Abstract
MicroRNAs (miRNAs) encoded by the miR-15/16 cluster are known to act as tumor suppressors. Expression of these miRNAs inhibits cell proliferation, promotes apoptosis of cancer cells, and suppresses tumorigenicity both in vitro and in vivo. miR-15a and miR-16-1 function by targeting multiple oncogenes, including BCL2, MCL1, CCND1, and WNT3A. Down-regulation of these miRNAs has been reported in chronic lymphocytic lymphoma (CLL), pituitary adenomas, and prostate carcinoma. This review summarizes the discovery, functions, and clinical relevance of these miRNAs in cancer, particularly CLL.
Similar content being viewed by others
Main
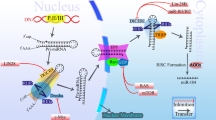
MicroRNAs (miRNAs) are a relatively recently identified class of regulatory noncoding RNAs, typically 20–23 nt in length, which function primarily by targeting specific messenger RNAs (mRNAs) for degradation or inhibition of translation and hence decreasing the expression of the resulting protein.1, 2 Their regulatory functions involve a variety of biological processes, including development, differentiation, apoptosis, survival, senescence, and metabolism.3, 4 Several hundred miRNAs have been identified and studied so far, but it is predicted that there are many more miRNAs whose significance and role in homeostasis and pathology is yet to be identified. Intriguingly, a significant fraction of miRNA genes map to regions implicated in cancer susceptibility and at fragile sites, suggesting a role of miRNAs in carcinogenesis.5, 6, 7 In 2002, we identified miR-15 and miR-16 as potential cancer genes in the pathogenesis of chronic lymphocytic leukemia (CLL).8 Specifically, miR-15a and miR-16-1, located at 13q14.3, were frequently deleted and/or down-regulated in CLL.8 This finding provided the first evidence that miRNA genes might be important for tumorigenesis.
To facilitate the efficient and comprehensive profiling of miRNA genes, a microarray chip defining the global levels for each known or predicted member of the miRNome was developed.9 Other technologies, including macroarrays,10 bead-based flow cytometric miRNA expression,11 and quantitative reverse transcription PCR12, 13 also became available. Collectively, these approaches have shown the differential expression of miRNA in human tissues and identified significant miRNA genes that are of biological and clinical relevance in human diseases.14 In particular, several research groups have sought to identify unique signatures or fingerprints of miRNAs in human malignancies in the hope that such knowledge would be helpful in exploring new venues and pathways that regulate the process of cancer evolution and would thus enable the design of novel diagnostic, prognostic, and therapeutic tools. Indeed, several miRNAs have been shown to have a promoting role in the development of primary tumors (reviewed in Ref.15); miRNAs that are amplified or overexpressed in cancer could act as oncogenes, whereas several miRNAs have been implicated as tumor suppressors based on their physical deletion or reduced expression in human cancer. Additionally, miRNAs have been implicated as modulators of tumor progression and metastasis (reviewed in Ref.16). As a result, we realize that deregulation of, at least some, miRNAs might also contribute to tumorigenesis.
In this review, we will discuss the classification of the miR-15a and miR-16-1 cluster as cancer genes, the mechanisms by which miR-15a and miR-16-1 genes exert their tumor suppressor function, and the clinical relevance of their expression in cancer.
Identification of miR-15a and miR-16-1 as Potential Cancer Genes
Many miRNAs have been suggested as playing a key role in cancer owing to their location and expression profiles.5, 6, 7 The first experimental evidence that miRNAs are involved in mammalian carcinogenesis was reported in CLL, the most common form of adult leukemia in the Western world.8 Genetic alterations, including hemizygous and homozygous loss, at 13q14.3 occur in more than half of CLL cases and constitute the most frequent chromosomal abnormality in CLL.17 Deletions at 13q14.3 have also been reported in other malignancies, such as multiple myeloma, mantle cell lymphoma, and prostate carcinoma.18 Together, these observations and others suggested that the13q14.3 genomic region harbors one or more tumor suppressor genes that are involved in the pathogenesis of some malignancies. Nonetheless, the identification of cancer genes involved in the 13q14.3 deletions in CLL failed, despite the extensive effort of several research groups. A region of more than 1.0 Mb has been fully sequenced and characterized in detail and identified several genes (Figure 1), but none of them were found to be involved in the initiation or progression of CLL or other human tumors.19 These findings led to the hypothesis that 13q14.3 is dispensable for CLL development and that deletions at this region do not confer any selective advantage to the malignant cells. Another possibility would be that haploinsufficiency itself predisposes to CLL.20 Alternatively, unknown genes, such as those spanning large genomic regions with huge introns or, possibly, extremely small genes perhaps not detected by classic cloning methods, might reside in this region and be the real target of genomic alteration.
Schematic diagram showing the 13q14.3 deletions in CLL cells targeting the miR-15a/miR-16-1 cluster. Eleven genes were cloned in the 13q14.3 region (upper panel), inside or very close to the deleted region. The lower panel shows the minimal region of deletion from two CLL patients,8 identified by cell hybrid experiments and containing the miR-15a/miR-16-1 cluster. The genomic markers and gene orientation are indicated. The gene symbols are as follows: DLEU7 (deleted in lymphocytic leukemia 7), DLEU1 (deleted in lymphocytic leukemia 1), DLEU2 (deleted in lymphocytic leukemia 2), miR-15a and miR-16-1 (microRNA genes 15a and 16-1), KCNRG (potassium channel regulator), DLEU5 (RFP2, ret finger protein 2), CLLD6 (C13orf1, chromosome 13 open reading frame 1), KPNA3 (karyopherin alpha 3, importin alpha 4), ARLTS1 (ARL11, ADP-ribosylation factor–like 11), CLLD7 (RCBTB1, regulator of chromosome condensation [RCC1] and BTB [POZ] domain containing protein 1), NY-REN-34 antigen (PHF11, PHD finger protein 11), and CLLD8 (SETDB2, SET domain, bifurcated 2)
To pursue this latter hypothesis and to identify putative tumor suppressor genes at 13q14.3, we completely sequenced and characterized a segment of 790 kb spanning the minimal region of loss,21 and we also generated somatic cell hybrids between mouse LM-TK− and CLL cells carrying 13q14.3 translocations and/or deletions.8 Subsequent analysis of an ∼30-kb deletion in one case of CLL associated with retinoblastoma (RB) and the chromosomal breakpoint translocation t(2:13)(q32;q14) in a different patient led to the identification of the critical region that contained two tightly linked miRNAs, miR-15a and miR-16-1 (Figure 1), involved in tumor suppression in CLL. Northern blotting and microarray analyses have shown that the miR-15a and miR-16-1 genes are ubiquitously and highly expressed in normal CD5+ lymphocytes,8 the cells from which CLL originates. These data suggested that these genes might play an important role in normal CD5+ B-cell homeostasis and that they could be the genomic targets of 13q14.3 in CLL (see below). It is worth noting that a second cluster including miR-15b/miR-16-2 is located on chromosome 3, though these genes are expressed at a much lower level in B cells and in CD5/CD19 positive cells as the chromosome 3q26.1 cluster.8 Further refined quantitative RT-PCR studies should investigate the specific expression of these two similar clusters in various human normal and pathologic states.
The identification of miR-15a and miR-16-1 as candidate cancer genes led to the question of how general the involvement of miRNAs is in human cancer. Numerous studies have used high-throughput microarrays to identify cancer-specific miRNA fingerprints in every type of cancer analyzed, including breast,22 hepatocellular,23, 24 lung,25, 26, 27 colon,28, 29 and gastric30 carcinomas. These studies clearly showed that the spectrum of miRNAs expressed in malignant cells is significantly different from that in healthy counterpart cells, suggesting that alterations in miRNA genes play a critical role in the pathophysiology of many, and perhaps all, human cancers.
Deregulation of miR-15a and miR-16-1 in Cancer
The importance of miRNAs in cancer, in general, is highlighted by the observation that half of all miRNA genes are located in cancer-associated genomic regions or fragile sites,6, 7 which are frequently altered in cancer. Global repression of miRNA processing machinery, including the enzymes Drosha, Pasha, and Dicer, promotes cellular transformation in mouse models, implicating the role of mature miRNA genes in cancer-related processes.31 Furthermore, large-scale profiling of miRNAs has revealed that miRNA deregulation is a common event in cancer (reviewed in Ref. 32). Although much is known about the aberrant expression of miRNAs in human cancer, much less is known about the functional relevance of such aberrations.
The mapping of miR-15a and miR-16-1 in a region that is frequently altered in cancer suggested that these two miRNAs could be the target genes of 13q14.3. Several studies of CLL (reviewed in Refs. 33, 34) and solid tumors35, 36 have shown that miR-15a and miR-16-1 are deleted or down-regulated in tumor cells, suggesting that these two miRNA genes could be ‘hot spots’ in cancer transformation.33 A recent analysis of miR-15a and miR-16-1 expression in prostate cancer revealed these genes’ consistent down-regulation in ∼80% of tumor samples compared with that in healthy tissues.35 No correlation between loss of the RB gene, which is located in the vicinity, and deregulation of miR-15a and miR-16-1 has been reported, indicating that the loss of miR-15a and miR-16-1 expression is often independent of the absence of the gene encoding RB. miR-15a and miR-16-1 have been also shown to express at lower levels in pituitary adenomas than in normal pituitary tissue.36 These data led to the hypothesis that miR-15a and miR-16-1 function as tumor suppressors and that their inactivation by allelic loss may contribute to tumorigenesis.
Abnormally expressed cancer genes are frequently targets for mutations that can activate or inactivate their function.37 Therefore, screening of mutations in miRNAs has been a critical step in determining the contribution of this new class of genes to cancer development and in identifying the genetic mechanisms responsible for their deregulation. Given that miR-15a and miR-16-1 have been suggested to act as tumor suppressors, it is possible that genetic mutations could inactivate their expression. Indeed, as we reported in one study, analysis of 75 CLL cases identified germline and somatic mutations in 5 of 42 sequenced miRNAs in 11 of 75 CLL patients, but no such mutations were found in 160 subjects without cancer (P<0.0001).38 Of note, a germline mutation in the pri– miR-16-1 sequence of two CLL patients—a C → T homozygous substitution, 7 bp in the 3′ direction after the precursor—have been found.38 This mutation has been found to reduce the expression of miR-16-1 in vivo, as CLL cells expressing these mutants display reduced expression of miR-16-1 compared with expression by normal CD5+ cells. Furthermore, ectopic expression of mutated miR-16-1 in vitro in HEK293 cells revealed reduced expression compared with that of a wild-type construct.38 Intriguingly, in the New Zealand Black mouse, which is characterized by spontaneously occurring late-onset CLL, Raveche et al.39 described a point mutation in 3′ DNA adjacent to the miR-16-1 region. This particular mutation resulted in the down-regulation of miR-16-1 expression in this CLL-prone mouse model, further indicating the tumor suppressor function of miR-16-1 in CLL. These observations indicate that mutations in the miR-16-1 gene could be responsible, at least in part, for its altered expression. Other mechanisms, including loss of heterozygosity, epigenetic regulation, and defects in the miRNA biogenesis machinery, could also contribute to the deregulation in miRNA expression.40
Given that CLL is a disease with a frequent familial association (10–20% of patients who have at least one first-degree relative with CLL) and that other cancers also occur frequently in persons with CLL,41 miRNA mutations may be a predisposing factor for cancers associated with CLL. In fact, one of the CLL patients harboring the germline C → T miR-16-1 germline mutation not only had been diagnosed earlier with breast cancer but also her mother died of CLL and her sister of breast cancer.38 Altogether, these findings may suggest that the miR-15a and miR-16-1 sequence could be a target of inactivation by point mutations that contribute to the initiation and/or progression of cancer.
miR-15a and miR-16-1 Function in Cancer Cells
miRNA profiling was instrumental in identifying the differential expression of these genes both in homeostasis and carcinogenesis.32 These profiling efforts led to the identification of miRNA oncogenes (oncomirs), referring to elevated levels of miRNAs in human tumor cells compared with normal cells. Oncomirs target the 3′ untranslated region (UTR), the 5′ UTR, or the coding sequence of mRNA of tumor suppressor genes and thus contribute to their down-regulation. On the other hand, up-regulation of oncogenes could be due to loss or reduced expression of miRNAs that function as tumor suppressors.42 After we found that miR-15a and miR-16-1 expression is lost or down-regulated in CLL cells, we hypothesized that these two genes may target the expression of a known oncogene or survival factor that is essential for CLL development.
Using bioinformatics tools, we found that the miR-15a and miR-16-1 sequences and the BCL2 mRNA sequence share a complementary homology, which suggests that the Bcl2 oncoprotein may be a target of posttranscriptional repression by miR-15a and miR-16-1 tumor suppressors.43 Bcl2 is a central player in the genetic program of eukaryotic cells, favoring survival by inhibiting cell death.44 Overexpression of the Bcl2 protein has been reported in many types of human cancers, including leukemias, lymphomas, and carcinomas.45 In follicular lymphomas and in a fraction of diffuse B-cell lymphomas, the mechanism of BCL2 activation has been found to be the translocation t(14;18)(q32;q21), which places the BCL2 gene under the control of immunoglobulin heavy-chain enhancers, resulting in deregulated expression of the gene.46, 47 B cell-CLL overexpress Bcl-248; however, with the exception of the fewer than 5% of cases in which BCL2 is juxtaposed to immunoglobulin loci,49 no mechanism has been discovered to explain BCL2 deregulation in CLL.
That the BCL2 gene is overexpressed in 65–70% of B cell-CLLs, and that deletions or down-regulation of the miR-15a and miR-16-1 cluster have been reported in the same proportion of CLL samples led us to hypothesize that miR-15a and miR-16-1 could be responsible for the increased levels of Bcl2 in CLL.43 Indeed, our data showed that miR-15a and miR-16-1 expression are inversely correlated with Bcl2 expression in CLL samples and that both miRNAs negatively regulate Bcl2 at a posttranscriptional level. Bcl2 repression by these miRNAs induced apoptosis in a leukemic cell line model. In MEG-01 cells transfected with the wild-type miR-15/16, we observed an increased rate of apoptosis mediated by cleavage of pro-caspase 9 and of poly (ADP-ribose) polymerase, indicating that the reduction in Bcl2 protein levels caused by miRNAs is sufficient to initiate the apoptotic process.43 Given that Bcl2 is overexpressed in the majority of CLL malignant lymphocytes and no genetic alteration can explain this aberrancy, control of Bcl2 expression by the miR-15/16 cluster can be considered as one of the main molecular mechanisms of this aberration.
To further shed light on the tumor suppressor action of miR-15a and miR-16-1 cluster, we analyzed the effects of both miRNAs on transcriptome and proteome in CLL cells.50 Ectopic expression of miR-15a and miR-16-1 led to the up-regulation of 265 genes and down-regulation of 3307 genes, among which the MCL1 anti-apoptotic gene and others are predicted targets of miR-15 and miR-16 (Table 1). Of note, ∼20% of the down-regulated genes had AU-rich elements (AREs), which could further support the finding that ARE-mediated instability is implicated in the regulation of gene expression by miR-16-1.51 Analysis of the proteomic changes after overexpression of miR-15a and miR-16-1 cluster revealed 27 different proteins that exhibited at least four-fold change with respect to the control group. Intriguingly, the proteins Bcl2 and Wilms tumor 1, a confirmed and predicted target, respectively, were among the proteins with prominent changes. Other proteins found are involved in cell growth, the cell-cycle, oncogenesis, tumor suppression, and anti-apoptosis.51 These observations clearly support the fact that a single miRNA, for example, miR-15a or miR-16-1, can have far-reaching effects on the overall function of the cell.
The tumor suppressor function of miR-15a/16-1 has also been addressed in vivo. In immunocompromised nude mice, ectopic expression of miR-15a/16-1 was found to cause dramatic suppression of tumorigenicity of MEG-01 leukemic cells exhibiting a loss of endogenous expression of miR-15a/16-1.50 This observation confirms that miR-15a and miR-16-1 genes are tumor suppressors and are highly effective in preventing the growth of leukemic xenografts.
The miR-15a/16-1 cluster targeting of BCL2 at the posttranscriptional level implies that miRNA genes play an important role in regulating apoptosis. Our data show that in cancer cells lacking these miRNAs, restoration of miR-15a and miR-16-1 triggers apoptosis and suppresses tumorigenicity.43 On the other hand, oncomirs such as miR21 and the miR17-92 cluster inhibit the expression of tumor suppressor genes such as PDCD452 and BIM,53 reducing apoptosis and enhancing proliferation (reviewed in Ref. 54) and thus contributing to tumorigenesis. Deregulation of apoptosis is an important step in cancer, as it allows genetically unstable cells to survive and accumulate further mutations that eventually lead to tumorigenesis. Therefore, it is possible that perturbation of the expression of oncosuppressor miRNA genes, such as miR-15a and miR-16-1, may lead to tumorigenesis.
An miRNA gene could have several targets; therefore, it is likely that miR-15a and miR-16-1 may target other oncogenes that are relevant to cancer. Recently, Bonci et al.35 reported that the miR-15a and miR-16-1 cluster targets not only BCL2 but also CCND1 (encoding cyclin D1) and WNT3A mRNA, which promote several prostate tumorigenic features, including survival, proliferation, and invasion. In vivo knockdown of miR-15a and miR-16-1 has been shown to result in prostate hypoplasia associated with Ccnd1 and Wnt3a up-regulation, indicating that the loss of these miRNAs could be a relevant pathogenic event in prostate carcinogenesis.35 Although BCL2, CCND1, and WNT3A may play a key role in various types of human cancer driven by loss of miR-15a and miR-16-1 expression, there could be hundreds of predicted targets for these miRNAs that might affect the process of carcinogenesis. Together, these data suggest that miR-15a/16-1 genes are natural anti-sense interactors with BCL2 and probably other oncogenes and that they can be used to suppress tumor growth (Figure 2).
Genetic alteration in miR-15a/16-1 cluster triggers apoptosis, proliferation, and tumor progression. The main mechanisms of miR-15/16 alteration are deletion, down-regulation, and mutation. Experimentally proven targets of miR-15a and miR-16-1 are shown: the BCL2 antiapoptotic gene, cyclin D1 (CCND1), and the WNT gene WNT3A. In tumor cells, miR-15/16 genes are altered, which affects the expression of their targets and leads to enhanced survival, increased proliferation, and tumor progression. Restoration of miR-15/16 genes in tumor cells should inhibit expression of these oncogenes, resulting in caspase activation, apoptosis, and tumor regression
Deregulation of miR-15a and miR-16-1 in Cancer is Associated with Clinical Parameters
In addition to their role in promoting the development of primary tumors, miRNAs might also guide the choice of the best treatment strategy by providing prognostic information. CLL is characterized by the progressive accumulation of mostly nondividing CD5+ B lymphocytes.17, 41 Several clinical parameters are used to determine prognosis and the therapeutic course in CLL. For example, few or no mutations in the immunoglobulin heavy-chain variable-region gene (IgVH) or high expression of the 70-kD zeta-associated protein (ZAP-70+) predict an aggressive course, whereas mutated IgVH CLL clones or low ZAP-70+ expression predicts an indolent course.17 Analysis of miRNA expression profiles of 94 samples of CLL cells defined a unique miRNA expression signature composed of 13 genes (of 190 analyzed) that can differentiate between low and high ZAP-70 expression and between unmutated and mutated IgVH.38 The corresponding miRNA signature was associated with the presence or absence of disease progression. Given that allelic loss of 13q14.3 has been shown to be associated with a favorable outcome in B-cell–associated CLL, we speculated that deletion or down-regulation of miR-15a and miR-16-1 could also be associated with improved prognosis. In fact, deregulation of miR-15a and miR-16-1 in CLL has been correlated with a better prognosis.38
Lower levels of miR-15a and miR-16-1 have also been reported in pituitary adenomas, and their expression was inversely correlated with tumor diameter and tumor growth.36 Bonci et al.35 also noted that deletion of miR-15a and miR-16-1 genes was associated with the presence of more-advanced prostate tumors. Therefore, accumulating evidence suggests that the miRNA expression profile not only is different between malignant cells and their normal counterparts but also is associated with prognostic factors in human cancer.
Future Perspectives
We envision the possible future use of miRNAs or compounds interacting with miRNAs as new therapeutic agents in cancer. Compared with other RNA inhibition techniques (such as anti-sense oligonucleotides (ASOs), ribozymes, and siRNAs), the major advantage of using miRNAs to treat cancer patients is that a specific miRNA targets several genes involved in the same pathway. For example, among the components of the miR-15a/16-1 signature in CLL, a disease in which the main altered cell program is apoptosis, we observed significant enrichment in cancer genes (such as MCL1, BCL2, ETS1, and PDCD6IP) (Table 1) that directly or indirectly affect apoptosis and the cell cycle. Therefore, by targeting miRNAs, a more effective functional restoration of the abnormal pathway can be achieved than can be achieved with the usual one-agent, one-target effect characteristic of all other RNA inhibition agents. Given recently published results showing the efficacy of oblimersen sodium in patients with advanced CLL that is refractory to fludarabine,55 it would be feasible to design regimens using a cocktail of anti-BCL2 ASOs and miRNAs targeting BCL2, such as miR-15 and miR-16, for indolent CLL or of anti-MCL1 miRNAs, such as miR-29 and miR181, for aggressive forms of CLL. In this way, multiple drugs targeting the anti-apoptotic overexpressed regulators Bcl2 and Mcl1 could have a better chance of consistently and robustly reducing these proteins’ expression levels. Given that many more miRNAs have been predicted in silico and will probably be proven experimentally to interact with BCL2, the possible combinations of RNA inhibitory drugs will increase in the near future.
Abbreviations
- CLL:
-
chronic lymphocytic leukemia
- miRNAs:
-
MicroRNAs
- UTR:
-
untranslated region
References
Ambros V . MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 2003; 113: 673–676.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Ambros V . The functions of animal microRNAs. Nature 2004; 431: 350–355.
He L, Hannon GJ . MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 2004; 5: 522–531.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–866.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 2004; 101: 2999–3004.
Sevignani C, Calin GA, Siracusa LD, Croce CM . Mammalian microRNAs: a small world for fine-tuning gene expression. Mamm Genome 2006; 17: 189–202.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. Frequent deletions and down-regulation of micro- RNA genes miR-15 and miR-16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Liu CG, Calin GA, Meloon B, Gamliel N, Sevignani C, Ferracin M et al. An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues. Proc Natl Acad Sci USA 2004; 101: 9740–9744.
Chan JA, Krichevsky AM, Kosik KS . MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 2005; 65: 6029–6033.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–838.
Jiang J, Lee EJ, Gusev Y, Schmittgen TD . Real-time expression profiling of microRNA precursors in human cancer cell lines. Nucleic Acids Res 2005; 33: 5394–5403.
Schmittgen TD, Jiang J, Liu Q, Yang L . A high-throughput method to monitor the expression of microRNA precursors. Nucleic Acids Res 2004; 32: e43.
Liu CG, Calin GA, Volinia S, Croce CM . MicroRNA expression profiling using microarrays. Nat Protoc 2008; 3: 563–578.
Ventura A, Jacks T . MicroRNAs and cancer: short RNAs go a long way. Cell 2009; 136: 586–591.
Nicoloso MS, Spizzo R, Shimizu M, Rossi S, Calin GA . MicroRNAs—the micro steering wheel of tumour metastases. Nat Rev Cancer 2009; 9: 293–302.
Chiorazzi N, Rai KR, Ferrarini M . Chronic lymphocytic leukemia. N Engl J Med 2005; 352: 804–815.
Dong JT, Boyd JC, Frierson Jr HF . Loss of heterozygosity at 13q14 and 13q21 in high grade, high stage prostate cancer. Prostate 2001; 49: 166–171.
Calin GA, Trapasso F, Shimizu M, Dumitru CD, Yendamuri S, Godwin AK et al. Familial cancer associated with a polymorphism in ARLTS1. N Engl J Med 2005; 352: 1667–1676.
Migliazza A, Bosch F, Komatsu H, Cayanis E, Martinotti S, Toniato E et al. Nucleotide sequence, transcription map, and mutation analysis of the 13q14 chromosomal region deleted in B-cell chronic lymphocytic leukemia. Blood 2001; 97: 2098–2104.
Bullrich F, Fujii H, Calin G, Mabuchi H, Negrini M, Pekarsky Y et al. Characterization of the 13q14 tumor suppressor locus in CLL: identification of ALT1, an alternative splice variant of the LEU2 gene. Cancer Res 2001; 61: 6640–6648.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005; 65: 7065–7070.
Wang Y, Lee AT, Ma JZ, Wang J, Ren J, Yang Y et al. Profiling microRNA expression in hepatocellular carcinoma reveals microRNA-224 up-regulation and apoptosis inhibitor-5 as a microRNA-224-specific target. J Biol Chem 2008; 283: 13205–13215.
Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T et al. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene 2006; 25: 2537–2545.
Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S et al. A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 2005; 65: 9628–9632.
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006; 9: 189–198.
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004; 64: 3753–3756.
Michael MZ, SM OC, van Holst Pellekaan NG, Young GP, James RJ . Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 2003; 1: 882–891.
Schetter AJ, Leung SY, Sohn JJ, Zanetti KA, Bowman ED, Yanaihara N et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 2008; 299: 425–436.
Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I et al. E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 2008; 13: 272–286.
Kumar MS, Lu J, Mercer KL, Golub TR, Jacks T . Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 2007; 39: 673–677.
Barbarotto E, Schmittgen TD, Calin GA . MicroRNAs and cancer: profile, profile, profile. Int J Cancer 2008; 122: 969–977.
Calin GA, Croce CM . Genomics of chronic lymphocytic leukemia microRNAs as new players with clinical significance. Semin Oncol 2006; 33: 167–173.
Nicoloso MS, Kipps TJ, Croce CM, Calin GA . MicroRNAs in the pathogeny of chronic lymphocytic leukaemia. Br J Haematol 2007; 139: 709–716.
Bonci D, Coppola V, Musumeci M, Addario A, Giuffrida R, Memeo L et al. The miR-15a-miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat Med 2008; 14: 1271–1277.
Bottoni A, Piccin D, Tagliati F, Luchin A, Zatelli MC, degli Uberti EC . miR-15a and miR-16-1 down-regulation in pituitary adenomas. J Cell Physiol 2005; 204: 280–285.
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE et al. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 2005; 353: 1793–1801.
Raveche ES, Salerno E, Scaglione BJ, Manohar V, Abbasi F, Lin YC et al. Abnormal microRNA-16 locus with synteny to human 13q14 linked to CLL in NZB mice. Blood 2007; 109: 5079–5086.
Deng S, Calin GA, Croce CM, Coukos G, Zhang L . Mechanisms of microRNA deregulation in human cancer. Cell Cycle 2008; 7: 2643–2646.
Catovsky D . Definition and diagnosis of sporadic and familial chronic lymphocytic leukemia. Hematol Oncol Clin North Am 2004; 18: 783–794, vii.
Calin GA, Croce CM . MicroRNA-cancer connection: the beginning of a new tale. Cancer Res 2006; 66: 7390–7394.
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005; 102: 13944–13949.
Cory S, Adams JM . The Bcl2 family: regulators of the cellular life-or-death switch. Nat Rev Cancer 2002; 2: 647–656.
Sanchez-Beato M, Sanchez-Aguilera A, Piris MA . Cell cycle deregulation in B-cell lymphomas. Blood 2003; 101: 1220–1235.
Tsujimoto Y, Cossman J, Jaffe E, Croce CM . Involvement of the bcl-2 gene in human follicular lymphoma. Science 1985; 228: 1440–1443.
Tsujimoto Y, Yunis J, Onorato-Showe L, Erikson J, Nowell PC, Croce CM . Molecular cloning of the chromosomal breakpoint of B-cell lymphomas and leukemias with the t(11;14) chromosome translocation. Science 1984; 224: 1403–1406.
Kitada S, Andersen J, Akar S, Zapata JM, Takayama S, Krajewski S et al. Expression of apoptosis-regulating proteins in chronic lymphocytic leukemia: correlations with in vitro and in vivo chemoresponses. Blood 1998; 91: 3379–3389.
Adachi M, Tefferi A, Greipp PR, Kipps TJ, Tsujimoto Y . Preferential linkage of bcl-2 to immunoglobulin light chain gene in chronic lymphocytic leukemia. J Exp Med 1990; 171: 559–564.
Calin GA, Cimmino A, Fabbri M, Ferracin M, Wojcik SE, Shimizu M et al. MiR-15a and miR-16-1 cluster functions in human leukemia. Proc Natl Acad Sci USA 2008; 105: 5166–5171.
Jing Q, Huang S, Guth S, Zarubin T, Motoyama A, Chen J et al. Involvement of microRNA in AU-rich element-mediated mRNA instability. Cell 2005; 120: 623–634.
Lu Z, Liu M, Stribinskis V, Klinge CM, Ramos KS, Colburn NH et al. MicroRNA-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 2008; 27: 4373–4379.
Xiao C, Srinivasan L, Calado DP, Patterson HC, Zhang B, Wang J et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol 2008; 9: 405–414.
Wang Y, Lee CG . MicroRNA and cancer—focus on apoptosis. J Cell Mol Med 2009; 13: 12–23.
O’Brien SM, Cunningham CC, Golenkov AK, Turkina AG, Novick SC, Rai KR . Phase I to II multicenter study of oblimersen sodium, a Bcl-2 antisense oligonucleotide, in patients with advanced chronic lymphocytic leukemia. J Clin Oncol 2005; 23: 7697–7702.
Acknowledgements
RIA is supported by a Ma’of fellowship and NIH R01DK079217 grant. GAC is supported as a Fellow by The University of Texas M. D. Anderson Research Trust, as a Fellow of The University of Texas System Regents Research Scholar, and by the Ladjevardian Regents Research Scholar Fund. Work in CMC's laboratory is funded by the National Institutes of Health. The authors thank Virginia M Mohlere for the editing work on this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Aqeilan, R., Calin, G. & Croce, C. miR-15a and miR-16-1 in cancer: discovery, function and future perspectives. Cell Death Differ 17, 215–220 (2010). https://doi.org/10.1038/cdd.2009.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.69
Keywords
This article is cited by
-
Nutrient restriction-activated Fra-2 promotes tumor progression via IGF1R in miR-15a downmodulated pancreatic ductal adenocarcinoma
Signal Transduction and Targeted Therapy (2024)
-
Recent insights into the microRNA-dependent modulation of gliomas from pathogenesis to diagnosis and treatment
Cellular & Molecular Biology Letters (2022)
-
Factors influencing degradation kinetics of mRNAs and half-lives of microRNAs, circRNAs, lncRNAs in blood in vitro using quantitative PCR
Scientific Reports (2022)
-
Dynamical modeling of miR-34a, miR-449a, and miR-16 reveals numerous DDR signaling pathways regulating senescence, autophagy, and apoptosis in HeLa cells
Scientific Reports (2022)
-
Transcriptional and post-transcriptional regulation of checkpoint genes on the tumour side of the immunological synapse
Heredity (2022)