Abstract
Multiple oncogenes (in particular phosphatidylinositol 3-kinase, PI3K; activated Akt1; antiapoptotic proteins from the Bcl-2 family) inhibit autophagy. Similarly, several tumor suppressor proteins (such as BH3-only proteins; death-associated protein kinase-1, DAPK1; the phosphatase that antagonizes PI3K, PTEN; tuberous sclerosic complex 1 and 2, TSC1 and TSC2; as well as LKB1/STK11) induce autophagy, meaning that their loss reduces autophagy. Beclin-1, which is required for autophagy induction acts as a haploinsufficient tumor suppressor protein, and other essential autophagy mediators (such as Atg4c, UVRAG and Bif-1) are bona fide oncosuppressors. One of the central tumor suppressor proteins, p53 exerts an ambiguous function in the regulation of autophagy. Within the nucleus, p53 can act as an autophagy-inducing transcription factor. Within the cytoplasm, p53 exerts a tonic autophagy-inhibitory function, and its degradation is actually required for the induction of autophagy. The role of autophagy in oncogenesis and anticancer therapy is contradictory. Chronic suppression of autophagy may stimulate oncogenesis. However, once a tumor is formed, autophagy inhibition may be a therapeutic goal for radiosensitization and chemosensitization. Altogether, the current state-of-the art suggests a complex relationship between cancer and deregulated autophagy that must be disentangled by further in-depth investigation.
Similar content being viewed by others
Main
Autophagy constitutes one of the major responses of cells to external or internal stimuli. As any other major phenomenon of cell biology (such as division, differentiation and cell death), autophagy can be perturbed in cancer cells and is modulated by anticancer chemotherapies. The first functional link that has been established between autophagy and cancer originates from the landmark discovery that Beclin 1, a phylogenetically conserved protein that is essential for macroautophagy, is also a haploinsufficient tumor suppressor.1 On the basis of this discovery and the observation that multiple stress signals including DNA damage can induce macroautophagy2, 3, 4 both fundamental and clinical oncologists have become aware of the possible importance of ‘self-eating’ for oncogenesis and therapeutic responses, thus launching a new area of febrile investigation.
Any biomedical investigator and clinical oncologist can enumerate the six cell-intrinsic hallmarks of cancer, as formulated by Hanahan and Weinberg,5 provision of autonomous growth signals, insensitivity to negative signals, limitless replication, production of angiogenic factors and tissue invasion with metastasis, as well as disabled apoptosis. In this context, it has been well established that prominent oncogenes reduce the propensity of cancer cells to undergo cell death, whereas prominent tumor suppressor proteins have proapoptotic properties. However, the link between autophagy and cell death is highly ambiguous, and autophagy may serve as a mechanism of adaptation to stress (and hence avoidance of cell death), as well as, in special circumstances, as a route to cell death6, 7, 8, 9, 10 suggesting that the relationship between cancer and autophagy may be more complex than that between cancer and cell death.
The present review deals with autophagy regulation by oncogenes and tumor supporessor proteins and the contribution of deregulated autophagy may contribute to oncogenesis and tumor progression.
Regulation of Autophagy by the PI3K/Akt/mTOR Pathway
In cancer, essential signal-transducing molecules involved in growth signaling often are constitutively switched on due to activating mutation or due to the continuous presence of autocrine growth factors. Growth factors activate receptor tyrosine kinases (RTKs), which then stimulate two key signal transducing components: the small GTPase Ras and the phosphatidylinositol 3-kinase (PI3K). Ras and PI3K converge to activate mTOR for stimulating cell growth and for inhibiting autophagy. Most cancers harbor activating mutations of the master regulators (K-ras, H-Ras, N-ras, the p110a PI3K subunit and RTKs) or their downstream effectors (such as the kinases Akt and PDK1), or inactivating mutations in negative regulators of these proteins.11 Germline mutation of such negative regulators, namely the Ras antagonist neurofibromin 1 (NF1), the mTOR inhibitors tuberous sclerosis complex 1 and 2 (TSC1, TSC2), LKB1 (a kinase that activates the AMP-activated protein kinase, AMPK) or the phosphatase PTEN, which reverses the phosphorylation mediated by PI3K, results in familial cancer syndromes. These pathologies share clinical features (phakomatoses or ‘neurocutaneous syndromes’), characterized by hamartoma-type tumors (Peutz-Jegher's syndrome, tuberous sclerosis, Cowden's disease and neurofibromatosis type 1).11
As a result, many cancers exhibit an enhanced activation of mTOR, resulting into reduced autophagy. Elegant in vivo experiments performed by White and colleagues4 demonstrate that constitutively active Akt inhibits the induction of autophagy in vitro and in vivo, whereas accelerating the growth of tumors in vivo, in particular in cells that lack the expression of Bax and Bak. This enhanced tumor growth could be correlated with an elevated level of tumor cell necrosis and local inflammation, as well as a marked genomic instability.12 An important autophagy-regulatory gene, such as Beclin 1, functions as a haploinsufficient tumor suppressor gene. Deletion of one allele of Beclin-1 accelerates tumor growth in vivo, in the context of Bcl-2 overexpression.12 In addition, monoallelic knockout of beclin-1 or biallelic knockout of atg5 had similar effects on genomic instability as constitutively active Akt.13, 14 It appears plausible that the inhibition of autophagy is a major contributing factor in the oncogenic action of a constitutively active PI3K/Akt axis. Formal proof in favor of this possibility is however still elusive.
Regulation of Autophagy by Bcl-2 Family Proteins
Antiapoptotic proteins from the Bcl-2 family are prominent oncogenes, whereas proapoptotic proteins from this family are tumor suppressor proteins. Although Bcl-2 family proteins have been initially characterized as cell death regulators, it has recently become clear that they control the autophagic process as well. Thus, antiapoptotic proteins such as Bcl-2,15 Bcl-XL,16 Bcl-w17 and Mcl-116 can inhibit autophagy. Accordingly, the knockdown of Bcl-2 with antisense oligonucleotides18 or small interfering RNA (siRNA) heteroduplexes19 can induced autophagy in a range of different cell lines including leukemia and breast cancer cells. Conversely, proapoptotic BH3-only proteins from the Bcl-2 family, such as BNIP3L,20, 21 Bad,16 Noxa, Puma, BimEL22 and Bik,23 stimulate autophagy.
Beclin 1/Atg6 has initially been identified as a protein that interacts with Bcl-2.1 Today, it is known that the essential autophagy protein Atg6/Beclin 1 cooperates with several cofactors (including Ambra-1, Bif-1 and UVRAG) to activate the lipid kinase Vps34, thereby inducing autophagy.24, 25, 26, 27 In baseline conditions, Beclin 1 is bound to and inhibited by Bcl-2 or other antiapoptotic multidomain proteins of the Bcl-2 family including Bcl-XL,16 Bcl-w17 and Mcl-1.16 At the structural level, it is important to note that antiapoptotic multidomain proteins possess a hydrophobic cleft, the so-called BH3 receptor domain, which can accommodate BH3 domains from proapoptotic Bcl-2 protein family members. Such BH3 domains have an amphipathic α-helical structure.28
Three different approaches have proven that Beclin 1 possesses a BH3 motif (aa 112–123). First, crystal structures have been obtained from recombinant Bcl-XL protein complexed to a synthetic peptide that corresponds to the Bcl-2/Bcl-XL-binding domain of Beclin 1. Similarly to previously determined Bcl-XL–BH3 structures, the amphipathic BH3 helix of Beclin 1 binds to the BH3 receptor domain of Bcl-XL.29 Second, the X-crystallographic data have been confirmed by NMR spectroscopy.30 Third, replacements of several critical amino acids within the putative BH3-only domain of Beclin 1 (L116A, L116E, L116Q, G120E, D121A and F123A) reduce or abrogate the molecular interaction between Beclin 1 and Bcl-XL in GST pulldown, analytical gel filtration, fluorescence anisotropy measurements and/or coimmunoprecipitation assays.15, 16, 29, 30, 31 Altogether, these three technologies (X-crystallography, NMR and mutational analysis) convincingly demonstrate that Beclin 1 possesses a BH3 domain, if such domains are defined by their capacity to interact with the BH3 receptor domain of the antiapoptotic Bcl-2 proteins. It should be noted, however, that the overexpression of Beclin 1 cannot induce cell death (MCM and GK, unpublished observation), indicating that Beclin 1, in contrast to other BH3-only proteins, has no proapoptotic properties. Moreover, the incidence of tumors in mice carrying a knock-in mutation that modulates the BH3 domain of Beclin 1 has not been determined. Thus, the putative antioncogenic function of the BH3 domain of Beclin 1 remains elusive. In conclusion, the interaction between Beclin-1 and antiapoptotic proteins of the Bcl-2 family (in particular Bcl-2 and Bcl-XL) involves the Bcl-2 homology 3 (BH3) domain in Beclin-1 and the BH3 receptor domain of Bcl-2/Bcl-XL. Other proteins carrying BH3 domains can competitively disrupt the interaction between Beclin-1 and Bcl-2/Bcl-XL to induce autophagy.31 Starvation from essential nutrients, which is the physiologically most potent inducer of autophagy, stimulates the dissociation of Beclin-1 from its inhibitors, presumably by activating the BH3-only protein, Bad.16 The BH3-mimetic compound, ABT737, an agent designed to occupy the BH3 receptor domain of Bcl-2 or Bcl-XL (but not that of Mcl-1),32 also competitively disrupts the physical interaction between Beclin 1 and Bcl-2 or Bcl-XL.31 ABT737 pretreatment abolishes the immunoprecipitation between Beclin 1 and Bcl-2 or Bcl-XL (but not Mcl-1) in cells that are resistant to the proapoptotic action of ABT737. This effect correlates with the induction of massive autophagy. ABT737-induced autophagy cannot be inhibited by Bcl-2 or Bcl-XL overexpression, yet is abolished by transfection with Mcl-1 or by the siRNA-mediated knockdown of Beclin 1.16 These results demonstrate that autophagy is efficiently triggered by the disruption of the interaction between Beclin 1 and Bcl-2 or Bcl-XL.
Interestingly, ABT737 induces the autophagic sequestration of mitochondria (mitophagy) more efficiently than that of endoplasmic reticulum (ER; reticulophagy).13 Similarly, one particular BH3-only protein, BNIP3L, has been found to be required for the autophagic elimination of mitochondria that accompanies terminal erythroid differentiation.33 Mitophagy induced by hypoxia occurs through the HIF-1-dependent transcriptional activation of BNIP3L, which disrupts the interaction between Bcl-2 and Beclin.34 These results point to the possibility that interactions between Bcl-2 family proteins and Beclin 1 have a particularly important function in the regulation of mitophagy.
Regulation of Autophagy by p53
The tumor suppressor protein p53 is often inactivated in tumor cells, for instance due to mutations of p53 itself, due to mutations of the kinases that lead to its activation (such as ATM or Chk1) or due to the amplification of MDM2, the E3 ubiquitin ligase that targets p53 to proteasome-mediated destruction.35, 36 Hence, inactivation of the p53 system is one of the most frequent alterations that occur in cancer. p53 is mainly viewed as a transcription factor that transactivates proapoptotic and cell cycle-arresting genes,36 thereby favoring apoptosis and senescence of cancer cell precursors (which explains its tumor-suppressive effects as a ‘guardian of the genome’) or of cancer cells that respond to chemotherapy or radiotherapy (which explains its positive impact on anticancer treatment). p53 can also transactivate an autophagy-inducing gene, dram, which codes for a lysosomal protein,37 and p53-dependent induction of autophagy has been documented by several groups in response to DNA damage,38 Arf activation,39 or reexpression of p53 in p53-negative tumor cells.40
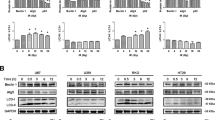
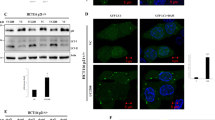
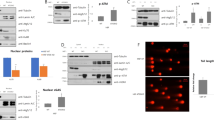
Recently, we observed that inactivation of p53 by deletion, depletion or inhibition also can trigger autophagy.41 Thus, human and mouse cells subjected to knockout, knockdown or pharmacological inhibition of p53 manifest signs of autophagy, such as depletion of p62/SQSTM1, LC3 lipidation (and hence conversion of LC3-I into LC3-II), redistribution of GFP-LC3 in cytoplasmic puncta and electron microscopic evidence of autophagosomes and autolysosomes,42 both in vitro and in vivo.41 This applies to a variety of methods for p53 inactivation: chemical inhibition with cyclic pifithrin-α (PFT-α), knockdown with siRNAs specific for human p53, mouse p53 or the Caenorhabditis elegans p53 ortholog cep1 or homologous recombination of p53 in human cancer cells, mice or nematodes. p53 inactivation was found to induce autophagy in several nontransformed or malignant human cell lines, namely HFFF2 fibroblasts, HCT116 colon cancer cells, SH-SY5Y neuroblastoma and HeLa cervical cancer cells, in mouse embryonic fibroblasts (MEF), in vivo in multiple mouse tissues (kidney, pancreas, liver, brain and heart), as well as in C. elegans embryos and adult pharyngeal cells.41 Electron microscopy and immunofluorescence experiments indicate that p53 inactivation induces both autophagy of the ER (reticulophagy) and mitochondria (mitophagy). When p53 was inhibited in an acute fashion by addition of PFT-α, reticulophagy was induced more rapidly than mitophagy, suggesting an intimate relationship between p53 inhibition and ER stress (which also induces preferential reticulophagy). Accordingly, p53 triggered the phosphorylation of eIF2α, which is a hallmark of ER stress. Moreover, the knockdown or knockout of IRE1, one of the quintessential ER stress effectors, reduced autophagy induction by p53 inactivation.41
Inhibition of p53 caused autophagy in enucleated cells, indicating that the cytoplasmic, nonnuclear pool of p53 can regulate autophagy. We also observed that retransfection of p53−/− cells with wild-type (WT) p53 suppressed autophagy and that this effect could be mimicked by a p53 mutant that is excluded from the nucleus, due to the deletion of the nuclear localization sequence. In contrast, retransfection of p53−/− cells with a nucleus-restricted p53 mutant (in which the nuclear localization sequence has been deleted) failed to inhibit autophagy. Hence, cytoplasmic (but not nuclear) p53 is responsible for the inhibition of autophagy. Several distinct autophagy inducers (e.g., nutrient depletion or addition of rapamycin, lithium, tunicamycin or thapsigargin) stimulated the rapid degradation of p53. The p53 protein was depleted both from the nucleus and from the cytoplasm, with similar kinetics. Inhibition of the p53-specific E3 ubiquitin ligase MDM2 (by two distinct pharmacological inhibitors or by knockdown) avoided p53 depletion and simultaneously prevented the activation of autophagy. Finally, a p53 mutant that lacks the MDM2 ubiquitinylation site and hence is more stable than WT p53 was particularly efficient in inhibiting autophagy.41
In conclusion, p53 has a dual function in the control of autophagy. On the one hand, nuclear p53 can induce autophagy through transcriptional effects. On the other hand, cytoplasmic p53 may act as a master repressor of autophagy. How this latter effect is achieved in mechanistic terms is not clear yet (Figure 1).
Cytoplasmic versus nuclear effects of p53. p53 has a Janus role in the control of autophagy. Nuclear p53 promotes the transcription of proapoptotic and cell cycle-arresting genes, and also can act as an autophagy-inducing transcription factor. In contrast, cytoplasmic p53 degradation exerts an autophagy-inhibitory function. Both of the p53 ‘faces’ are not completely known at molecular level
Regulation of Autophagy by DAPK
The death-associated protein kinase-1 (DAPK-1) is commonly silenced in human cancers by methylation, and has tumor and metastasis suppressor properties.43 DAPK-1 also induces autophagy and apoptosis, through independent mechanisms. DAPK-1 interacts with the microtubule-associated protein MAP1B, in particular in conditions of amino-acid starvation. Autophagy induced by DAPK-1 overexpression is attenuated by knockdown of MAP1B,44 suggesting that the interaction of DAPK1 with MAP1B (which in turn is capable of interacting with LC3) might be responsible for the induction of autophagy. MAP1B has been shown to inhibit autophagy in previous studies.45 Future investigation will clarify whether MAP1B is the sole route that links DAPK1 to autophagy. Indeed, DAPK1 is an extremely pleiotropic molecule43 and can activate the p53 system, suggesting that transcriptional programs influenced by DAPK1 might also affect the propensity of cells to undergo autophagy.
Deficient Autophagy as a Mechanism of Oncogenesis
Beclin 1 maps to a tumor susceptibility locus that is monoallelically deleted in a high percentage of human breast, ovarian and prostate cancers24 (Table 1). Decreased expression of Beclin 1 has been observed in human breast, ovarian and brain tumors.24, 50 The monoallelic deletions of beclin 1 in human cancer likely contribute tumorigenesis, as targeted mutant mice with heterozygous disruption of beclin 1 have decreased autophagy, spontaneously develop tumors including lymphomas, lung carcinomas, hepatocellular carcinomas and mammary precancerous lesions as they age, and undergo accelerated hepatitis B virus-induced carcinogenesis.51, 52 Moreover, immortalized kidney and mammary epithelial cells derived from beclin 1 heterozygous-deficient mice are more aggressive than control cells derived from WT mice when they are inoculated into immunodeficient recipient mice.13, 14 Other proteins that form part of the Beclin 1/class III PI3K complex, such as UVRAG and Bif1 may also act as tumor suppressors. UVRAG is monoallelically deleted at a high frequency in human colon cancers and, when overexpressed, suppresses the proliferation and tumorigenicity of human colon cancer cells.24 The deletion of Bif1 in mice results in the development of spontaneous tumors. Finally, knockout of atg4c, a cysteine protease involved in the processing of LC3/Atg8, accelerates the development of methyl-cholanthrene-induced fibrosarcomas in mice53 (Table 2). This suggests that tumor suppression may be a shared property of several distinct autophagy proteins that act at different steps in the pathway.
White and colleagues.4 have proposed two alternative (and nonexclusive) hypotheses to explain how the loss of autophagy may stimulate oncogenesis in spite of the fact that the suppression of autophagy enhances the frequency of necrotic cell deaths within tumors. First, enhanced necrosis due to suppressed autophagy might exacerbate local inflammation and thereby stimulate tumor growth.12 Second, invalidated autophagy may promote genomic instability in metabolically stressed cells, leading to oncogene activation and tumor progression.13, 14 Indeed, immortalized mouse epithelial cells that exhibit a genetically determined defect in autophagy due to monoallelic loss of beclin 1 or biallelic loss of atg5 display increased DNA damage foci, centrosome abnormalities, numerical and structural chromosomal abnormalities and gene amplification, especially after ischemic stress. The mechanisms through which deficient autophagy may induce genomic instability are elusive. As a possibility, autophagy might simply be required for an optimal management of the bioenergetic flux, thus facilitating the minimum amount of ATP levels (and that of other metabolites) required for DNA repair.8 It is also possible that autophagy indirectly contributes to the enforcement of cell-cycle checkpoints by degrading specific organelles or proteins that are essential for cell growth regulation.41 Finally, autophagy may be required to remove damaged organelles (such as mitochondria) and hence to regulate the levels of endogenous stress (including ROS production by uncoupled mitochondria),26 meaning that deficient autophagy might affect genomic instability in a rather indirect fashion (Figure 2a).
Autophagy in oncogenesis and anticancer therapy. The role of autophagy in oncogenesis is manifold. At the early stages of tumor, autophagy acts as a tumor suppressor. During tumor progression, autophagy can contribute to tumor growth. Further studies are necessary to elucidate the molecular mechanisms of this switch (a). Autophagy as a response to cancer therapy can promote/suppress tumor (b). Autophagy can be considered as an unusual target to improve anticancer therapy (c)
Enhanced Autophagy as a Mechanism of Tumor Cell Survival
A large series of clinically approved and experimental anticancer therapies induce the accumulation of autophagosomes in tumor cell lines in vitro.7 For many years, it was thought that these therapies kill cells through autophagy and hence induce the so called ‘autophagic cell death’. However, specific inhibition of autophagy with siRNA's targeted against ATG genes usually accelerates, rather than prevents, cell death in these settings,7 indicating that autophagy activation likewise represents a cellular attempt to cope with stress induced by cytotoxic agents.26 Along these lines, inhibition of autophagy can be used for experimental cancer treatment. For example, in mice harboring c-Myc-induced lymphomas, the lysosomotropic drug chloroquine enhanced the efficacy of either p53 or a DNA alkylating agent to induce tumor cell death and tumor regression.40
Inhibition of autophagy by siRNAs that target essential autophagy genes also sensitizes cancer cells to the induction of cell death by radiotherapy54 and a wide range of chemotherapeutic agents including cyclophosphamide40 and N-(4-hydroxypheny)retinamide.55 Inhibition of autophagy also sensitizes breast cancer cells to killing by the estrogen receptor antagonist tamoxifen,56 prostate cancer cells to androgen deprivation,57 colon cancer cells to amino-acid or glucose deprivation,58 Bax−/− HCT116 cells to TRAIL-induced apoptosis59 and multiple different cell lines to anoikis (apoptosis due to detachment from the extracellular matrix),60 suggesting that suppression of autophagy could constitute a general strategy for rendering cancer cells more susceptible to the induction of cell death (Figure 2b and c).
Conclusions and Perspectives
As indicated in the introduction to this paper, cancer is characterized by six well-established hallmarks. There have been attempts to identify additional hallmarks, such as avoidance of immune control61 or increased anabolic reactions.62 As discussed here, however, the relationship between cancer and autophagy cannot be reduced to a simple formulation. As it stands, reduced autophagy maybe an oncogenic event and contribute to tumor progression, whereas enhanced autophagy may rather constitute a mechanisms through which tumor cells survive hypoxic, metabolic, detachment-induced or therapeutic stress.
Induction of autophagy, as it occurs in response to starvation, is responsible for the beneficial effects of caloric restriction on longevity, at least in C. elegans.63 It is tempting to speculate that periodic induction of autophagy may also be responsible for the cancer preventive effect of caloric restriction. If this were true, pharmacological induction of autophagy might be used for cancer chemoprevention. Therapeutic inhibition of autophagy, if feasible, might enhance the efficacy of radiotherapy or chemotherapy, yet might have an intrinsically co-oncogenic potential and stimulate tumor progression. Future will tell us how clinical oncologists will succeed to navigate between Scylla and Charybdis and in which particular tumors, in combination with which set of drugs, and in which therapeutic regimes, autophagy inhibitors or inducer will become efficient anticancer agents.
Abbreviations
- GFP:
-
green fluorescent protein
- NF:
-
nutrient-free medium
- PI:
-
propidum iodine
- siRNA:
-
small interfering RNA
- WT:
-
wild type
References
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402: 672–676.
Levine B . Cell biology: autophagy and cancer. Nature 2007; 446: 745–747.
Amaravadi RK, Thompson CB . The roles of therapy-induced autophagy and necrosis in cancer treatment. Clin Cancer Res 2007; 13: 7271–7279.
Mathew R, Karantza-Wadsworth V, White E . Role of autophagy in cancer. Nat Rev Cancer 2007; 7: 961–967.
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Kroemer G, Jaattela M . Lysosomes and autophagy in cell death control. Nat Rev Cancer 2005; 5: 886–897.
Maiuri MC, Zalckvar E, Kimchi A, Kroemer G . Self-eating and self-killing: cross talk between autophagy and apoptosis. Nat Rev Mol Cell Biol 2007; 8: 741–752.
Boya P, Gonzalez-Polo RA, Casares N, Perfettini JL, Dessen P, Larochette N et al. Inhibition of macroautophagy triggers apoptosis. Mol Cell Biol 2005; 25: 1025–1040.
Galluzzi L, Miguel J, Kepp VO, Tasdemir E, Maiuri MC, Kroemer G . To die or not to die: that is the autophagic question. Curr Mol Med 2008; 8: 78–91.
Tsujimoto Y, Shimizu S . Another way to die: autophagic programmed cell death. Cell Death Differ 2005; 2: 1528–1534.
Shaw RJ, Cantley LC . Ras, PI(3)K and mTOR signalling controls tumour cell growth. Nature 2006; 441: 424–430.
Degenhardt K, Mathew R, Beaudoin B, Bray K, Anderson D, Chen G et al. Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 2006; 10: 51–64.
Mathew R, Kongara S, Beaudoin B, Karp CM, Bray K, Degenhardt K et al. Autophagy suppresses tumor progression by limiting chromosomal instability. Genes Dev 2007; 21: 1367–1381.
Karantza-Wadsworth V, Patel S, Kravchuk O, Chen G, Mathew R, Jin S et al. Autophagy mitigates metabolic stress and genome damage in mammary tumorigenesis. Genes Dev 2007; 21: 1621–1635.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
Maiuri MC, Le Toumelin G, Criollo A, Rain JC, Gautier F, Juin P et al. Functional and physical interaction between Bcl-X(L) and a BH3-like domain in Beclin-1. EMBO J 2007; 26: 2527–2539.
Erlich S, Mizrachy L, Segev O, Lindenboim L, Zmira O, Adi-Harel S et al. Differential interactions between Beclin 1 and Bcl-2 family members. Autophagy 2007; 3: 561–568.
Saeki K, Yuo A, Okuma E, Yazaki Y, Susin SA, Kroemer G et al. Bcl-2 down-regulation causes autophagy in a caspase-independent manner in human leukemic HL60 cells. Cell Death Differ 2000; 7: 1263–1269.
Akar U, Chaves-Reyes A, Barria M, Tari A, Saungino A, Kondo Y et al. Silencing of Bcl-2 expression by small interfering RNA induces autophagic cell death in MCF-7 breast cancer cells. Autophagy 2008; 4: 669–679.
Daido S, Kanzawa T, Yamamoto A, Takeuchi H, Kondo Y, Kondo S . Pivotal role of the cell death factor BNIP3 in ceramide-induced autophagic cell death in malignant glioma cells. Cancer Res 2004; 64: 4286–4293.
Hamacher-Brady A, Brady NR, Logue SE, Sayen MR, Jinno M, Kirshenbaum LA et al. Response to myocardial ischemia/reperfusion injury involves Bnip3 and autophagy. Cell Death Differ 2007; 14: 146–157.
Abedin MJ, Wang D, McDonnell MA, Lehmann U, Kelekar A . Autophagy delays apoptotic death in breast cancer cells following DNA damage. Cell Death Differ 2007; 14: 500–510.
Rashmi R, Pillai SG, Vijayalingam S, Ryerse J, Chinnadurai G . BH3-only protein BIK induces caspase-independent cell death with autophagic features in Bcl-2 null cells. Oncogene 2008; 27: 1366–1375.
Liang C, Feng P, Ku B, Dotan I, Canaani D, Oh BH et al. Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat Cell Biol 2006; 8: 688–699.
Fimia GM, Stoykova A, Romagnoli A, Giunta L, Di Bartolomeo S, Nardacci R et al. Ambra1 is a novel regulator of autophagy and controls nervous system development. Nature 2007; 447: 1121–1125.
Rodriguez-Enriquez S, Kim I, Currin RT, Lemasters JJ . Tracker dyes to probe mitochondrial autophagy (mitophagy) in rat hepatocytes. Autophagy 2006; 2: 39–46.
Takahashi Y, Coppola D, Matsushita N, Cualing HD, Sun M, Sato Y et al. Bif-1 interacts with Beclin 1 through UVRAG and regulates autophagy and tumorigenesis. Nat Cell Biol 2007; 9: 1142–1151.
Letai AG . Diagnosing and exploiting cancer’s addiction to blocks in apoptosis. Nat Rev Cancer 2008; 8: 121–132.
Oberstein A, Jeffrey P, Shi Y . Crystal structure of the BCL-XL-beclin 1 peptide complex: Beclin 1 is a novel BH3-only protein. J Biol Chem 2007; 282: 13123–13132.
Feng W, Huang S, Wu H, Zhang M . Molecular basis of Bcl- XL’s target recognition versatility revealed by the structure of Bcl-XL in complex with the BH3 domain of Beclin-1. J Mol Biol 2007; 372: 223–235.
Maiuri MC, Criollo A, Tasdemir E, Vicencio JM, Tajeddine N, Hickman JA et al. BH3-only proteins and BH3 mimetics induce autophagy by competitively disrupting the interaction between beclin 1 and Bcl-2/Bcl-X(L). Autophagy 2007; 3: 374–376.
Oltersdorf T, Elmore SW, Shoemaker AR, Armstrong RC, Augeri DJ, Belli BA et al. An inhibitor of Bcl-2 family proteins induces regression of solid tumours. Nature 2005; 435: 677–681.
Schweers RL, Zhang J, Randall MS, Loyd MR, Li W, Dorsey FC et al. NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc Natl Acad Sci USA 2007; 104: 19500–19505.
Zhang H, Bosch-Marce M, Shimoda LA, Tan YS, Baek JH, Wesley JB et al. Mitochondrial autophagy is an HIF-1-dependent adaptive metabolic response to hypoxia. J Biol Chem 2008; 283: 10892–10903.
Vogelstein B, Lane D, Levine AJ . Surfing the p53 network. Nature 2000; 408: 307–310.
Vousden KH, Lane DP . p53 in health and disease. Nat Rev Mol Cell Biol 2007; 8: 275–283.
Crighton D, Wilkinson S, O’Prey J, Syed N, Smith P, Harrison PR et al. DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell 2006; 126: 121–134.
Feng Z, Zhang H, Levine AJ, Jin S . The coordinate regulation of the p53 and mTOR pathways in cells. Proc Natl Acad Sci USA 2005; 102: 8204–8209.
Abida WM, Gu W . p53-Dependent and p53-independent activation of autophagy by ARF. Cancer Res 2008; 68: 352–357.
Amaravadi RK, Yu D, Lum JJ, Bui T, Christophorou MA, Evan GI et al. Autophagy inhibition enhances therapy-induced apoptosis in a Myc-induced model of lymphoma. J Clin Invest 2007; 117: 326–336.
Tasdemir E, Maiuri MC, Galluzzi L, Vitale I, Djavaheri-Mergny M, D’Amelio M et al. Regulation of autophagy by cytoplasmic p53. Nat Cell Biol 2008; 10: 676–687.
Klionsky DJ, Cuervo AM, Seglen PO . Methods for monitoring autophagy from yeast to human. Autophagy 2007; 3: 181–206.
Gozuacik D, Kimchi A . DAPk protein family and cancer. Autophagy 2006; 2: 74–79.
Harrison B, Kraus M, Burch L, Stevens C, Craig A, Gordon-Weeks P et al. DAPK-1 binding to a linear peptide motif in MAP1B stimulates autophagy and membrane blebbing. J Biol Chem 2008; 283: 9999–10014.
Wang QJ, Ding Y, Kohtz DS, Mizushima N, Cristea IM, Rout MP et al. Induction of autophagy in axonal dystrophy and degeneration. J Neurosci 2006; 26: 8057–8068.
Cully M, You H, Levine AJ, Mak TW . Beyond PTEN mutations: the PI3K pathway as an integrator of multiple inputs during tumorigenesis. Nat Rev Cancer 2006; 6: 184–192.
Schwartz RA, Fernandez G, Kotulska K, Jozwiak S . Tuberous sclerosis complex: advances in diagnosis, genetics, and management. J Am Acad Dermatol 2007; 57: 189–202.
Ji H, Ramsey MR, Hayes DN, Fan C, McNamara K, Kozlowski P et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 2007; 448: 807–810.
Liang J, Shao SH, Xu ZX, Hennessy B, Ding Z, Larrea M et al. The energy sensing LKB1-AMPK pathway regulates p27(kip1) phosphorylation mediating the decision to enter autophagy or apoptosis. Nat Cell Biol 2007; 9: 218–224.
Miracco C, Cosci E, Oliveri G, Luzi P, Pacenti L, Monciatti I et al. Protein and mRNA expression of autophagy gene Beclin 1 in human brain tumors. Int J Oncol 2007; 30: 429–436.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest 2003; 112: 1809–1820.
Yue Z, Jin S, Yang C, Levine AJ, Heintz N . Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci USA 2003; 100: 15077–15082.
Marino G, Salvador-Montoliu N, Fueyo A, Knecht E, Mizushima N, Lopez-Otin C . Tissue-specific autophagy alterations and increased tumorigenesis in mice deficient in Atg4C/Autophagin-3. J Biol Chem 2007; 282: 18573–18583.
Apel A, Herr I, Schwarz H, Rodemann HP, Mayer A . Blocked autophagy sensitizes resistant carcinoma cells to radiation therapy. Cancer Res 2008; 68: 1485–1494.
Tiwari M, Bajpai VK, Sahasrabuddhe AA, Kumar A, Sinha RA, Behari S et al. Inhibition of N-(4-hydroxyphenyl)retinamide-induced autophagy at a lower dose enhances cell death in malignant glioma cells. Carcinogenesis 2008; 29: 600–609.
Qadir MA, Kwok B, Dragowska WH, To KH, Le D, Bally MB et al. Macroautophagy inhibition sensitizes tamoxifen-resistant breast cancer cells and enhances mitochondrial depolarization. Breast Cancer Res Treat 2008; in press.
Li M, Jiang X, Liu D, Na Y, Gao GF, Xi Z . Autophagy protects LNCaP cells under androgen deprivation conditions. Autophagy 2008; 4: 54–60.
Sato K, Tsuchihara K, Fujii S, Sugiyama M, Goya T, Atomi Y et al. Autophagy is activated in colorectal cancer cells and contributes to the tolerance to nutrient deprivation. Cancer Res 2007; 67: 9677–9684.
Han J, Hou W, Goldstein LA, Lu C, Stolz DB, Yin XM et al. Involvement of protective autophagy in TRAIL-resistance of apoptosis defective tumor cells. J Biol Chem 2008; 283: 19665–19677.
Fung C, Lock R, Gao S, Salas E, Debnath J . Induction of autophagy during extracellular matrix detachment promotes cell survival. Mol Biol Cell 2008; 19: 797–806.
Zitvogel L, Apetoh L, Ghiringhelli F, Kroemer G . Immunological aspects of cancer chemotherapy. Nat Rev Immunol 2008; 8: 59–73.
Kroemer G, Pouyssegur J . Altered tumor cell metabolism; cancer’s Achilles heel. Cancer Cell 2008; 13: 472–482.
Toth ML, Sigmond T, Borsos E, Barna J, Erdelyi P, Takacs-Vellai K et al. Longevity pathways converge on autophagy genes to regulate life span in caenorhabditis elegans. Autophagy 2008; 4: 330–338.
Acknowledgements
GK is supported by Ligue contre le Cancer, Agence Nationale pour la Recherche (ANR), Cancéropôle Ile-de-France, European Commission (Active p53, Apo-Sys, Chemores, TransDeath, Right, Death-Train), Fondation pour la Recherche Médicale, and Institut National du Cancer (INCa).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Maiuri, M., Tasdemir, E., Criollo, A. et al. Control of autophagy by oncogenes and tumor suppressor genes. Cell Death Differ 16, 87–93 (2009). https://doi.org/10.1038/cdd.2008.131
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.131
Keywords
This article is cited by
-
A novel tumor suppressor encoded by a 1p36.3 lncRNA functions as a phosphoinositide-binding protein repressing AKT phosphorylation/activation and promoting autophagy
Cell Death & Differentiation (2023)
-
Redox balance and autophagy regulation in cancer progression and their therapeutic perspective
Medical Oncology (2022)
-
Novel lysosome-targeted anticancer fluorescent agents used in zebrafish and nude mouse tumour imaging
Frontiers of Chemical Science and Engineering (2022)
-
CMTM7 as a novel molecule of ATG14L-Beclin1-VPS34 complex enhances autophagy by Rab5 to regulate tumorigenicity
Cell Communication and Signaling (2021)
-
Measuring Cancer Hallmark Mediation of the TET1 Glioma Survival Effect with Linked Neural-Network Based Mediation Experiments
Scientific Reports (2020)