of the DRG from the trunk level (between the forelimb bud and the hindlimb bud) of E11.5 Wnt1-Cre/Sox9flox/flox and Sox9flox/flox embryos, and found that DRG volume in Wnt1-Cre/Sox9flox/flox was only 60% of that in Sox9flox/flox (Supplementary Figure 1A). The percentages of neurons in DRG in both genotypes were comparable (Supplementary Figure 1B), implying that, as in the conditional Hoxb5 mutants, the shrinkage of DRG in the absence of Sox9 activity was not solely attributable to loss of neurons.
We also investigated the migration of enteric neuroblasts and skin melanoblasts in Wnt1-Cre/Sox9flox/flox using X-gal staining. At E14.5, neuroblasts had colonized the entire gut and reached the distal colon in Wnt1-Cre/R26R (Figure 4c). In contrast, the neuroblasts had populated only up to the cecum and aganglionosis was observed in the distal colon of Wnt1-Cre/R26R/Sox9flox/flox embryos (Figure 4c).
The expression of Dct in the skin epidermis of Wnt1-Cre/Sox9flox/flox and Sox9flox/flox embryos at E10.5 was comparable in terms of both pattern and intensity (Figure 4d). By E12.5, melanoblasts were found between the epidermis and dermis of both Wnt1-Cre/R26R/Sox9flox/flox and Wnt1-Cre/R26R at the trunk level (compare Figures 4e and 2b), indicating that melanoblast development was largely unaffected by the Sox9 deletion in NCC.
Perturbation of Hoxb5 function disrupts Sox9 expression in NCC
Blocking Sox9 activity in NCC along the A–P axis of the NT leads to NCC apoptosis and developmental defects of DRG, sympathetic ganglion and ENS, which resemble most of the developmental defects observed when Hoxb5 activity is blocked in the same regions. This suggested that Hoxb5 and Sox9 might function in the same developmental pathway.
To investigate the relationship between Hoxb5 and Sox9 in our mutants, we assessed Sox9 expression. In E9.5 enb5, Sox9-expressing NCCs were localized in the dorsal NT, along the dorsal-lateral migration pathway under the surface ectoderm and ventral-medial migration pathway to the cardinal vein (Figure 5a). In contrast, in Wnt1-Cre/enb5 mice, no Sox9 immuno-positive NCCs were seen in the dorsal NT or under the surface ectoderm; and only a few Sox9-expressing NCCs were found in the vicinity of the cardinal vein (Figure 5b). These results suggested that Sox9 might act downstream of Hoxb5 in the same signaling pathway.
Reduction of Sox9 expression in Wnt1-Cre/enb5 mice. (a and b) Sox9-expressing cells (green; arrows) were localized in E9.5 Wnt1-Cre or enb5 (a) and Wnt1-Cre/enb5 (b) embryos by immunofluorescence using anti-Sox9 serum on sections of the trunk level. Regions highlighted were magnified and shown as insets. Arrowhead denoted the notochord. Number of embryos analyzed was indicated by ‘n’. cv, cardinal vein; da, dorsal aorta; ht, heart; nt, neural tube
HOXB5 binds to SOX9 promoter
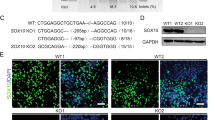
In silico analysis predicted three potential HOX-binding sites in the SOX9 promoter: R1A (−483 to −466), R1B (−468 to −450) and R2 (−35 to −16) (Figure 6a). We evaluated the binding of HOXB5 to the SOX9 promoter using electro-mobility shift assay (EMSA) with a glutathione S-transferase (GST)-HOXB5 fusion protein and polymerase chain reaction (PCR) products spanning these binding sequences (Figure 6a). A retarded band was observed only in the lanes containing the GST-HOXB5 and the PCR fragments (probe) of R1 and R2. No retarded band was observed if GST-HOXB5 was replaced with GST protein, or if excess unlabeled probe was included in the reaction mixtures. More importantly, the specific binding between GST-HOXB5 and the probe was completely abolished if the respective HOX-binding sequence was deleted from the probe.
Hoxb5 binds and trans-activates the Sox9 promoter. (a) Human SOX9 promoter sequence with transcription start site (bent arrow) and putative HOX-binding sequences (boxes) indicated. Dotted lines demarcate region 1 (R1) and region 2 (R2) fragments for EMSA. Arrows denote primers for quantitative PCR. EMSA for HOXB5 binding to SOX9 promoter. ‘+’ or ‘–’ indicate the presence or absence of a reagent in EMSA. (b and c) ChIP and quantitative PCR for HOXB5 binding to SOX9 promoter in transfected cells (b), and in the CNS of E9.5 wild-type mouse embryos (c). y axis indicated fold enrichment (mean±S.D.) normalized to control IgG. (d) HOXB5 trans-activation from full-length (FL; −1034 to +67) or HOX-binding sequence deleted (−R1A; −R1B; −R2) SOX9 promoter. Fold changes (mean±S.D.) were determined relative to luciferase unit of pRC/CMV. (e) HOXB5 trans-activation from full-length SOX9 promoter with or without enb5 inhibition. Fold increment (mean±S.D.) was determined relative to luciferase unit of pRC/CMV (EV). Trans-activation of full-length SOX9 promoter by HOXB5 with or without Flag-enb5. The relative luciferase activity in cells transfected with HOXB5 alone was arbitrarily regarded as 100%. (f) ChIP and quantitative PCR for Flag-enb5 binding to SOX9 promoter. y axis indicated fold enrichment (mean±S.D.) normalized to control IgG. *P<0.05; **P<<0.05
We extended this analysis by looking at the binding of HOXB5 to the SOX9 promoter in a neuroblastoma cell line (SK-N-SH) transfected with HOXB5, using chromatin immunoprecipitation (ChIP) followed by quantitative PCR. Binding of HOXB5 to R1 and R2 was significantly enriched compared with the nonspecific IgG control (Figure 6b). Furthermore, we confirmed the binding of Hoxb5 to the Sox9 promoter in vivo in the central nervous system (CNS) of E9.5 wild-type embryos (Figure 6c).
HOXB5 trans-activation from SOX9 promoter is suppressed by enb5
To study whether HOXB5 protein trans-activates the SOX9 promoter, we used a luciferase reporter construct consisting of 1100 bp (−1034 to +67) of the SOX9 gene 5′ of the luciferase gene in SK-N-SH cells. HOXB5 increased the transcription by 5.8±0.1 (mean±S.D.) fold compared with the pRC/CMV control (Figure 6d). Deletion of R1A and R1B from the SOX9 promoter reduced the induction (to 4.98±0.10 and 3.61±0.20 (mean±S.D.) fold, respectively). Conversely, deletion of R2 increased the induction to 13.52±1.52 (mean±S.D.) fold. Our data revealed that binding of HOXB5 to these elements of the SOX9 promoter increases overall transcription from the SOX9 promoter, which could be attributed to the differential binding of HOXB5 onto R1 to R2.
Transfection of the dominant-negative form of Hoxb5, enb5, prominently suppressed the trans-activation of SOX9 by HOXB5 (Figure 6e). To test the mechanism, we generated a vector expressing a Flag-tagged enb5 (Flag-enb5). As expected, Flag-enb5 suppressed HOXB5-transactivated SOX9 promoter activity by 50% (Figure 6e). ChIP and quantitative PCR analysis revealed a modest but significant enrichment (1.98±0.08 and 1.25±0.06 (mean±S.E.M.) fold for R1 and R2, respectively) of binding of Flag-enb5 to the SOX9 promoter compared with a nonspecific IgG control. This suggested that Flag-enb5, thus enb5, binds to the same regions of the SOX9 promoter as HOXB5 (Figure 6f). Therefore, HOXB5 binds to the SOX9 promoter and this can be suppressed by the dominant-negative chimera, enb5.
Sox9 alleviates enb5-induced cell death and Hoxb5 induces Sox9 expression in ovo
In ovo electroporation of chick NT with enb5 resulted in obvious cell death in the transfected region but only a few apoptotic cells were detected in the contralateral non-electroporated side. Co-electroporation of Hoxb5 or Sox9 notably reduced enb5-induced cell death (Figure 7a), again suggesting that Sox9 and Hoxb5 act in the same signaling pathway. When chick NT was electroporated with Hoxb5, ectopic expression of Sox9 was induced on the transfected side at 6–12 h, confirming the trans-activation action of Sox9 by Hoxb5 (Figure 7b).
Sox9 alleviates enb5-induced cell death and Hoxb5 induces Sox9 expression in chick NT. (a) In ovo electroporation with enb5, Hoxb5 and Sox9, either alone or in combination. Chick NT was assayed for TUNEL (red) and immunofluorescence for GFP (green) 24 h post-transfection on transverse sections at trunk level. Number of embryos analyzed was indicated by ‘n’. (b) In ovo electroporation with chick Hoxb5, chick NT was assayed for GFP (green) and Sox9 expression (purple) 12 h post-transfection by immunofluorescence for GFP (green) and in situ hybridization for Sox9 (purple) on transverse sections at trunk level. (n/n) indicated number of embryos showing positive detection (GFP or Sox9) versus number of transfected embryos. (c) Schematic diagram of the HH10-11 chick embryo 12–24 h after electroporation, dotted line indicated the level of sections used for immunofluorescence or in situ hybridization analysis
Discussion
In this study, we used mice expressing a dominant-negative form of Hoxb5 (enb5) in NCC from all along the NT to investigate the impact of abnormal Hoxb5 signaling in the development of NCC-derived tissues. These Wnt1-Cre/enb5 mice displayed defects in multiple NCC-derived tissues, suggesting that NCCs were affected before lineage specification. TUNEL analysis revealed that pre-migratory and migratory NCC underwent apoptosis in E9.5 Wnt1-Cre/enb5 embryos. Taken together, our previous report22 and this study indicate that Hoxb5 regulates the survival of vagal and trunk NCC.
In mouse embryos, vagal NCCs enter the foregut at E9.5 and colonize the gut reaching distal hindgut at E14.5. During migration, NCCs proliferate, interact with gut mesenchyme, and differentiate into neurons and glia. NCC growth defects and errors in their interactions with gut mesenchyme contribute to abnormal ENS.28, 29 In Wnt1-Cre/enb5 embryos, expression of enb5 by Wnt1-Cre led to apoptosis of pre-migratory and early migratory NCC. Expression of enb5 in vagal NCC by another Cre mouse (b3-IIIa-Cre) resulted in NCC migration defect in the intestine.22 In b3-IIIa-Cre/enb5 mice, enb5-expressing neuroblasts survive, proliferate and differentiate normally in the intestine, and NCC apoptosis was not observed. The lack of NCC apoptosis in b3-IIIa-Cre/enb5 embryos is attributed to a more restricted and late expression of enb5 in b3-IIIa-Cre/enb522 than that in Wnt1-Cre/enb5. All these indicate that the differentiation and proliferation of neuroblasts in the intestine is largely unaffected by enb5. ENS defect in Wnt1-Cre/enb5 mice is mainly attributed to NCC apoptosis, which leads to reduction of the population size of the NCC entering the gut. Whether enb5 affects NCC formation requires further investigation. Hoxb5 is also expressed in gut mesenchyme, suggesting it may influence the gut environment.17 However, enb5 is only expressed in NCC in this and previous22 studies, indicating that ENS anomalies in these mice are due to defective NCC growth.
Sox9 is required for NCC development, and we have demonstrated that blocking Hoxb5 downregulated Sox9 expression in these cells. HOXB5 binds to SOX9 promoter and induces SOX9 expression, and this induction of SOX9 promoter activity was substantially reduced by enb5. The human and mouse Sox9 promoters share high sequence conservation (>70%), and Hoxb5-binding sites were also identified within the mouse Sox9 promoter (Supplementary Figure 2). ChIP assay on CNS confirmed that Hoxb5 bound to the mouse Sox9 promoter. More importantly, expression of enb5 in NCC reduced Sox9 expression in pre-migratory and migratory NCC in Wnt1-Cre/enb5 embryos. Our findings revealed that apoptosis of trunk NCC and defects in multiple trunk NCC-derived tissues in Wnt1-Cre/enb5 were consequences of downregulated Sox9 expression in NCC.
Wnt1-Cre/Sox9flox/flox mice displayed craniofacial defects resembling campomelic dysplasia in human.30 Cranial NCC-derived cartilages and endochondral bones of the head were completely absent in Wnt1-Cre/Sox9flox/flox mice, but development of trunk NCC-derived structures was not addressed. Wnt1-Cre/enb5 mice also display domed skull and short snout as Wnt1-Cre/Sox9flox/flox mice, but the cranial NCC-derived skeletal elements are not absent but malformed in Wnt1-Cre/enb5 heads (unpublished data). Different fates of NCC are dictated by both the intrinsic properties and external environmental factors. Trunk NCCs show overlapping Hoxb5 and Sox9 expression pattern and they committed to apoptosis when there is a loss of Hoxb5 or Sox9 functions in respective mutant mice. However, in cranial NCC, the expression patterns of Hoxb520, 21, 24 and Sox931, 32 in mice are not completely overlapping, which indicates that Hoxb5 alone is neither sufficient nor essential for Sox9 expression. Moreover, the apoptotic patterns seen in the developing heads of Wnt1-Cre/Sox9flox/flox and Wnt1-Cre/enb5 mice were not exactly the same too (unpublished data). These suggested that the regulation of Hoxb5 on Sox9 in trunk NCC may not be identical to that in cranial NCC. Therefore, it is not totally unexpected to observe phenotypic differences in some NCC-derived tissues between Wnt1-Cre/Sox9flox/flox and Wnt1-Cre/enb5 mice, and Sox9 deletion or enb5 expression by Wnt1-Cre may affect different developmental aspects of cranial NCC.
Sox9 provides the competence for trunk NCC to migrate from the NT; it is also required for NCC survival.27 However, not all the trunk NCC died in Sox9−/− mutants, as some NCC-derived neuro-glial components were still present in the DRG and peripheral nerve as shown by the residual expression of Brn3.0 (neuron marker) and Sox10 (glia marker),27 indicating that the generation and survival of a subpopulation of trunk NCC was independent of Sox9 function.
Wnt1-Cre/enb5 and Wnt1-Cre/Sox9flox/flox mutants display some common phenotypes including NCC apoptosis, defective ENS, reduction of DRG and sympathetic ganglia, however, the pigmentation defect was observed only in Wnt1-Cre/enb5. The ENS defects in Wnt1-Cre/Sox9flox/flox mutants denoted that Sox9 was also required for vagal NCC development in addition to its inductive and survival roles in trunk NCC. In mouse, trunk NCCs migrate to their target organs in three waves: the first two migrate ventral-medially to generate the neuro-glial derivatives at E9.0-E10.5, including the DRG and sympathetic ganglion; the third wave migrates dorsal-laterally along the surface ectoderm and dermomyotome to form the skin melanoblasts at E10.5–E13.5.33 In line with the role of Sox9 in NCC survival, deletion of Sox9 in Wnt1-Cre expressing cells led to NCC apoptosis, DRG, sympathetic ganglion and ENS defects. The lack of a pigmentation defect in Wnt1-Cre/Sox9flox/flox mice implies that the survival of trunk NCC at the later embryonic stage, which give rise to skin pigment cells, is not so dependent on Sox9.
Another SoxE gene, Sox10, is essential for the specification of NCC into melanoblast and glia cells.34, 35, 36, 37 In silico analysis identified potential Hox-binding sites in the Sox10 promoter (unpublished data), future investigation could address the relationship between Hoxb5, Sox10 and pigmentation defect in Wnt1-Cre/enb5 mice.
Hirschsprung disease (HSCR, MIM142623) is a neurocristopathy of ENS deficiency, characterized by the absence of enteric ganglia from variable lengths of the gut. Approximately 30% of HSCR patients exhibit additional NCC-associated anomalies, known as syndromic HSCR.28, 38, 39, 40 Mutations in SOX10, EDN3 or EDNRB have been identified to be responsible for 65–85% of the syndromic HSCR cases with pigmentation defects, whereas genes responsible for the remaining cases still left undetermined.41 Especially for EDN3, a reduction in spinal sensory innervation of the rectum was reported in mice with disruption of Edn3 gene expression.42 In line with this, this study also showed that perturbation of Hoxb5 function in NCC of mice causes multiple neurological phenotypes and hypopigmentation, resembling some of the phenotypes of syndromic HSCR. Nevertheless, the causal relationship between abnormal HOXB5 function and HSCR in human deserves further investigation, in particular, the identification of a dominant-negative type of HOXB5 mutation in these patients.
Materials and Methods
Mouse lines
The enb5,22 R26R,43 Sox9flox/flox44 and Wnt1-Cre26 mice were maintained on mixed genetic backgrounds. Genotyping of mice was performed by PCR (Supplementary Information). All experimental procedures were approved by the Committee on the Use of Live Animals at the University of Hong Kong (CULTRA 2038-009).
Immunofluorescence staining
Immunofluorescence staining was performed according to standard protocol. Antibodies used were: Islet1/2 (Developmental Studies Hybridoma Bank, Iowa City, IA, USA; 51.4H9, 1 : 100); Sox9 (Chemicon, Billerica, MA, USA; ab5535, 1 : 500); GFP (AbDirect, Oxford, UK; 4745–1051, 1 : 1000).
TUNEL assay and immunofluorescence staining
Apoptotic cells on sections detected using In Situ Cell Death Detection Kit (Roche Applied Science, Indianapolis, IN, USA). After TUNEL, sections were incubated with antibody for GFP (AbDirect, 4745–1051, 1 : 1000) or p75NTR (Chemicon, 07–476, 1 : 2000). Images were taken with Nikon Eclipse 80i microscope (Melville, NY, USA) mounted with SPOT RT3 microscope digital camera (DIAGNOSTIC Instruments, Inc., Sterling Heights, MI, USA). Photos were compiled using Adobe Photoshop 7.
Determination of the sizes of DRG in embryos
Transverse sections (10 μm in thickness) from the trunk level (between the forelimb bud and the hindlimb bud) were mounted onto glass slides, stained with hematoxylin and eosin. Photos of 300 consecutive sections of each embryo were taken at 100 × on a Nikon Eclipse E600 microscope (Tokyo, Japan) fitted with Sony Digital Camera DSM1200F (Tokyo, Japan). DRG on either side of the NT in the photos were demarcated manually. The area of DRG on each section was determined using the software ImageJ (NIH, Bethesda, MD, USA) and an arbitrary value was given. The volume of DRG on each section was then determined by multiplying the total area of the DRG (on each side of the NT) by 10 (each section was of 10 μm in thickness).
In situ hybridization
In situ hybridization was performed on whole embryos to detect the expression of Sox10 and Islet1/2 as previously described.45
Electro-mobility shift assay
PCR products spanning the putative HOX-binding sites of the human SOX9 promoter were labeled with Biotin-11-UTP (Pierce, Thermo Fisher Scientific, Rockland, MA, USA), EMSA was performed as previously described.46
ChIP and quantitative PCR assay
ChIP assay on human neuroblastoma cell line SK-N-SH (#HTB-11) (ATCC, Manassas, VA, USA) transfected with HOXB5 or Flag-enb5 was performed as previously described.46 ChIP assay on mouse developing CNS was performed with head and NT tissues from 27 E9.5 WT mouse embryos with minor modifications.46, 47 See Supplementary Information for quantitative PCR primers for human and mouse Sox9 promoters.
Transient transfection and dual-luciferase reporter assay
A SOX9-luciferase reporting construct containing the 1100 bp (−1034 to +36) DNA fragment of the human SOX9 promoter48 and the luciferase gene was used for the study. Mutated SOX9-luciferase constructs with the predicted HOX-binding sites deleted were generated using QuikChange Site-Directed Mutagenesis Kit (Stratagene, Santa Clara, CA, USA). The dual-luciferase reporter assay was performed as previously described.46 At least two independent triplicate or quintuplicate experiments were performed, and the luciferase activity was presented as relative luciferase unit normalized with the Renilla luciferase internal control.
Chick in ovo electroporation
Enb5 and chick Hoxb5 were cloned into pCIG expression vector,49 upstream of an internal ribosomal entry site (IRES) and a nuclear localization sequence-tagged enhanced green fluorescent protein (EGFP). HH10-11 chick embryos (SIPAFASI, Jinan, China) were electroporated with Hoxb5, enb5 or Sox950 (4 μg/μl) either alone or in combination. pCAGGS-IRES-nls-EGFP (pCIG) was co-injected at a concentration of 1 μg/μl to monitor the injection and electroporation.
Statistical analyses
ANOVA was performed for all experiments to calculate the differences between groups, and P-value <0.05 was regarded as statistical significant.
Abbreviations
- NCC:
-
neural crest cell
- A–P:
-
anterior–posterior
- ENS:
-
enteric nervous system
- enb5:
-
engrailed-Hoxb5
- NT:
-
neural tube
- X-gal:
-
beta-galactosidase
- DRG:
-
dorsal root ganglion
- Dct:
-
dopachrome tautomerase
- E:
-
embryonic day
- GST:
-
glutathione S-transferase
- ChIP:
-
chromatin immunoprecipitation
- PCR:
-
polymerase chain reaction
- CNS:
-
central nervous system
- HSCR:
-
Hirschsprung disease
References
Kalcheim C, Le Douarin N . The Neural Crest. Cambridge University Press: Cambridge, UK, 1999.
Le Douarin NM, Creuzet S, Couly G, Dupin E . Neural crest cell plasticity and its limits. Development 2004; 131: 4637–4650.
Sauka-Spengler T, Bronner-Fraser M . A gene regulatory network orchestrates neural crest formation. Nat Rev Mol Cell Biol 2008; 9: 557–568.
Anderson RB, Stewart AL, Young HM . Phenotypes of neural-crest-derived cells in vagal and sacral pathways. Cell Tissue Res 2006; 323: 11–25.
Burns AJ, Douarin NM . The sacral neural crest contributes neurons and glia to the post-umbilical gut: spatiotemporal analysis of the development of the enteric nervous system. Development 1998; 125: 4335–4347.
Kapur RP . Colonization of the murine hindgut by sacral crest-derived neural precursors: experimental support for an evolutionarily conserved model. Dev Biol 2000; 227: 146–155.
Nagy N, Brewer KC, Mwizerwa O, Goldstein AM . Pelvic plexus contributes ganglion cells to the hindgut enteric nervous system. Dev Dyn 2007; 236: 73–83.
Serbedzija GN, Burgan S, Fraser SE, Bronner-Fraser M . Vital dye labelling demonstrates a sacral neural crest contribution to the enteric nervous system of chick and mouse embryos. Development 1991; 111: 857–866.
Wang X, Chan AK, Sham MH, Burns AJ, Chan WY . Analysis of the sacral neural crest cell contribution to the hindgut enteric nervous system in the mouse embryo. Gastroenterology 2011; 141: 992–1002 e1-6.
Akin ZN, Nazarali AJ . Hox genes and their candidate downstream targets in the developing central nervous system. Cell Mol Neurobiol 2005; 25: 697–741.
Carpenter EM . Hox genes and spinal cord development. Dev Neurosci 2002; 24: 24–34.
Duboule D, Morata G . Colinearity and functional hierarchy among genes of the homeotic complexes. Trends Genet 1994; 10: 358–364.
Durston AJ, Jansen HJ, In der Rieden P, Hooiveld MH . Hox collinearity—a new perspective. Int J Dev Biol 2011; 55: 899–908.
Lumsden A, Krumlauf R . Patterning the vertebrate neuraxis. Science 1996; 274: 1109–1115.
Trainor PA, Krumlauf R . Patterning the cranial neural crest: hindbrain segmentation and Hox gene plasticity. Nat Rev Neurosci 2000; 1: 116–124.
Trainor PA, Manzanares M, Krumlauf R . Genetic interactions during hindbrain segmentation in the mouse embryo. Results Probl Cell Differ 2000; 30: 51–89.
Fu M, Lui VC, Sham MH, Cheung AN, Tam PK . HOXB5 expression is spatially and temporarily regulated in human embryonic gut during neural crest cell colonization and differentiation of enteric neuroblasts. Dev Dyn 2003; 228: 1–10.
Hogan BL, Holland PW, Lumsden A . Expression of the homeobox gene, Hox 2.1, during mouse embryogenesis. Cell Differ Dev 1988; 25 (Suppl): 39–44.
Holland PW, Hogan BL . Spatially restricted patterns of expression of the homeobox-containing gene Hox 2.1. during mouse embryogenesis. Development 1988; 102: 159–174.
Krumlauf R, Holland PW, McVey JH, Hogan BL . Developmental and spatial patterns of expression of the mouse homeobox gene, Hox 2.1. Development 1987; 99: 603–617.
Kuratani SC, Wall NA . Expression of Hox 2.1 protein in restricted populations of neural crest cells and pharyngeal ectoderm. Dev Dyn 1992; 195: 15–28.
Lui VC, Cheng WW, Leon TY, Lau DK, Garcia-Barcelo MM, Miao XP et al. Perturbation of hoxb5 signaling in vagal neural crests down-regulates ret leading to intestinal hypoganglionosis in mice. Gastroenterology 2008; 134: 1104–1115.
Pitera JE, Smith VV, Thorogood P, Milla PJ . Coordinated expression of 3' hox genes during murine embryonal gut development: an enteric Hox code. Gastroenterology 1999; 117: 1339–1351.
Wall NA, Jones CM, Hogan BL, Wright CV . Expression and modification of Hox 2.1 protein in mouse embryos. Mech Dev 1992; 37: 111–120.
Rancourt DE, Tsuzuki T, Capecchi MR . Genetic interaction between hoxb-5 and hoxb-6 is revealed by nonallelic noncomplementation. Genes Dev 1995; 9: 108–122.
Danielian PS, Muccino D, Rowitch DH, Michael SK, McMahon AP . Modification of gene activity in mouse embryos in utero by a tamoxifen-inducible form of Cre recombinase. Curr Biol 1998; 8: 1323–1326.
Cheung M, Chaboissier MC, Mynett A, Hirst E, Schedl A, Briscoe J . The transcriptional control of trunk neural crest induction, survival, and delamination. Dev Cell 2005; 8: 179–192.
Amiel J, Sproat-Emison E, Garcia-Barcelo M, Lantieri F, Burzynski G, Borrego S et al. Hirschsprung disease, associated syndromes and genetics: a review. J Med Genet 2008; 45: 1–14.
Tam PK, Garcia-Barcelo M . Genetic basis of Hirschsprung’s disease. Pediatr Surg Int 2009; 25: 543–558.
Mori-Akiyama Y, Akiyama H, Rowitch DH, de Crombrugghe B . Sox9 is required for determination of the chondrogenic cell lineage in the cranial neural crest. Proc Natl Acad Sci USA 2003; 100: 9360–9365.
Morais da Silva S, Hacker A, Harley V, Goodfellow P, Swain A, Lovell-Badge R . Sox9 expression during gonadal development implies a conserved role for the gene in testis differentiation in mammals and birds. Nat Genet 1996; 14: 62–68.
Wright E, Hargrave MR, Christiansen J, Cooper L, Kun J, Evans T et al. The Sry-related gene Sox9 is expressed during chondrogenesis in mouse embryos. Nat Genet 1995; 9: 15–20.
Ruhrberg C, Schwarz Q . In the beginning: generating neural crest cell diversity. Cell Adh Migr 2010; 4: 622–630.
Paratore C, Goerich DE, Suter U, Wegner M, Sommer L . Survival and glial fate acquisition of neural crest cells are regulated by an interplay between the transcription factor Sox10 and extrinsic combinatorial signaling. Development 2001; 128: 3949–3961.
Potterf SB, Furumura M, Dunn KJ, Arnheiter H, Pavan WJ . Transcription factor hierarchy in Waardenburg syndrome: regulation of MITF expression by SOX10 and PAX3. Hum Genet 2000; 107: 1–6.
Potterf SB, Mollaaghababa R, Hou L, Southard-Smith EM, Hornyak TJ, Arnheiter H et al. Analysis of SOX10 function in neural crest-derived melanocyte development: SOX10-dependent transcriptional control of dopachrome tautomerase. Dev Biol 2001; 237: 245–257.
Southard-Smith EM, Kos L, Pavan WJ . Sox10 mutation disrupts neural crest development in Dom Hirschsprung mouse model. Nat Genet 1998; 18: 60–64.
Bonnet JP, Till M, Edery P, Attie T, Lyonnet S . Waardenburg-Hirschsprung disease in two sisters: a possible clue to the genetics of this association? Eur J Pediatr Surg 1996; 6: 245–248.
Toki F, Suzuki N, Inoue K, Suzuki M, Hirakata K, Nagai K et al. Intestinal aganglionosis associated with the Waardenburg syndrome: report of two cases and review of the literature. Pediatr Surg Int 2003; 19: 725–728.
Yoder BJ, Prayson RA . Shah-Waardenburg syndrome and Dandy-Walker malformation: an autopsy report. Clin Neuropathol 2002; 21: 236–240.
Pingault V, Ente D, Dastot-Le Moal F, Goossens M, Marlin S, Bondurand N . Review and update of mutations causing Waardenburg syndrome. Hum Mutat 2010; 31: 391–406.
Zagorodnyuk VP, Kyloh M, Nicholas S, Peiris H, Brookes SJ, Chen BN et al. Loss of visceral pain following colorectal distension in an endothelin-3 deficient mouse model of Hirschsprung's disease. J Physiol 2011; 589: 1691–1706.
Soriano P . Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat Genet 1999; 21: 70–71.
Akiyama H, Chaboissier MC, Behringer RR, Rowitch DH, Schedl A, Epstein JA et al. Essential role of Sox9 in the pathway that controls formation of cardiac valves and septa. Proc Natl Acad Sci USA 2004; 101: 6502–6507.
Cheng Y, Cheung M, Abu-Elmagd MM, Orme A, Scotting PJ . Chick sox10, a transcription factor expressed in both early neural crest cells and central nervous system. Brain Res Dev Brain Res 2000; 121: 233–241.
Zhu J, Garcia-Barcelo MM, Tam PK, Lui VC . HOXB5 cooperates with NKX2-1 in the transcription of human RET. PLoS One 2011; 6: e20815.
Liu W, Lagutin OV, Mende M, Streit A, Oliver G . Six3 activation of Pax6 expression is essential for mammalian lens induction and specification. EMBO J 2006; 25: 5383–5395.
Piera-Velazquez S, Hawkins DF, Whitecavage MK, Colter DC, Stokes DG, Jimenez SA . Regulation of the human SOX9 promoter by Sp1 and CREB. Exp Cell Res 2007; 313: 1069–1079.
Niwa H, Yamamura K, Miyazaki J . Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 1991; 108: 193–199.
Cheung M, Briscoe J . Neural crest development is regulated by the transcription factor Sox9. Development 2003; 130: 5681–5693.
Acknowledgements
Plasmids for riboprobe synthesis were provided as gifts: Dct (I Jackson, MRC Human Genetics Unit, Western General Hospital, Edinburgh); Islet1/2 (R Lovell-Badge, MRC NIMR, London, UK); chick Sox9 (PT Sharpe, King's College, London, UK); and Sox10 (M Tani, National Cancer Center Research Institute, Japan). The SOX9-lucifrease construct was provided by S Piera (Thomas Jefferson University, Philadelphia, PA, USA). Mouse strains used in this study were provided as gifts: R26R (PM Soriano, Mount Sinai School of Medicine), Sox9flox/flox (A Schedl and M-C Chaboissier, Centre de Biochimie, Parc Valrose, France), Wnt1-Cre (AP McMahon, Harvard University, USA). We thank Patrick Tam for comments on the manuscript, and J Marsh for proofreading the manuscript. This work was partly supported by the Hong Kong RGC GRF (HKU 7245/02M) and HKU Seed Funding for Basic Research (200811159088) to VCH Lui.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by RA Knight
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Kam, M., Cheung, M., Zhu, J. et al. Perturbation of Hoxb5 signaling in vagal and trunk neural crest cells causes apoptosis and neurocristopathies in mice. Cell Death Differ 21, 278–289 (2014). https://doi.org/10.1038/cdd.2013.142
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2013.142
Keywords
This article is cited by
-
Hirschsprung’s disease: clinical dysmorphology, genes, micro-RNAs, and future perspectives
Pediatric Research (2017)