Abstract
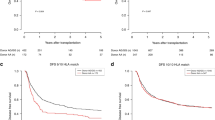
CTLA-4 is a negative regulator of activated T cells and the association of CTLA-4 polymorphisms with autoimmune diseases and transplant outcome has been reported. We evaluated the effect of donor CTLA-4 polymorphisms on outcome after allogeneic hematopoietic SCT (HSCT). We analyzed 147 Japanese HLA-matched sibling recipients and their donors who had undergone allogeneic HSCT. Genotyping of three single-nucleotide polymorphisms in CTLA-4 (−318, +49, CT60) was performed using TaqMan-PCR. According to the international HapMap database, only these three CTLA-4 haplotypes, classified as C-G-G, C-A-A and T-A-G, are present in the Japanese population. In this study, percentage expression of the C-G-G, C-A-A and T-A-G haplotypes was 59.5, 30.6 and 9.9%, respectively. Recipients of the C-A-A haplotype donor showed a significantly lower risk of relapse (HR: 0.54, 95% CI: 0.30–0.97, P=0.040) and a trend toward higher OS (HR: 0.61, 95% CI: 0.36–1.0, P=0.054) than did recipients of a donor without the C-A-A haplotype. The presence or absence of the C-A-A haplotype did not affect GVHD or non-relapse mortality. As the presence of the C-A-A haplotype reduced relapse risk and improved survival after allogeneic HSCT, this CTLA-4 haplotype may provide useful information for donor selection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ferrara JL, Deeg HJ . Graft-versus-host disease. N Engl J Med 1991; 324: 667–674.
Horowitz MM, Gale RP, Sondel PM, Goldman JM, Kersey J, Kolb HJ et al. Graft-versus-leukemia reactions after bone marrow transplantation. Blood 1990; 75: 555–562.
Dickinson AM . Non-HLA genetics and predicting outcome in HSCT. Int J Immunogenet 2008; 35: 375–380.
Hansen JA, Petersdorf EW, Lin MT, Wang S, Chien JW, Storer B et al. Genetics of allogeneic hematopoietic cell transplantation. Role of HLA matching, functional variation in immune response genes. Immunol Res 2008; 41: 56–78.
Goulmy E, Schipper R, Pool J, Blokland E, Falkenburg JH, Vossen J et al. Mismatches of minor histocompatibility antigens between HLA-identical donors and recipients and the development of graft-versus-host disease after bone marrow transplantation. N Engl J Med 1996; 334: 281–285.
Nishida T, Akatsuka Y, Morishima Y, Hamajima N, Tsujimura K, Kuzushima K et al. Clinical relevance of a newly identified HLA-A24-restricted minor histocompatibility antigen epitope derived from BCL2A1, ACC-1, in patients receiving HLA genotypically matched unrelated bone marrow transplant. Br J Haematol 2004; 124: 629–635.
Lin MT, Storer B, Martin PJ, Tseng LH, Gooley T, Chen PJ et al. Relation of an interleukin-10 promoter polymorphism to graft-versus-host disease and survival after hematopoietic-cell transplantation. N Engl J Med 2003; 349: 2201–2210.
Keen LJ, DeFor TE, Bidwell JL, Davies SM, Bradley BA, Hows JM . Interleukin-10 and tumor necrosis factor alpha region haplotypes predict transplant-related mortality after unrelated donor stem cell transplantation. Blood 2004; 103: 3599–3602.
Inamoto Y, Murata M, Katsumi A, Kuwatsuka Y, Tsujimura A, Ishikawa Y et al. Donor single nucleotide polymorphism in the CCR9 gene affects the incidence of skin GVHD. Bone Marrow Transplant 2010; 45: 363–369.
Sugimoto K, Murata M, Onizuka M, Inamoto Y, Terakura S, Kuwatsuka Y et al. Decreased risk of acute graft-versus-host disease following allogeneic hematopoietic stem cell transplantation in patients with the 5,10-methylenetetrahydrofolate reductase 677TT genotype. Int J Hematol 2008; 87: 451–458.
Ahmed S, Ihara K, Kanemitsu S, Nakashima H, Otsuka T, Tsuzaka K et al. Association of CTLA-4 but not CD28 gene polymorphisms with systemic lupus erythematosus in the Japanese population. Rheumatology (Oxford) 2001; 40: 662–667.
Hudson LL, Rocca K, Song YW, Pandey JP . CTLA-4 gene polymorphisms in systemic lupus erythematosus: a highly significant association with a determinant in the promoter region. Hum Genet 2002; 111: 452–455.
Lee YH, Harley JB, Nath SK . CTLA-4 polymorphisms and systemic lupus erythematosus (SLE): a meta-analysis. Hum Genet 2005; 116: 361–367.
Haller K, Kisand K, Pisarev H, Salur L, Laisk T, Nemvalts V et al. Insulin gene VNTR, CTLA-4 +49A/G and HLA-DQB1 alleles distinguish latent autoimmune diabetes in adults from type 1 diabetes and from type 2 diabetes group. Tissue Antigens 2007; 69: 121–127.
Balic I, Angel B, Codner E, Carrasco E, Perez-Bravo F . Association of CTLA-4 polymorphisms and clinical-immunologic characteristics at onset of type 1 diabetes mellitus in children. Hum Immunol 2009; 70: 116–120.
Kouki T, Sawai Y, Gardine CA, Fisfalen ME, Alegre ML, DeGroot LJ . CTLA-4 gene polymorphism at position 49 in exon 1 reduces the inhibitory function of CTLA-4 and contributes to the pathogenesis of Graves’ disease. J Immunol 2000; 165: 6606–6611.
Perez-Garcia A, De la Camara R, Roman-Gomez J, Jimenez-Velasco A, Encuentra M, Nieto JB et al. CTLA-4 polymorphisms and clinical outcome after allogeneic stem cell transplantation from HLA-identical sibling donors. Blood 2007; 110: 461–467.
Azarian M, Busson M, Lepage V, Charron D, Toubert A, Loiseau P et al. Donor CTLA-4 +49 A/G*GG genotype is associated with chronic GVHD after HLA-identical haematopoietic stem-cell transplantations. Blood 2007; 110: 4623–4624.
Wu J, Tang JL, Wu SJ, Lio HY, Yang YC . Functional polymorphism of CTLA-4 and ICOS genes in allogeneic hematopoietic stem cell transplantation. Clin Chim Acta 2009; 403: 229–233.
Przepiorka D, Weisdorf D, Martin P, Klingemann HG, Beatty P, Hows J et al. 1994 Consensus Conference on Acute GVHD Grading. Bone Marrow Transplant 1995; 15: 825–828.
Shulman HM, Sullivan KM, Weiden PL, McDonald GB, Striker GE, Sale GE et al. Chronic graft-versus-host syndrome in man. A long-term clinicopathologic study of 20 Seattle patients. Am J Med 1980; 69: 204–217.
Ligers A, Teleshova N, Masterman T, Huang WX, Hillert J . CTLA-4 gene expression is influenced by promoter and exon 1 polymorphisms. Genes Immun 2001; 2: 145–152.
Wang XB, Zhao X, Giscombe R, Lefvert AK . A CTLA-4 gene polymorphism at position −318 in the promoter region affects the expression of protein. Genes Immun 2002; 3: 233–234.
Sato S, Fujimoto M, Hasegawa M, Komura K, Yanaba K, Hayakawa I et al. Serum soluble CTLA-4 levels are increased in diffuse cutaneous systemic sclerosis. Rheumatology (Oxford) 2004; 43: 1261–1266.
Wong CK, Lit LC, Tam LS, Li EK, Lam CW . Aberrant production of soluble costimulatory molecules CTLA-4, CD28, CD80 and CD86 in patients with systemic lupus erythematosus. Rheumatology (Oxford) 2005; 44: 989–994.
Oaks MK, Hallett KM, Penwell RT, Stauber EC, Warren SJ, Tector AJ . A native soluble form of CTLA-4. Cell Immunol 2000; 201: 144–153.
Saverino D, Brizzolara R, Simone R, Chiappori A, Milintenda-Floriani F, Pesce G et al. Soluble CTLA-4 in autoimmune thyroid diseases: relationship with clinical status and possible role in the immune response dysregulation. Clin Immunol 2007; 123: 190–198.
Ueda H, Howson JM, Esposito L, Heward J, Snook H, Chamberlain G et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 2003; 423: 506–511.
Mäurer M, Loserth S, Kolb-Mäurer A, Ponath A, Wiese A, Kruse N et al. A polymorphism in the human cytotoxic T-lymphocyte antigen 4 (CTLA4) gene (exon 1 +49) alters T-cell activation. Immunogenetics 2002; 54: 1–8.
Oh H, Loberiza Jr FR, Zhang MJ, Ringden O, Akiyama H, Asai T et al. Comparison of graft-versus-host-disease and survival after HLA-identical sibling bone marrow transplantation in ethnic populations. Blood 2005; 105: 1408–1416.
Stem Cell Trialists’ Collaborative Group. Allogeneic peripheral blood stem-cell compared with bone marrow transplantation in the management of hematologic malignancies: an individual patient data meta-analysis of nine randomized trials. J Clin Oncol 2005; 23: 5074–5087.
Acknowledgements
This study was supported in part by a Grant-in-Aid for Scientific Research (20890096) from Japan Society for the Promotion of Science, and a Health and Labour Science Research Grant (Research on Human Genome and Tissue Engineering) from the Ministry of Health, Labour and Welfare of Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Murase, M., Nishida, T., Onizuka, M. et al. Cytotoxic T-lymphocyte antigen 4 haplotype correlates with relapse and survival after allogeneic hematopoietic SCT. Bone Marrow Transplant 46, 1444–1449 (2011). https://doi.org/10.1038/bmt.2010.319
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bmt.2010.319
Keywords
This article is cited by
-
Donor single nucleotide polymorphism in ACAT1 affects the incidence of graft-versus-host disease after bone marrow transplantation
International Journal of Hematology (2020)
-
CTLA-4 polymorphisms: influence on transplant-related mortality and survival in children undergoing allogeneic hematopoietic stem cell transplantation
Journal of Cancer Research and Clinical Oncology (2018)