Abstract
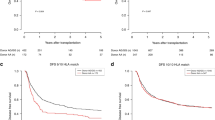
Uncertainty still exists on the role of polymorphisms outside the HLA-DRB1 binding site or inside the HLA-DRB3 binding groove in unrelated hematopoietic SCT (HSCT). The ideal model to solve the conundrum consists of the transplants mismatched for HLA-DRB1*14:01/*14:54 and/or for HLA-DRB3*02:01/*02:02. A task force was set up in Italy to recruit transplanted pairs defined as HLA-DRB1*14:01 before 2006, the year crucial for the proper definition of the HLA-DRB1*14:54 allele in molecular biology. Out of 2723 unrelated pairs, 189 transplanted in Italy from 1995 to 2006 were HLA-DRB1*14:01 positive; 103/189 pairs with good historical DNA were retyped for HLA-DRB1*14 and HLA-DRB3 at-high resolution level; 31/103 pairs had HLA-DRB1*14 and/or HLA-DRB3 mismatched; 99/103, having complete clinical data, underwent statistical analysis for OS, TRM, disease-free survival and acute and chronic GvHD. No significant involvement of HLA-DRB1*14:01/*14:54 or HLA-DRB3*02:01/*02:02 mismatches was found, either alone or combined. Our findings suggest that disparities at exon 3 of the HLA-DRB1 gene seem unlikely to influence the outcome after HSCT. The same may be envisaged for HLA-DRB3*02:01 and *02:02 alleles which, although differing in the Ag binding site, seem unable to modulate an appreciable immune response in an HSCT setting.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lee SJ, Klein J, Haagenson M, Baxter-Lowe LA, Confer DL, Eapen M et al. High-resolution donor-recipient HLA matching contributes to the success of unrelated donor marrow transplantation. Blood 2007; 110: 4576–4583.
Carreras E, Jiménez M, Gómez-García V, de la Cámara R, Martín C, Martínez F et al. Donor age and degree of HLA matching have a major impact on the outcome of unrelated donor haematopoietic cell transplantation for chronic myeloid leukaemia. Bone Marrow Transplant 2006; 37: 33–40.
Lechler R, Batchelor R, Lombardi G . The relationship between MHC restricted and allospecific T cell recognition. Immunol Lett 1991; 29: 41–50.
Brown JH, Jardetzky TS, Gorga JC, Stern LJ, Urban RG, Strominger JL et al. Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 1993; 364: 33–39.
Marsh S, Albert E, Bodmer W, Bontrop R, Dupont B, Erlich H et al. Nomenclature for factors of the HLA system, 2004. Tissue Antigens 2005; 65: 301–369.
Xiao Y, Lazaro AM, Masaberg C, Haagenson M, Vierra-Green C, Spellman S et al. Evaluating the potential impact of mismatches outside the antigen recognition site in unrelated hematopoietic stem cell transplantation: HLA-DRB1*1454 and DRB1*140101. Tissue Antigens 2009; 73: 595–598.
Yang KL, Chen MJ, Lee SK, Lin CC, Tsai MJ, Chiu HM et al. New allele name of some HLA-DRB1*1401: HLA-DRB1*1454. Int J Immunogenet 2009; 36: 119–120.
Horn PA, Albis-Camps M, Verboom M, Bunce M, Yousaf K, Williams S et al. The nature of diversity of HLA-DRB1 exon 3. Tissue Antigens 2007; 70: 335–337.
Dai S, Crawford F, Marrack P, Kappler JW . The structure of HLA-DR52c: comparison to other HLA-DRB3 alleles. Proc Natl Acad Sci USA 2008; 105: 11893–11897.
Teutsch SM, Bennetts BH, Buhler MM, Heard RN, Stewart GJ . The DRB1 Val86/Val86 genotype associates with multiple sclerosis in Australian patients. Hum Immunol 1999; 60: 715–722.
Menconi F, Monti MC, Greenberg DA, Oashi T, Osman R, Davies TF et al. Molecular amino acid signatures in the MHC class II peptide-binding pocket predispose to autoimmune thyroiditis in humans and in mice. Proc Natl Acad Sci USA 2008; 105: 14034–14039.
Emery P, Mach B, Reith W . The different level of expression of HLA-DRB1 and-DRB3 genes is controlled by conserved isotypic differences in promoter sequence. Hum Immunol 1993; 38: 137–147.
Faner R, James E, Huston L, Pujol-Borrel R, Kwok WW, Juan M . Reassessing the role of HLA-DRB3 T-cell responses: evidence for significant expression and complementary antigen presentation. Eur J Immunol 2010; 40: 91–102.
Mack SJ, Tu B, Lazaro A, Yang R, Lancaster AK, Cao K et al. HLA-A, -B, -C, and -DRB1 allele and haplotype frequencies distinguish Eastern European Americans from the general European American population. Tissue Antigens 2009; 73: 17–32.
Kaplan EL, Meier P . Non-parametric estimation from incomplete observations. J Am Stat Assoc 1958; 53: 457–481.
Glucksberg H, Storb R, Fefer A, Buckner CD, Neiman PE, Clift RA et al. Clinical manifestations of graft-versus-host disease in human recipients of marrow from HL-A-matched sibling donors. Transplantation 1974; 18: 295–304.
Cox DR . Regression models and life tables. J Royal Stat Soc 1972; 34: 187–220.
Maeng CY, Kim KH, Kang JH, Han H, Kim KL . A novel HLA-DR12 allele (DRB1*1206) found in a Korean B-cell line. Tissue Antigens 1999; 53: 516–518.
Lazaro AM, Steiner NK, Cao K, Slack R, Chen DS, Xiao Y et al. Searching for HLA-DRB1*1206 in volunteer marrow donors in four US population groups. Tissue Antigens 2006; 68: 439–441.
Lee KW, Jung YA . Additional sequence analysis outside exon 2 clarifies DRB1*12 and DRB1*14 allelic frequencies in Koreans. Hum Immunol 2009; 70: 464–467.
Fürst D, Solgi G, Schrezenmeier H, Mytilineos J . The frequency of DRB1*1454 in South German Caucasians. Tissue Antigens 2010; 76: 57–59.
Zino E, Frumento G, Marktel S, Sormani MP, Ficara F, Di Terlizzi S et al. A T-cell epitope encoded by a subset of HLA-DPB1 alleles determines non permissive mismatches for hematologic stem cell transplantation. Blood 2004; 103: 1417–1424.
Duquesnoy RJ, Askar M . HLAMatchmaker: a molecularly based algorithm for histocompatibility determination. V. Eplet matching for HLA-DR, HLA-DQ, and HLA-DP. Hum Immunol 2007; 68: 12–25.
Stern LJ, Calvo-Calle JM . HLA-DR: molecular insights and vaccine design. Curr Pharm Des 2009; 15: 3249–3261.
Duquesnoy R, Spellman S, Haagenson M, Wang T, Horowitz MM, Oudshoorn M . HLAMatchmaker-defined triplet matching is not associated with better survival rates of patients with class I HLA allele mismatched hematopoietic cell transplants from unrelated donors. Biol Blood Marrow Transplant 2008; 14: 1064–1071.
Acknowledgements
We are grateful to Gruppo Italiano Trapianti Midollo Osseo, who gave us the clinical data of the recipients, to the Italian Bone Marrow Donor Registry, who selected the pairs suitable for the analyses and to the Immunogenetics Italian Laboratories for their precious support throughout the conduct of this study. This work was supported by grants from the Associazione Italiana per la Ricerca sul Cancro, the Current Research Projects 2008–2011 and 2009–2010 (Responsible Dr M Martinetti) of the IRCCS Foundation Policlinico San Matteo of Pavia, the Cariplo Foundation and the Telethon Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Pasi, A., Crocchiolo, R., Bontempelli, M. et al. The conundrum of HLA-DRB1*14:01/*14:54 and HLA-DRB3*02:01/*02:02 mismatches in unrelated hematopoietic SCT. Bone Marrow Transplant 46, 916–922 (2011). https://doi.org/10.1038/bmt.2010.246
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bmt.2010.246
Keywords
This article is cited by
-
HLA mismatches that are identical for the antigen recognition domain are less immunogenic
Bone Marrow Transplantation (2018)
-
High-resolution HLA matching in unrelated donor transplantation in Switzerland: differential impact of class I and class II mismatches may reflect selection of nonimmunogenic or weakly immunogenic DRB1/DQB1 disparities
Bone Marrow Transplantation (2015)