Abstract
Damage to endoplasmic reticulum (ER) homeostasis that cannot be corrected by the unfolded protein response activates cell death. Here, we identified death-associated protein kinase (DAPk) as an important component in the ER stress-induced cell death pathway. DAPk−/− mice are protected from kidney damage caused by injection of the ER stress-inducer tunicamycin. Likewise, the cell death response to ER stress-inducers is reduced in DAPk−/− primary fibroblasts. Both caspase activation and autophagy induction, events that are activated by ER stress and precede cell death, are significantly attenuated in the DAPk null cells. Notably, in this cellular setting, autophagy serves as a second cell killing mechanism that acts in concert with apoptosis, as the depletion of Atg5 or Beclin1 from fibroblasts significantly protected from ER stress-induced death when combined with caspase-3 depletion. We further show that ER stress promotes the catalytic activity of DAPk by causing dephosphorylation of an inhibitory autophosphorylation on Ser308 by a PP2A-like phosphatase. Thus, DAPk constitutes a critical integration point in ER stress signaling, transmitting these signals into two distinct directions, caspase activation and autophagy, leading to cell death.
Similar content being viewed by others
Main
Disturbances of endoplasmic reticulum (ER) homeostasis that lead to extensive and irreparable damage activate ER-specific cell death mechanisms.1 These involve the upregulation of the CHOP transcription regulator by the PERK/eIF2α/ATF-4 pathway, jun N-terminus kinase (JNK) activation by an ER-based IRE1/TRAF2/ASK1 complex, and in mouse cells, direct activation of ER-localized caspase-12. Additionally, cytosolic calcium concentrations are increased during ER stress, contributing to cell death activation.1 ER-stress-induced cell death has been, in the past, described as apoptotic in its nature, involving the activation of caspase-dependent pathways.1 More recently, several reports have been published describing how certain ER stress inducers, including polyglutamine repeats associated with neurodegenerative disorders, DTT and various ER stress-inducing drugs, can also activate autophagy.2, 3, 4 Autophagy is characterized by sequestration of bulk cytoplasm, proteins and organelles in double or multimembrane autophagic vesicles, and their delivery to, and subsequent degradation by, the cell's own lysosomal/vacuolar system.5, 6 During metabolic stress, such as amino-acid starvation, the upregulation of autophagy serves as a means to provide an alternative source of intracellular building blocks and energy, allowing the cell to remain viable under unfavorable circumstances.7 In other cellular settings, however, increased autophagic activity has been associated with cell death.8, 9 Under such circumstances, the RNAi knockdown of key autophagy genes blocked cell death, indicating that autophagy can actually cause cellular demise.10, 11, 12, 13 Notably, this functional duality is also apparent during ER stress, in which autophagy has been shown to protect from further cell death in some scenarios,2, 4, 14, 15 and to promote cell killing in others.4 These studies raise several unresolved issues regarding the cross-talk between apoptosis and autophagy during ER stress-induced cell death.
We have previously shown that overexpression of the calcium-activated calmodulin (CaM)-regulated serine/threonine kinase death-associated protein kinase (DAPk) induces autophagic cell death in cancer cell lines.16 DAPk was originally cloned as a positive mediator of HeLa cell death induced by interferon-γ, and this death was later characterized as caspase-independent autophagy.16, 17 In different experimental settings, DAPk was also shown to be causal to caspase-dependent apoptosis in response to various stimuli.18 This raised the possibility that DAPk may be a positive mediator of both apoptotic and autophagic programmed cell death pathways.
Here, we report that DAPk is a mediator of ER stress-induced cell death, whose catalytic activity is increased by ER stress signals. Knockout of DAPk provides significant protection from ER stress-induced cell death, and both apoptosis and autophagy activation are attenuated. Both processes, shown here to be activated concurrently within the same cells, lead to cellular demise in response to ER stress, as simultaneous disruption of apoptosis and autophagy is required to obtain maximal cell viability. Thus, these studies implicate DAPk as a common upstream integrator of the two types of programmed cell death pathways that are induced by ER stress.
Results
ER stress leads to both apoptosis and autophagy in the same cell
A cell culture model of ER stress was used to thoroughly assess the role of autophagy in this phenomenon. This involved treatment of primary mouse embryonic fibroblasts (MEFs) with drugs that induce ER stress through accumulation of misfolded proteins, including thapsigargin, an ER calcium channel blocker, and tunicamycin, an N-linked glycosylation inhibitor. An assay measuring the autophagy-mediated degradation of radiolabelled long-lived proteins showed a 3.5-fold acceleration in ER-stressed cells, compared to controls at 16–20 h post-treatment (Figure 1A). Furthermore, immunoblots of extracts from tunicamycin treated MEFs for the autophagy marker microtubule-associated protein (MAP) light chain 3 (LC3) indicated a progressive conversion of the free LC3-I form to its lipidated, autophagosome-associated LC3-II form (Figure 1B). This was apparent as early as 8 h after induction of ER stress, when ∼50% of the endogenous LC3 was present in its lipidated form, and continued to increase at 16 and 24 h, with a concomitant induction in total LC3 (Figure 1B, left and middle panels). The accumulation of LC3-II was significantly augmented by addition of the lysosomal inhibitors E64d and pepstatin A, indicating that the shift observed resulted from an increase in autophagic activity and not from a block in lysosomal-mediated degradation of autophagosomes. (Figure 1B, right panel). The presence of autophagosomes was confirmed by transmission electron microscopic (TEM) analysis (Figure 1C). Typical double or multimembrane autophagic vesicles engulfing cytoplasmic components and organelles accumulated in cells treated with ER stress inducers (Figure 1Cd–f, arrowheads).
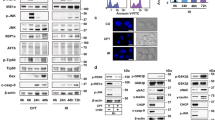
ER stress induces autophagy and caspase activation. (A) Increased rate of autophagic degradation of long-lived proteins after tunicamycin treatment. MEFs were metabolically labeled with radiolabeled valine, treated with DMSO or tunicamycin for 16 h and protein degradation was measured 4 h later (after 20 h of tunicamycin treatment). The results are presented as percent 3-MA inhibited degradation of total-labeled protein. Data are shown as mean±S.D. (n=2). (B) Western blots of total cell lysate from MEFs treated with 1 μg/ml tunicamycin or DMSO and reacted with antibodies to LC3, active caspase-3 and actin, as a loading control. Left panel, kinetics of LC3-II and active caspase-3 accumulation following tunicamycin treatment. Graph in the middle shows quantification of the ratio of LC3-II to LC3-I at each time point, to indicate the extent of conversion to the autophagic LC3-II form. Right panel, the effect of lysosomal protease inhibitors on the tunicamycin-induced LC3-II conversion. Cells were treated for 8 h with DMSO or tunicamycin (TUN), in the absence or presence of the inhibitors (INH): E64d (10 μg/ml) and pepstatin A (10 μg/ml). (C) Ultrastructural analysis of ER stress-induced autophagy by transmission electron microscopy (TEM). TEM images of DMSO (a), thapsigargin (1 μM, 24 h) (THAP, b, d and e) or tunicamycin (1 μg/ml, 24 h) (TUN, c and f) treated MEFs. Scale bars, a–c, 500 nm; d–f, 100 nm; m, mitochondrion; n, nucleus; black arrow, condensed chromatin; white arrowheads, double membrane autophagic vesicles; black arrowheads, multimembrane autophagic vesicles. (D) Simultaneous activation of autophagy and caspase-3 by ER stress. MEFs isolated from GFP-LC3 (green) transgenic mice were treated with DMSO, 1 μM thapsigargin (THAP), 1 μg/ml tunicamycin (TUN) or UV light (UV) (15 kJ/cm2) and viewed 24 h later following immunostaining with a specific antibody to active caspase-3 (red). (E) Quantitation of the percent of cells with punctate GFP-LC3 following treatment of MEFs with tunicamycin or DMSO. A minimum of 200 cells were scored at each time point, and only cells with more than five puncta were considered positive. Values shown are the means±S.D. (n=3)
Endoplasmic reticulum stress in this system also induced markers of the apoptotic pathway. Western blotting indicated proteolytic activation of caspase-3 starting at 16 h after addition of tunicamycin (Figure 1B, left panel). The fact that apoptosis appeared later than the earliest indications of autophagy suggests that the autophagy observed here is not a reaction to, or a compensatory mechanism for, the apoptotic response. To determine whether the two phenomena occurred within the same cell populations, MEFs from a transgenic mouse line expressing the autophagy marker protein GFP-LC319 were used. In non-treated MEFs, GFP-LC3 was diffusively distributed throughout the cell. The treatment of cells with either thapsigargin or tunicamycin caused a punctate staining of GFP-LC3, reflecting the recruitment of LC3 to autophagic vesicles (Figure 1D), similar to the pattern observed in cells starved of amino acids and serum (data not shown). The number of cells with such puncta increased with time, from ∼50% at 8 h, to over 80% at 20 h (Figure 1E). Thus a significant portion of the population exhibited signs of increased autophagic activity following ER stress. Notably, immunostaining of GFP-LC3 fibroblasts with an antibody recognizing only the cleaved, activated form of caspase-3 showed that ER stress caused simultaneous activation of autophagy and caspase-3 cleavage in the same cell (Figure 1D). This contrasted with UV irradiation, which in these cells induced caspase-3 activation only, without the localization shift of GFP-LC3 (Figure 1D). Likewise, close examination of the electron micrographs of cells containing autophagosomes revealed the presence of several hallmarks of apoptosis, such as chromatin condensation (Figure 1Cb, c, black arrows), mitochondrial condensation (Figure 1Cb, c) and membrane blebbing (data not shown). Thus autophagy and apoptosis are not mutually exclusive events, but rather occur within the same cell following induction of ER stress.
Importantly, evidence of autophagy and apoptosis was observed in a mouse model of ER stress-induced death as well. Systemic injection of tunicamycin into mice causes ER stress-mediated death of kidney tubular cells.20 Four days after injection of tunicamycin, mice showed clinical signs of disease, including weight loss, lassitude and lack of grooming. Tdt-mediated dUDP-nick end labeling (TUNEL) staining of kidney sections derived from these mice revealed a high level of apoptosis in the ER-stressed tissue (Figure 5B, DAPk+/+ kidneys). Furthermore, analysis of kidney sections prepared from GFP-LC3 transgenic mice injected with tunicamycin revealed strong punctate and ring-shaped fluorescent staining reflecting the presence of a high level of autophagic activity in the tubular cells (Figure 2A). EM sections of these kidneys clearly exhibited the presence of double or multilayer membrane vesicles engulfing a variety of intracellular components, indicative of the presence of autophagosomes (Figure 2Bc–f). The damaged tubular cells also contained many empty vacuoles (e.g. Figure 2Bc), some of which were engulfed together with the surrounding cytosol by the autophagosomes (Figure 2Be, f). They also contained many condensed mitochondria (Figure 2Bc, d). Notably, advanced chromatin condensation characteristic of apoptosis could be detected in damaged tubular cells, which also display autophagic vesicle accumulation suggesting that autophagy and apoptosis develop in the same cells (Supplementary Figure 1). Taken together these data demonstrate that ER stress activates both apoptosis and autophagy at a single-cell level in cultured primary fibroblasts and in a mouse model system of kidney damage.
Attenuation of tunicamycin-induced kidney tubular cell death in DAPk−/− mice. (A) Hematoxylin and eosin-stained sections of DAPk+/+ (a, b, e, f) or DAPk−/− (c, d, g, h) kidneys from mice injected with vehicle (DMSO, a, e, c, g) or tunicamycin (b, f, d, h). Right panels (scale bar, 50 μ) are higher magnifications of selected areas from the left panels (scale bar, 200 μ). Arrows indicate vacuolated cells (f, h). (B) The TUNEL (upper panels) or DAPI (lower panels) staining of kidneys from DMSO or tunicamycin (TUN)-injected mice. (C) Quantification of TUNEL-positive nuclei per high power field in kidney sections. The results are from five individual DAPk+/+ and DAPk−/− mice. Each point represents the mean of multiple high power fields from an individual mouse
ER stress induces hallmarks of apoptosis and autophagy in vivo in the intact kidney. (A) Intracellular localization of GFP-LC3 in the renal tubule upon injection of tunicamycin to GFP-LC3 transgenic mice. Note the collapse of the tubule structure following tunicamycin treatment, with the disintegration of the tubular cells (d), loss of nuclei (n) and distortion of the lumen (L). Arrows indicate autophagic vesicles within individual tubular cells. Arrowheads indicate brush border of tubular cells. (B) Ultrastructural analysis of ER stress-induced autophagy in the kidney by transmission electron microscopy (TEM). Shown are images of kidneys from DMSO (a, b) or tunicamycin-injected mice (c–f). Scale bar, a, c, 5 μ; b, d, 1 μ; e, f 200 nm. m, mitochondrion, n, nucleus, mv, microvilli. Arrows, double (d, f) and multimembraned (e) autophagic vesicles. Note mitochondrial condensation/darkening (m) in cells with autophagic vesicles
Both caspase activation and autophagy contribute to ER stress-induced cell death
To determine whether ER-induced autophagy positively mediates cell death or alternatively protects from cell death, the effects of 3-MA (3-methyladenine) (a commonly used inhibitor of autophagy) on the cell death response to tunicamycin were evaluated. Notably, administration of 3-MA did not accelerate cell death. Instead, it significantly augmented cell viability, but only when added in combination with zVAD, a pancaspase inhibitor. By itself, zVAD had no discernible protective effect on cell viability, suggesting that the combined action of caspase activation and autophagy induction contribute to cellular killing (Figure 3a). Similarly, as recently reported,21 addition of 3-MA to Bax/Bak double-knockout fibroblasts, which fail to activate the caspase pathway in response to ER stress, further increased cell viability (Supplementary Figure 2). These observations were confirmed by utilizing double genetic disruptions, which target both the autophagic and apoptotic pathways. To this end, siRNA was used to knock down caspase-3 or Beclin1, on the background of Atg5−/− or caspase-3−/− MEFs, respectively. Atg5 and Beclin1 are essential components of the basic autophagy machinery, and Atg5 knockout cells or Beclin1-depleted cells are unable to activate autophagy in response to classical stimuli such as amino acid starvation.22 Atg5 knockout cells are not able to lipidate LC3 in response to tunicamycin, an essential step in the formation of autophagic vesicles (Supplementary Figure 3). Knocking out Atg5 or knocking down Beclin-1 had no significant effect on the death response to ER stress (Figure 3b and c). The fact that inhibition of autophagy by itself did not accelerate cell death, ruled out the possibility that autophagy is death protective in this cellular setting. In contrast, however, in combination with caspase-3 perturbation (by either knockout or knockdown) the perturbation of Atg5 or of Beclin-1 was clearly cell death protective (Figure 3b and c). Notably, the single perturbation of caspase-3 by itself either protected to some extent from cell death or had no effect pending on whether the gene was knocked out or down, respectively (Figure 3b and c). This difference may emerge from residual levels of caspase-3 in the knocked-down cells. Yet, in spite of these differences, in both cases the second perturbation of an autophagic gene further protected from ER-induced cell death. These findings imply that in primary fibroblasts, autophagy is a death-promoting mechanism, the contribution of which becomes apparent when the caspase-dependent pathway is blocked.
Autophagy is necessary for ER stress-induced cell death. (a) MEFs were treated with 1 μg/ml tunicamycin in combination with zVAD (50 μM), 3-MA (5 mM), or both, and cell survival was evaluated after 24 h by the neutral red uptake assay. Data are shown as mean±S.D. (n=3, *P<0.05). (b) ATG5+/+ or ATG5−/− MEFs were transfected with control (HcRed) or caspase-3 siRNAs and cell survival was evaluated after 48 h tunicamycin treatment by propidium iodide exclusion. Data are shown as mean±S.D. (n=3, *P<0.01). Western blot shows successful knockdown of caspase-3 in ATG5+/+ or −/− MEFs. (c) Caspase-3+/+ or caspase-3−/− MEFS were transfected with control (HcRed) or Beclin1 siRNAs and cell survival was evaluated after 48 h tunicamycin treatment. Data are shown as mean±S.D. (n=3, *P<0.02). Western blot shows successful knockdown of Beclin1 in caspase-3 −/−MEFS. In (b and c), nonspecific (NS) band recognized by the antibodies was used as loading controls
DAPk knockout fibroblasts are resistant to ER stress-induced cell death
Several signaling molecules, including DAPk, have been shown to mediate apoptotic or autophagic death pathways, depending on the death stimulus and cell type. Presumably, such dual-nature proteins are likely to have a significant function in ER stress-induced death, which has both apoptotic and autophagic components. To study the function of DAPk, we used a mouse line generated in our laboratory in which the DAPk gene was deleted (DAPk−/− mice) (see Supplementary Figure 4 for generation of knockout mice). Previous studies have indicated that knock-out of DAPk confers protection from several external stresses, including glutamate toxicity to retinal ganglion cells in the intact animal,23 ceramide treatment of cultured hippocampal neurons24 and oncogene activation in isolated MEFs.25 To explore the possible involvement of DAPk in ER stress-induced cell death, the response of DAPk−/− MEFs to various ER stress inducers, including brefeldin A, thapsigargin and tunicamycin, was assessed (Figure 4a and c). Strikingly, in each case, DAPk−/− MEFs displayed attenuated cell death as compared to DAPk+/+ controls. The reduction in cell death in response to tunicamycin was similar to that observed in the double knockout of Bax and Bak (Figure 4b), previously shown to be necessary for ER stress-induced cell death.26 In marked contrast, the death responses to UV light, adriamycin and staurosporine were indistinguishable between the wild-type and DAPk-null fibroblasts (Figure 4a). A colony-formation assay using different doses of tunicamycin confirmed the resistance of DAPk−/− MEFs to ER stress-induced cell death compared to wild-type MEFs (Figure 4d). Altogether, these results indicate that DAPk has a significant function in cell death activated by ER stress.
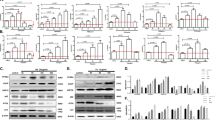
Attenuation of cell death in DAPk−/− MEFs in response to ER stress inducers. (a) Survival of DAPk+/+ versus DAPk−/− MEFs after treatment with ultra violet (UV) light (60 kJ/cm2), adriamycin (ADM, 1 μg/ml), staurosporine (STS, 1 μM) or ER stress inducers brefeldin A (BrefA, 10 μg/ml), thapsigargin (THAP, 1 μM) and tunicamycin (TUN, 1 μg/ml). Data are shown as mean±S.D. (n=3, *P<0.02). (b) Survival of Bax/Bak+/+ versus Bax/Bak −/− MEFs after tunicamycin (1 μg/ml) treatment. Data are shown as mean±S.D. (n=4, *P<0.005). (c) Phase contrast microscopy of DAPk+/+ or DAPk−/− MEFs treated with DMSO or tunicamycin. (d) Colony-formation assay performed after tunicamycin treatment of MEFs at indicated doses. Data are shown as mean±S.D. (n=3, *P<0.05). (e) Immunoblot showing procaspase-12 cleavage following 16 h tunicamycin treatment of DAPk+/+ or DAPk−/− MEFs. Tubulin is used as a loading control. (f) Immunoblot showing CHOP induction following tunicamycin treatment of DAPk+/+ or DAPk−/− MEFs for the indicated times. (g) Kinetics of JNK activation by phosphorylation in tunicamycin (TUN, 1 μg/ml) treated MEFs analyzed by immunoblotting for phospho JNK (P-JNK) or total JNK
Interestingly, we found that DAPk deficiency does not interfere with the previously identified ER stress-induced pathways, namely induction of CHOP, activation of caspase-12 or JNK. As shown in Figure 4e, caspase-12 cleavage occurred to a similar degree regardless of the DAPk status of the cells and induction of CHOP expression was not attenuated in DAPk−/− MEFs (Figure 4f). In both DAPk+/+ and DAPk−/− MEFs, the activated phosphorylated form of JNK protein was observed as early as 15 min after tunicamycin treatment (data not shown), peaking at 30–90 min (Figure 4g). Thapsigargin treatment of DAPk knockout cells also resulted in comparable levels of JNK phosphorylation (data not shown). Thus, DAPk is not necessary for the immediate cellular response to ER stress that involves activation of JNK, CHOP or caspase-12.
DAPk knockout mice are resistant to kidney toxicity caused by systemic tunicamycin injection
As DAPk is highly expressed in the kidney, and especially in renal tubular cells,27 we assessed whether DAPk is necessary for ER stress-induced tubular cell death in vivo upon injection of tunicamycin. Analysis of hematoxylin and eosin-stained sections of the DAPk+/+ kidneys revealed multifocal coalescing degeneration of proximal tubules with the presence of cellular debris in the lumen (Figure 5Ab, f). Many tubular cells were filled with vacuoles occupying almost the entire cytoplasm, pushing the nucleus to the side (Figure 5Af, arrows). In contrast, tubular damage in the DAPk−/− kidneys was more limited. In some areas, DAPk−/− tubular cells showed the presence of small vacuoles in their cytoplasm (Figure 5Ad, h); yet even there, the cellular and tubular architecture was preserved (Figure 5Ah, arrows). No signs of kidney toxicity were observed in DAPk+/+ (Figure 5Aa, e) and DAPk−/− (Figure 5Ac, g) mice injected with the same volume of the vehicle DMSO. TUNEL staining was used to quantify cell death. While minimal TUNEL positivity was observed following DMSO injection, extensive TUNEL-positive staining was detected in the kidney sections of tunicamycin injected DAPk+/+ mice (43.6±10 per high power field) (Figure 5B and C). In contrast, the number of TUNEL-positive nuclei in DAPk−/− kidneys was significantly reduced (17.5±13). Therefore, lack of DAPk conferred resistance to ER stress-induced cell death in vivo.
ER stress activates DAPk by causing Ser308 dephosphorylation
In growing cells, DAPk activity is under strict negative control involving auto-inhibition of the catalytic cleft by interaction with its CaM regulatory-binding segment, and an inhibitory autophosphorylation on Ser308 within this domain.28 DAPk is activated when these controls are relieved, respectively, by binding of CaM to the CaM-regulatory domain and dephosphorylation of the inhibitory Ser.308 To study the possible activation of DAPk by ER stress, we developed monoclonal antibodies specifically recognizing the inactive, phosphorylated Ser308 form of DAPk (Supplementary Figure 5). These anti-phosphoSer308 antibodies were used to immunoprecipitate endogenous DAPk from HEK293T cells treated with tunicamycin to determine the abundance of the phosphorylated inactive form relative to total DAPk. A pronounced, reproducible decrease in the levels of phosphorylated DAPk was observed at 4–8 h (Figure 6a) and was detectable as early as 1 h after tunicamycin treatment (Figure 6b). Thus, DAPk is dephosphorylated before LC3 is lipidated (Figure 1B and Supplementary Figure 6A), and before cell death morphology is first evident in 293T cells, at 18–24 h. Notably, the dephosphorylation persisted for at least 24 h (data not shown). These data suggest that a major portion of the intracellular DAPk pool is activated by ER stress. This was further supported by measuring kinase activity in vitro using myosin II regulatory light chain (MLC) as a substrate. In these assays, endogenous DAPk protein was immunoprecipitated with antibodies to the C-terminus of the protein, which recognize both phosphorylated and non-phosphorylated DAPk. DAPk immunoprecipitated from tunicamycin-treated cells showed significantly higher kinase activity in the presence of 0.1–1 nM calmodulin, compared to DMSO-treated cells (Figure 6c).
DAPk is activated by ER stress through dephosphorylation of Ser.308 (a) Time-course of tunicamycin-dependent dephosphorylation of DAPk. DAPk was immunoprecipitated (IP) from DMSO (0) or from 1 μg/ml tunicamycin (TUN)-treated HEK293T cells at the indicated time points using anti-phosphoSer308 (P-Ser308) antibody and immunoblotted with anti-DAPk antibody. TCL, total cell lysate. Tubulin and a non-relevant protein are used as loading controls. The graph at the right shows quantitation of the extent of dephosphorylation normalized to levels of DAPk in total cell lysate at representative time points (n=3; *P<0.05). (b) DAPk was immunoprecipitated and blotted as in (a) from DMSO- (0) or 1 μg/ml tunicamycin (TUN)-treated cells at early time points. (c) Stimulation of DAPk activity by ER stress. DAPk was immunoprecipitated with anti-DAPk antibodies from DMSO- or tunicamycin-treated cells and an in vitro kinase assay was performed using MLC (2 μg) as a substrate in the presence of EGTA, 0.1 or 1 nM calmodulin (CaM). Kinase activity, calculated by quantifying the relative MLC phosphorylation levels after normalization to DAPk protein levels, is shown as mean±S.D. (n=4, *P<0.05). (d) Abrogation of ionomycin-induced DAPk Ser308 dephosphorylation by PP2A inhibition. HEK293T cells transfected with DAPk were treated for 30 min with DMSO (D), 5 μM ionomycin or 5 μM ionomycin with 1 μM calyculin (Caly), 1 μM cyclosporine A (CycA) or increasing concentrations of okadaic acid (OA). Protein extracts were immunoblotted with anti-phosphoSer,308 anti-DAPk (DAPk) or anti-tubulin antibodies. (e) In vitro dephosphorylation of DAPk by PP2A but not by PP2B or PP1. 100 ng purified recombinant DAPk was incubated in the absence or presence of purified PP2A (0.5 U), PP2B (1 μg) or PP1 (0.5 U) and immunoblotted with anti-phosphoSer308 and anti-DAPk antibodies
To identify the phosphatase responsible for DAPk Ser308 dephosphorylation, various phosphatase inhibitors were screened in cells. As the combination of ER stress-inducers with specific inhibitors of major serine/threonine phosphatases was highly toxic, ionomycin was used as a stimulus. Ionomycin, which causes a fast and robust increase in cytosolic calcium concentrations, has been shown to induce autophagy in MCF7 cells, similar to the calcium mobilizer thapsigargin.29 LC3 mobility shift assays confirmed that it induces autophagy in 293T cells as well (Supplementary Figure 6B). Ionomycin treatment led to a rapid dephosphorylation of DAPk Ser308 after 30 min (Figure 6d, lanes 1, 2). In this setting, the addition of the PP2B inhibitor cyclosporine A had no effect on Ser308 dephosphorylation (Figure 6d, lane 4), whereas protein phosphatase 1 (PP1) and PP2A inhibitors calyculin A and okadaic acid blocked it (Figure 6d, lanes 3, 7). Okadaic acid is a more potent inhibitor of PP2A and its closely related family members than of PP1, and concentrations lower than 10 nM fail to inhibit PP1, while inhibiting the PP2A family. It was found that even low concentrations of okadaic acid in the nanomolar range were able to potently inhibit DAPk Ser308 dephosphorylation, suggesting that a PP2A-like phosphatase is responsible for DAPk activation (Figure 6d, lanes 5–7). It should be noted that siRNA-mediated knockdown experiments of PP2A were inconclusive as the depletion of PP2A had additional indirect effects on cells. It strongly reduced the steady state levels of DAPk protein in cells, irrespective of whether they were treated or not by tunicamycin, thus preventing an accurate assessment of the dephosphorylation event (Supplementary Figure 7). Finally, to confirm that PP2A directly dephosphorylates DAPk, an in vitro dephosphorylation assay using purified proteins was performed. As shown in Figure 6e, incubation of recombinant DAPk with purified PP2A enzyme caused dephosphorylation of Ser.308 In contrast, neither PP2B nor PP1 was effective in these assays. Thus, the cell-based okadaic acid effects and the in vitro dephosphorylation assays are consistent with the possibility that PP2A or a closely related phosphatase may be the serine/threonine phosphatase activating DAPk during ER stress.
DAPk promotes ER stress-induced caspase-activation and autophagy
A quantitative analysis of the effects of DAPk deletion on ER stress-induced autophagy and caspase activation was then undertaken. A DEVDase assay showed that caspase-3-like activity in DAPk−/− MEFs was attenuated when compared to WT controls (Figure 7a). In line with this data, accumulation of the active form of caspase-3 was attenuated in DAPk−/− MEFs (Supplementary Figure 8A). In contrast, the deletion of DAPk did not block caspase activation upon treatment of MEFs with the classical apoptosis inducer staurosporine (Supplementary Figure 8A) consistent with the lack of protection from staurosporine-induced cell death observed in Figure 4a. To assess the effect of DAPk deletion on the autophagic process, DAPk−/− mice were crossed to GFP-LC3 transgenic mice. GFP-LC3 DAPk +/+ and −/− MEFs derived from these mice were used to measure the number of LC3 dots/mm2 of cellular area after 8, 16 or 24 h exposure to tunicamycin. GFP-LC3 dots accumulated in wild-type MEFs in a time-dependent manner, whereas in the DAPk−/− cells, this process was significantly inhibited (Figure 7b). The number of GFP-LC3 dots/mm2 was also significantly reduced in DAPk−/− MEFs treated with thapsigargin for 24 h, compared to DAPk+/+ counterparts (Figure 7c). Two different knockout fibroblast preparations gave similar results (data not shown). Again, these protective effects of DAPk deletion were unique to ER stress. The lack of DAPk in MEFs reduced the accumulation of the LC3-II form in response to tunicamycin and not in response to amino acid starvation (Supplementary Figure 8B). These data demonstrate that activation of DAPk by ER stress signals mediates caspase activation and autophagic induction in primary fibroblasts.
Attenuation of both caspase activity and autophagy in cells lacking DAPk. (a) Decreased DEVDase activity in 20 h tunicamycin-treated DAPk−/− MEF extracts. Data are shown as mean±S.D. (n=4, *P<0.0001). (b) GFP-LC3 dots/mm2 in MEFs treated 8, 16 or 24 h with 1 μg/ml tunicamycin. Data are shown as mean±S.D. (n=3, *P<0.005, **P<0.05). (c) GFP-LC3 dots/mm2 in MEFs treated 24 h with 1 μM thapsigargin. Data are shown as mean±S.D. (n=4, *P<0.0005)
Discussion
In this study, we showed that ER stress induces a mixed type of cell death with characteristics of both apoptosis (i.e., caspase activation, chromatin and mitochondrial condensation) and autophagic cell death (i.e., accumulation of autophagic vesicles). To gain insight into the molecular mechanism, which integrates these two pathways, we used cells and mice that are deficient of DAPk. DAPk was previously reported to induce cell death with apoptotic or autophagic characteristics in independent cellular and experimental contexts.18 Indeed, we found here that DAPk is activated during ER stress and is positively connected to these two processes, which develop within the same cells. We proved that ER stress led to the activation of DAPk catalytic activity by dephosphorylation of a key residue in its calmodulin-regulatory domain. Quantitative measurements of caspase activity and accumulation of autophagic vesicles demonstrated that the hallmarks of both apoptotic and autophagic pathways activated during ER stress were attenuated in DAPk-deficient cells. The requirement for DAPk during ER stress-induced cell death was further confirmed by the resistance of DAPk-null cells to cell death caused by agents leading to prolonged ER stress. This was shown in vitro using isolated fibroblasts in culture and in vivo in a kidney toxicity model. The major death-related pathways activated by ER stress, involving caspase-12, transcriptional upregulation of CHOP and early JNK activation, however, were not altered in DAPk-null cells. Thus, DAPk may act downstream to these molecules in the known ER stress-induced signaling pathways leading to cell death, or alternatively, may define a novel signaling pathway, which gauges the stress response and activates cell death. Hence, DAPk is one of the key molecules activated by ER stress and is necessary for cellular demise by a mixed type of cell death under these conditions.
The capacity of ER stress to induce autophagy in cultured cells has been recently addressed by several groups. In yeast, where autophagy mainly acts as a survival-promoting response, ER stress induction led to autophagy activation.3, 15 In studies using mammalian cells in culture which were exposed to various ER stress-inducing drugs or polyglutamine aggregates, autophagy was alternatively described as either death protective or death promoting.4, 14, 30, 31 It has been proposed that autophagy contributes to cell death in non-transformed cells, while it serves to attenuate apoptotic cell death in transformed cancer cell lines.4 Here, we found that the ER-induced autophagy in primary fibroblasts had prodeath characteristics. Notably, increased cell viability could be detected only if both cell death modalities were attenuated simultaneously by knocking out/down critical genes along these pathways (e.g., caspase3, Atg5). A recent publication also provided evidence for enhanced cell survival upon Atg5 knockdown or 3-MA treatment in cells in which apoptotic cell death was blocked by overexpression of Bcl-XL or knockout of Bax/Bak.21 Similar to our study, the protection from ER stress-induced cell death by the combined perturbation of apoptotic and autophagic genes was always partial. This further suggests the existence of additional mechanisms of cell death such as necrosis, which may be activated in parallel by the ER stress or may function as a backup mechanism called into action when the other two pathways are attenuated.
DAPk has been linked to apoptosis through the phosphorylation of MLC,32, 33 which promotes membrane blebbing, through the inhibition of integrin function and induction of anoikis34 and through the activation of p53.25 In this system, however, we can exclude the involvement of p53 in the DAPk/ER stress pathway, as neither p53 nor its target genes, including p21, Mdm2 and PUMA, were upregulated by ER stress in either WT or DAPk −/− cells, and p53 −/− MEFs retained their sensitivity to ER stress-induced death (Supplementary Figure 9). The molecular mechanisms linking DAPk to autophagy have not yet been characterized. It is possible that the previously identified DAPk substrate syntaxin-1A, a component of the v-SNARE complex, may be involved.35 Notably, the CaM-dependent kinase kinase β (CaMKKβ), another substrate of DAPk,36 has been implicated in mediating autophagy induced by ionomycin and inducers of ER stress.29 However, phosphorylation of CaMKKβ by DAPk on Ser511 attenuates the former's CaM-dependent autophosphorylation, a step necessary for its activation, suggesting that DAPk actually inhibits CaMKK function. Thus, it is unlikely to be a downstream mediator of DAPk in the autophagy-signaling pathway. Future characterization of the direct targets of DAPk will clarify how the kinase is linked to both apoptotic and autophagic death-related pathways.
To our knowledge, the DAPk knockout mouse is the first model where two death-related processes, caspase activation and autophagy, are attenuated by the knockout of a single gene. This indicates that during ER stress-induced cell death, DAPk functions as node of integration between apoptosis and autophagy, which may mediate coordinated activation and cooperativity between the pathways. Yet, deletion of DAPk did not block either pathway entirely, nor was complete viability restored in DAPk−/− cells. Thus, just as the presence of multiple executioners, such as the caspases, and regulators, such as the Bcl-2 family, provide compensatory redundancy to ensure a robust and fine-tuned response to an apoptotic signal, so too, several layers of signal sensors, DAPk comprising one, would ensure activation of the proper mixed apoptotic and autophagic responses. The data presented here underline the importance of combinatorial drug treatment approaches for human diseases manifesting hallmarks of ER stress, apoptosis and autophagy, including neurodegenerative diseases and stroke. These approaches should target not only apoptotic pathways, but also pathways involving alternative types of cell death, and molecules like DAPk, which regulate multiple death pathways, are therefore the most promising targets for such treatment approaches.
Materials and Methods
Generation of DAPk-deficient mice
The DAPk gene was disrupted with a targeting vector derived from a 20 kb genomic fragment encompassing the first exon, which was isolated by screening a 129-SVJ genomic phage library (Stratagene, La Jolla, CA, USA) with a 460 bp 5′ fragment of mouse DAPk cDNA. The positive selection marker, neomycin, was inserted into the first exon, flanked by homologous arms of 3.5 kb and a 1.8 kb, respectively, and the diphtheria toxin gene was added for negative selection. The targeting construct was electroporated into ES cells of 129-SVJ background, which were microinjected into wild-type C57Bl/6J blastocysts to generate chimeric mice. Gene deletion was confirmed by PCR and by northern blot analysis using 2.5 μg polyA+ RNA, and probes generated from radiolabeled PCR fragments corresponding to nt 3187–4293 of mouse DAPk sequence or to full-length GAPDH. Germ line transmission was confirmed by mating chimeras with wild-type C57Bl/6J mice, and 129/C57Bl/6J mixed background siblings were obtained. C57Bl/6J background mice were obtained after 10 generations of consecutive backcrossing. GFP-LC3 transgenic DAPk−/− or +/+ mice were obtained by crossing C57Bl/6J DAPk−/− mice to GFP-LC3 mice19 of the same background.
Cell culture
HEK293T cells and MEFs were cultured as described.16, 25 MEFs were isolated from the embryos of identical genotype (pools of 6–9 embryos per genotype) as described25 and assayed at passage 3–6. Immortalized polyclonal ATG5−/− and ATG5+/+ fibroblasts were obtained by transfection of primary MEFs22 with the whole SV40 genome. Bax/Bak−/− and +/+ fibroblasts were obtained by SV40 immortalization of primary MEFs as described.26 Caspase-3−/− fibroblasts were previously described.37 For gene-depletion experiments, control Hc-Red siRNA (5′-GUAUGCGCAUCAAGAUGUA-3′), siGENOME commercial 19mer siRNAs targeting two different regions of the mouse caspase-3 sequence (Cat no. D-043042-03 and D043042-04) and siRNA targeting Beclin1 (Cat no. D-55895-01) were purchased from Dharmacon (Lafayette, CO, USA). MEFs (3 × 106) were transfected with 5 μg siRNA by electroporation using the Amaxa (Gaithersburg, MD, USA) MEF-II kit and Amaxa Nucleofector program A023.
Cell death experiments
Cell death inducers (Sigma-Aldrich, Rehovot, Israel) were administered to MEFs as indicated in the text, and cell viability was assessed 24 h later by measuring the uptake of neutral red dye (Sigma-Aldrich). Propidium iodide uptake (Sigma-Aldrich, 4 μl/ml) was measured by counting the number of positive cells in the whole cell population. For starvation experiments, MEFs were incubated in Earl's balanced salt solution without serum for 5 h. For colony-formation assay, 4 × 105 cells were plated on 100 mm dishes and 24 h later, equal plating was confirmed by the neutral red assay. Triplicate dishes were treated for 24 h with the indicated concentrations of tunicamycin, and were then washed. Twelve days later, plates were fixed and stained with crystal violet and the number of colonies containing >50 cells were counted. The DEVDase assay was performed as described using 40 μg MEF extract per reaction.38 DEVDase activity was expressed as the difference between the crude DEVDase activity and the activity in the presence of 50 μM ZVAD (Enzyme Systems Products, MP Biomedicals, Solon, OH, USA).
Autophagic activity was quantified by counting GFP-LC3 dots by fluorescent microscopy (see below). GFP-LC3 transgenic fibroblasts were treated with 1 μg/ml tunicamycin or 1 μM thapsigargin. Forty cells per genotype were analyzed from different fields and a micrograph of each counted cell was recorded. After determining the surface area of each cell digitally using Photoshop (Adobe Systems, San Jose, CA, USA) software, the total number of dots per total mm2 cell area was calculated. (A more detailed explanation of the method is presented as Supplementary Methods.) Results shown represent 3–4 independent experiments (120–160 cells per genotype). The number of dots/mm2 in DMSO-treated cells was negligible.
For long-lived protein degradation, MEFs were grown in valine-free α-MEM medium supplemented with 10% FBS and [14C]valine (1.5 μCi/ml) for 24 h and incubated with tunicamycin (1 μg/ml) or DMSO for an additional 13 h. Cells were washed with PBS and incubated 3 h in chase medium (α-MEM supplemented with 10% FBS, 15 mM cold valine and tunicamycin (1 μg/ml) or DMSO, with or without 3-MA (5 mM)) to allow degradation of short-lived proteins. After PBS wash, cells were incubated 4 h in the corresponding fresh chase medium. Long-lived protein degradation was calculated as the percentage of the ratio of total acid-soluble radioactivity in the media and cells to the total TCA-precipitable radioactivity. 3-MA-inhibitable degradation was considered as autophagic degradation.
Immunoblotting
Immunoblotting was performed using monoclonal antibodies to DAPk, β-tubulin and actin (Sigma-Aldrich), CHOP/GADD153 (Santa Cruz Biotechnology, Santa Cruz, CA, USA), PP2Ac (Millipore), phospho-JNK and JNK (gift of R Seger, Weizmann Institute of Science) and rabbit polyclonal antibodies to caspase-12 (Cell Signaling Technology, Beverly, MA, USA), active caspase-3 (Promega, Madison WI, USA), procaspase-3 (Santa Cruz) and anti-LC3 polyclonal antibodies (either contributed by N Mizushima or Sigma-Aldrich).
Histology, immunostaining and microscopy
Two–month-old C57Bl/6 mice were given a single 1 μg/g body weight intraperitoneal injection of tunicamycin. Four days after injection, mice were perfused with 3.7% paraformaldehyde (PFA) in PBS. Hematoxylin and eosin-stained paraffin sections were used for histopathological analysis. Frozen tissue sections were analyzed by TUNEL staining using the ApopTag Red in situ apoptosis detection kit, according to the manufacturer's instructions (Millipore, Billerica, MA, USA). The TUNEL-positive nuclei were counted in 5–10 random fields at the subcortical/proximal tubular area of kidney sections by fluorescent microscopy (BX41, Olympus, Center Valley, PA) at × 400 magnification ( × 40 UPlanFL objective, N.A.=0.75) at room temperature, and the correspondence of each dot to a nucleus was confirmed by comparing the DAPI nuclear staining of the same field. GFP-LC3 MEFs were fixed with 3.7% PFA in PBS, permeabilized with 0.05% Triton X-100/PBS and stained with anti-active caspase-3 antibody (Cell Signaling Technology) followed by rhodamine-conjugated anti-rabbit antibody (Jackson ImmunoResearch, West Grove, PA, USA). Slides were examined at room temperature with the same microscope, using × 60 (N.A.=1.25) and × 100 (N.A.=1.3) UPlan-Fl oil immersion objectives. Digital images were generated with an Olympus DP50 CCD camera using Viewfinder Lite and Studio Lite software. Final composites were prepared using Adobe Photoshop (Adobe Systems). TEM of MEFs and of mouse kidneys were performed as previously described.16
Analysis of Ser308 phosphorylation
HEK293T cells were treated with 1 μg/ml tunicamycin or equal volume of the vehicle DMSO for the indicated times. Cells were lysed in PLB (10 mM NaPO4 pH 7.5, 5 mM EDTA, 100 mM NaCl, 1% Triton, 0.5% Na deoxycholate, 0.1% SDS) containing 100 nM okadaic acid, 1 μM cyclosporine A, 1 mM NaF, 50 mM β-glycerophosphate and protease inhibitors. After preclearing for 30 min on empty protein G-agarose beads (Santa Cruz), cell lysate was incubated at 4°C for 30 min with 1 μl anti-phosphoSer308 antibody prebound to protein-G beads. After three washes in the same buffer, samples were immunoblotted using anti-DAPk antibody. The anti-phosphoSer308 mouse monoclonal antibodies were produced in collaboration with Sigma-Aldrich using peptide CKKWKQS-PO3VRLI as the immunogen (Supplementary Figure 5). Alternatively, for phosphoSer308 analysis of exogenous DAPk, 293T cells transfected with sublethal concentrations of Flag-DAPk were treated for 30 min with 5 μM ionomycin (Sigma-Aldrich) in the presence of DMSO or various phosphatase inhibitors as indicated, lysed in PLB as described above and subjected to western blotting with anti-phosphoSer308 antibody.
In vitro kinase and phosphatase assays
Endogenous DAPk was immunoprecipitated for 2 h as described above using protein A-agarose beads (Santa Cruz) bound to a rabbit polyclonal antibody raised against the DAPk C-terminal tail region.39 Beads were incubated for 20 min at 30°C in 30 μl reaction buffer (50 mM HEPES pH 7.5, 8 mM MgCl2, 2 mM MnCl2 and 0.1 mg/ml BSA, 0.5 mM CaCl2, 50 μM ATP, 2 μg MLC (Sigma-Aldrich), 100 nM okadaic acid, 1 μM cyclosporine A, 1 mM NaF, 50 mM β-glycerophosphate and protease inhibitors) containing 15 μCi [γ-32P]ATP, and the indicated amounts of bovine calmodulin (Sigma-Aldrich) or 5 mM EGTA. Samples were analyzed by immunoblotting and autoradiography. Densitometric values of MLC phosphorylation and DAPk protein levels in each sample were quantified by NIH software. DAPk activity was expressed as the ratio of kinase activity per quantity of DAPk protein in the immunoprecipitate.
Alternatively, Flag-tagged DAPk was purified from transiently transfected HEK293T cells as described.39 100 ng DAPk was incubated for 30 min at 30°C in PP2A reaction buffer (20 mM MOPS, pH 7.5, 150 mM NaCl, 60 mM 2-mercaptoethanol, 1 mM MgCl2, 1 mM EGTA, 0.1 mM MnCl2, 1 mM DTT, 10% glycerol and 0.1 mg/ml serum albumin) or in PP2B reaction buffer (50 mM Tris-HCl, pH 7.5, 2.5 mM DTT, 1 mM CaCl2, 1 mM MnCl2 and 2 μM bovine calmodulin (Sigma-Aldrich)) or in PP1 reaction buffer (50 mM Tris-HCl, pH 7.5, 0.1 mM EDTA, 5 mM DTT, 1 mM MnCl2) in the presence or absence of purified PP2A (0.5 U), PP2B (1 μg) or PP1 (0.5 U) (Upstate, Billerica, MA, USA). Activity of the recombinant enzymes was confirmed by pNPP test before experimentation. Samples were analyzed by immunoblotting as described above.
Statistical analysis
The statistical significance of differences between groups was assessed by two-tailed Student's t-test. Values of P<0.05 were considered significant.
Abbreviations
- 3-MA:
-
3-methyladenine
- CaM:
-
calcium-activated calmodulin
- CaMKKβ:
-
CaM-dependent protein kinase kinase β
- DAPk or DAP-kinase:
-
death-associated protein kinase
- ER:
-
endoplasmic reticulum
- JNK:
-
jun N-terminus kinase
- LC3:
-
microtubule-associated protein (MAP) light chain 3
- MEF:
-
mouse embryonic fibroblast
- MLC:
-
myosin II regulatory light chain
- PP1:
-
protein phosphatase 1
- PP2A:
-
protein phosphatase 2A
- PP2B:
-
protein phosphatase 2B
- PP4:
-
protein phosphatase 4
- TEM:
-
transmission electron microscopy
- TUNEL:
-
Tdt-mediated dUDP-nick end labeling
References
Wu J, Kaufman R . From acute ER stress to physiological roles of the unfolded protein response. Cell Death Differ 2006; 13: 374–384.
Kouroku Y, Fujita E, Tanida I, Ueno T, Isoai A, Kumagai H et al. ER stress (PERK/eIF2alpha phosphorylation) mediates the polyglutamine-induced LC3 conversion, an essential step for autophagy formation. Cell Death Differ 2007; 14: 230–239.
Yorimitsu T, Nair U, Yang Z, Klionsky DJ . Endoplasmic reticulum stress triggers autophagy. J Biol Chem 2006; 281: 30299–30304.
Ding WX, Ni HM, Gao W, Hou YF, Melan MA, Chen X et al. Differential effects of endoplasmic reticulum stress-induced autophagy on cell survival. J Biol Chem 2007; 282: 4702–4710.
Ohsumi Y . Molecular dissection of autophagy: two ubiquitin-like systems. Nat Rev Mol Cell Biol 2001; 2: 211–216.
Shintani T, Klionsky DJ . Autophagy in health and disease: a double-edged sword. Science 2004; 306: 990–995.
Levine B, Yuan J . Autophagy in cell death: an innocent convict? J Clin Invest 2005; 115: 2679–2688.
Baehrecke EH . Autophagic programmed cell death in Drosophila. Cell Death Differ 2003; 10: 940–945.
Gozuacik D, Kimchi A . Autophagy as a cell death and tumor suppressor mechanism. Oncogene 2004; 23: 2891–2906.
Yu L, Alva A, Su H, Dutt P, Freundt E, Welsh S et al. Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 2004; 304: 1500–1502.
Shimizu S, Kanaseki T, Mizushima N, Mizuta T, Arakawa-Kobayashi S, Thompson CB et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat Cell Biol 2004; 6: 1221–1228.
Xu Y, Kim SO, Li Y, Han J . Autophagy contributes to caspase-independent macrophage cell death. J Biol Chem 2006; 281: 19179–19187.
Reef S, Zalckvar E, Shifman O, Bialik S, Sabanay H, Oren M et al. A short mitochondrial form of p19ARF induces autophagy and caspase-independent cell death. Mol Cell 2006; 22: 463–475.
Ogata M, Hino S, Saito A, Morikawa K, Kondo S, Kanemoto S et al. Autophagy is activated for cell survival after endoplasmic reticulum stress. Mol Cell Biol 2006; 26: 9220–9231.
Bernales S, McDonald K, Walter P . Autophagy counterbalances endoplasmic reticulum expansion during the unfolded protein response. PLoS Biol 2006; 4: e423.
Inbal B, Bialik S, Sabanay I, Shani G, Kimchi A . DAP kinase and DRP-1 mediate membrane blebbing and the formation of autophagic vesicles during programmed cell death. J Cell Biol 2002; 157: 455–468.
Deiss LP, Feinstein E, Berissi H, Cohen O, Kimchi A . Identification of a novel serine/threonine kinase and a novel 15-kD protein as potential mediators of the gamma interferon-induced cell death. Genes Dev 1995; 9: 15–30.
Bialik S, Kimchi A . The death-assoicated protein kinases: structure, function and beyond. Annu Rev Biochem 2006; 75: 189–210.
Mizushima N, Yamamoto A, Matsui M, Yoshimori T, Ohsumi Y . In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell 2004; 15: 1101–1111.
Zinszner H, Kuroda M, Wang X, Batchvarova N, Lightfoot RT, Remotti H et al. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev 1998; 12: 982–995.
Ullman E, Fan Y, Stawowczyk M, Chen H, Yue Z, Zong W . Autophagy promotes necrosis in apoptosis-deficient cells in response to ER stress. Cell Death Differ 2007; 15: 422–425.
Kuma A, Hatano M, Matsui M, Yamamoto A, Nakaya H, Yoshimori T et al. The role of autophagy during the early neonatal starvation period. Nature 2004; 432: 1032–1036.
Schori H, Yoles E, Wheeler LA, Raveh T, Kimchi A, Schwartz M . Immune-related mechanisms participating in resistance and susceptibility to glutamate toxicity. Eur J Neurosci 2002; 16: 557–564.
Pelled D, Raveh T, Riebeling C, Fridkin M, Berissi H, Futerman AH et al. Death-associated protein (DAP) kinase plays a central role in ceramide-induced apoptosis in cultured hippocampal neurons. J Biol Chem 2002; 277: 1957–1961.
Raveh T, Droguett G, Horwitz MS, DePinho RA, Kimchi A . DAP kinase activates a p19ARF/p53-mediated apoptotic checkpoint to suppress oncogenic transformation. Nat Cell Biol 2001; 3: 1–7.
Wei MC, Zong WX, Cheng EH, Lindsten T, Panoutsakopoulou V, Ross AJ et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 2001; 292: 727–730.
Yukawa K, Shirasawa N, Ohshima A, Mune M, Kimura A, Bai T et al. Death-associated protein kinase localization to human renal tubule cells, and increased expression of chronic obstructive uropathy in rats. J Nephrol 2004; 17: 26–33.
Shohat G, Spivak-Kroizman T, Cohen O, Bialik S, Shani G, Berrisi H et al. The pro-apoptotic function of death-associated protein kinase is controlled by a unique inhibitory autophosphorylation-based mechanism. J Biol Chem 2001; 276: 47460–47467.
Hoyer-Hansen M, Bastholm L, Szyniarowski P, Campanella M, Szabadkai G, Farkas T et al. Control of macroautophagy by calcium, calmodulin-dependent kinase kinase-beta, and Bcl-2. Mol Cell 2007; 25: 193–205.
Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol 2006; 8: 1124–1132.
Wang Y, Singh R, Massey A, Kane S, Kaushik S, Grant T et al. Loss of macroautophagy promotes or prevents fibroblast apoptosis depending on the death stimulus. J Biol Chem 2008; 238: 4766–4777.
Bialik S, Bresnick A, Kimchi A . DAP-kinase-mediated morphological changes are localization dependent and involve myosin-II phosphorylation. Cell Death Differ 2004; 11: 631–644.
Kuo JC, Lin JR, Staddon JM, Hosoya H, Chen RH . Uncoordinated regulation of stress fibers and focal adhesions by DAP kinase. J Cell Sci 2003; 116: 4777–4790.
Wang WJ, Kuo JC, Yao CC, Chen RH . DAP-kinase induces apoptosis by suppressing integrin activity and disrupting matrix survival signals. J Cell Biol 2002; 159: 169–179.
Tian JH, Das S, Sheng ZH . Ca2+-dependent phosphorylation of syntaxin-1A by the death-associated protein (DAP) kinase regulates its interaction with Munc18. J Biol Chem 2003; 278: 26265–26274.
Schumacher AM, Schavocky JP, Velentza AV, Mirzoeva S, Watterson DM . A calmodulin-regulated protein kinase linked to neuron survival is a substrate for the calmodulin-regulated death-associated protein kinase. Biochem 2004; 43: 8116–8124.
Kuida K, Zheng TS, Na S, Kuan C, Yang D, Karasuyama H et al. Decreased apoptosis in the brain and premature lethality in CPP32-deficient mice. Nature 1996; 384: 368–372.
Haviv R, Lindenboim L, Li H, Yuan J, Stein R . Need for caspases in apoptosis of trophic factor-deprived PC12 cells. J Neurosci Res 1997; 50: 69–80.
Shani G, Marash L, Gozuacik D, Bialik S, Teitelbaum L, Shohat G et al. Death-associated protein kinase phosphorylates ZIP kinase, forming a unique kinase hierarchy to activate its cell death functions. Mol Cell Biol 2004; 24: 8611–8626.
Acknowledgements
We thank Mitsubishi Chemical Corporation (Japan) for critical technical help in the generation of the DAPk−/− mice, Hanna Berissi for technical assistance during the production and testing of anti-phosphoserine308 antibodies, and Sharon Reef for reading the manuscript and fruitful discussions. We also thank Rony Seger (Weizmann Institute of Science) for generously providing anti-JNK and anti-phospho-JNK antibodies, and Atan Gross (Weizmann Institute of Science) for generously providing Bax/Bak−/−, caspase-3−/− and +/+ fibroblasts. The electron microscopy studies were conducted at the Irving and Cherna Moskowitz Center for Nano and Bio-Nano Imaging at the Weizmann Institute of Science. This study was supported by the Israel Science Foundation administered by the Israel Academy of Sciences and Humanities and by the Kahn Fund for System Biology at the Weizmann Institute of Science. AK is the incumbent of Helena Rubinstein Chair of Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by DC Rubinsztein
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Gozuacik, D., Bialik, S., Raveh, T. et al. DAP-kinase is a mediator of endoplasmic reticulum stress-induced caspase activation and autophagic cell death. Cell Death Differ 15, 1875–1886 (2008). https://doi.org/10.1038/cdd.2008.121
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.121
Keywords
This article is cited by
-
Impact of context-dependent autophagy states on tumor progression
Nature Cancer (2023)
-
Structure-Based Virtual Screening and Discovery of New Bi-functional DAPK1 Inhibitors
Molecular Biotechnology (2023)
-
Quantitative Proteomic and Phosphoproteomic Analyses Reveal a Role of Death-Associated Protein Kinase 1 in Regulating Hippocampal Synapse
Molecular Neurobiology (2023)
-
The role of autophagy and apoptosis in early brain injury after subarachnoid hemorrhage: an updated review
Molecular Biology Reports (2022)
-
Endoplasmic reticulum stress and unfolded protein response in cardiovascular diseases
Nature Reviews Cardiology (2021)