Abstract
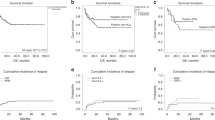
We determined whether assessment of the immunogenicity of individual donor–recipient HLA mismatches based on differences in their amino-acid sequence and physiochemical properties predicts clinical outcome following haematopoietic SCT (HSCT). We examined patients transplanted with 9/10 single HLA class I-mismatched grafts (n=171) and 10/10 HLA-A-, -B-, -C-, -DRB1- and -DQB1-matched grafts (n=168). A computer algorithm was used to determine the physiochemical disparity (electrostatic mismatch score (EMS) and hydrophobic mismatch score (HMS)) of mismatched HLA class I specificities in the graft-versus-host direction. Patients transplanted with HLA-mismatched grafts with high EMS/HMS had increased incidence of ⩾grade II acute GVHD (aGVHD) compared with patients transplanted with low EMS/HMS grafts; patients transplanted with low and medium EMS/HMS grafts had similar incidence of aGVHD to patients transplanted with 10/10 HLA-matched grafts. Mortality was higher following single HLA-mismatched HSCT but was not correlated with HLA physiochemical disparity. Assessment of donor–recipient HLA incompatibility based on physiochemical HLA disparity may enable better selection of HLA-mismatched donors in HSCT.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Furst D, Muller C, Vucinic V, Bunjes D, Herr W, Gramatzki M et al. High resolution HLA-matching in hematopoietic stem cell transplantation: a retrospective collaborative analysis. Blood 2013; 122: 3220–3229.
Claas FH, Roelen DL, Mulder A, Doxiadis II, Oudshoorn M, Heemskerk M . Differential immunogenicity of HLA class I alloantigens for the humoral versus the cellular immune response: "towards tailor-made HLA mismatching". Hum Immunol 2006; 67: 424–429.
Dankers MK, Heemskerk MB, Duquesnoy RJ, Doxiadis II, Oudshoorn M, Roelen DL et al. HLAMatchmaker algorithm is not a suitable tool to predict the alloreactive cytotoxic T-lymphocyte response in vitro. Transplantation 2004; 78: 165–167.
Heemskerk MB, Doxiadis II, Roelen DL, Claas FH, Oudshoorn M . The HistoCheck algorithm does not predict T-cell alloreactivity in vitro. Bone Marrow Transplant 2005; 36: 927–928.
Joris MM, van Rood JJ, Roelen DL, Oudshoorn M, Claas FH . A proposed algorithm predictive for cytotoxic T cell alloreactivity. J Immunol 2012; 188: 1868–1873.
Joris MM, Lankester AC, von dem Borne PA, Kuball J, Bierings M, Cornelissen JJ et al. Translating in vitro prediction of cytotoxic T cell alloreactivity to hematopoietic stem cell transplantation outcome. Transpl Immunol 2014; 30: 59–64.
Kosmoliaptsis V, Chaudhry AN, Sharples LD, Halsall DJ, Dafforn TR, Bradley JA et al. Predicting HLA class I alloantigen immunogenicity from the number and physiochemical properties of amino acid polymorphisms. Transplantation 2009; 88: 791–798.
Kosmoliaptsis V, Sharples LD, Chaudhry AN, Halsall DJ, Bradley JA, Taylor CJ . Predicting HLA class II alloantigen immunogenicity from the number and physiochemical properties of amino acid polymorphisms. Transplantation 2011; 91: 183–190.
Oudshoorn M, Doxiadis II, van den Berg-Loonen PM, Voorter CE, Verduyn W, Claas FH . Functional versus structural matching: can the CTLp test be replaced by HLA allele typing? Hum Immunol 2002; 63: 176–184.
Heemskerk M, Roelen D, Dankers M, van Rood J, Claas F, Doxiadis I et al. Allogeneic MHC class I molecules with numerous sequence differences do not elicit a CTL response. Hum Immunol 2005; 66: 969–976.
Bjorkman PJ, Saper MA, Samraoui B, Bennett WS, Strominger JL, Wiley DC . The foreign antigen binding site and T cell recognition regions of class I histocompatibility antigens. Nature 1987; 329: 512–518.
Garboczi DN, Ghosh P, Utz U, Fan QR, Biddison WE, Wiley DC . Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 1996; 384: 134–141.
Saper MA, Bjorkman PJ, Wiley DC . Refined structure of the human histocompatibility antigen HLA-A2 at 2.6 A resolution. J Mol Biol 1991; 219: 277–319.
Hopp TP, Woods KR . Prediction of protein antigenic determinants from amino acid sequences. Proc Natl Acad Sci USA 1981; 78: 3824–3828.
Fine JP, Gray RJ . A proportional hazards model for the subdistribution of a competing risk. J Am Stat Assoc 1999; 94: 496–509.
Pidala J, Wang T, Haagenson M, Spellman SR, Askar M, Battiwalla M et al. Amino acid substitution at peptide-binding pockets of HLA class I molecules increases risk of severe acute GVHD and mortality. Blood 2013; 122: 3651–3658.
Kawase T, Morishima Y, Matsuo K, Kashiwase K, Inoko H, Saji H et al. High-risk HLA allele mismatch combinations responsible for severe acute graft-versus-host disease and implication for its molecular mechanism. Blood 2007; 110: 2235–2241.
Lee SJ, Klein J, Haagenson M, Baxter-Lowe LA, Confer DL, Eapen M et al. High-resolution donor-recipient HLA matching contributes to the success of unrelated donor marrow transplantation. Blood 2007; 110: 4576–4583.
Shaw BE, Gooley TA, Malkki M, Madrigal JA, Begovich AB, Horowitz MM et al. The importance of HLA-DPB1 in unrelated donor hematopoietic cell transplantation. Blood 2007; 110: 4560–4566.
Acknowledgements
VK, CJT and JAB were funded by the NIHR Cambridge Biomedical Research Centre. We thank the staff of the section Immunogenetics and Transplantation Immunology of the Leiden University Medical Centre for HLA typing all donors and patients.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Bone Marrow Transplantation website
Rights and permissions
About this article
Cite this article
Kosmoliaptsis, V., Jöris, M., Mallon, D. et al. Physiochemical disparity of mismatched HLA class I alloantigens and risk of acute GVHD following HSCT. Bone Marrow Transplant 50, 540–544 (2015). https://doi.org/10.1038/bmt.2014.305
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bmt.2014.305