Abstract
Background:
Major histocompatibility complex (MHC) class I chain-related protein A (MICA) and MHC class I chain-related protein B (MICB) are polymorphic proteins that are induced upon stress, damage or transformation of cells which act as a ‘kill me’ signal through the natural-killer group 2, member D receptor expressed on cytotoxic lymphocytes. MICA/B are not thought to be constitutively expressed by healthy normal cells but expression has been reported for most tumour types. However, it is not clear how much of this protein is expressed on the cell surface.
Methods:
Using a novel, well-characterised antibody and both standard and confocal microscopy, we systematically profiled MICA/B expression in multiple human tumour and normal tissue.
Results:
High expression of MICA/B was detected in the majority of tumour tissues from multiple indications. Importantly, MICA/B proteins were predominantly localised intracellularly with only occasional evidence of cell membrane localisation. MICA/B expression was also demonstrated in most normal tissue epithelia and predominantly localised intracellularly. Crucially, we did not observe qualitative differences in cell surface expression between tumour and MICA/B expressing normal epithelia.
Conclusions:
This demonstrates for the first time that MICA/B is more broadly expressed in normal tissue and that expression is mainly intracellular with only a small fraction appearing on the cell surface of some epithelia and tumour cells.
Similar content being viewed by others
Main
Immunosurveillance is an important part of the immune response not only against pathogens but also against cancer. As one mechanism, cytotoxic lymphocytes, including NK cells and CD8+ T cells, can detect cell surface molecules whose expression is induced in infected or transformed cells. One sensor of such ‘kill me’ signals is the natural-killer group 2, member D (NKG2D) receptor expressed on natural killer (NK), CD8+ T cells and on γδ T cells.
NKG2D ligands are reported usually not to be expressed on the cell surface of healthy cells, although a few studies have shown expression of major histocompatibility complex (MHC) class I chain-related protein A/B (MICA/B) on healthy tissues such as the gastrointestinal epithelium (Groh et al, 1996; Allegretti et al, 2013) but can be induced in response to various stressors, including chromatin remodelling, infection or transformation and render stressed cells susceptible to killing by cytotoxic cells (Raulet et al, 2013; Ullrich et al, 2013). However, NKG2D ligands, such as MICA and MICB, can also be shed from the surface of transformed cells, and soluble MICA and MICB have been reported in the sera of patients with a wide variety of malignancies (Salih et al, 2008; Zhang et al, 2015).
Shedding of NKG2D ligands leads not only to reduced cell surface ligand density and reduced NK cell recognition but also continual exposure to soluble ligands also results in downregulation of surface NKG2D expression and subsequent reduction of NK and CD8+ T-cell cytotoxicity (Groh et al, 2001; Salih et al, 2002; Coudert et al, 2005; Hanaoka et al, 2010). Moreover by inducing caspase-mediated CD3ζ degradation in T and NK cells, soluble MICA has also been shown to negatively affect other activating receptors, such as the T-cell receptor, CD16 and NKp46, thereby inducing a more general tolerisation of cytotoxic cells (Hanaoka et al, 2010). In addition, NKG2D expression has been reported on some tumours, where soluble NKG2D ligands may stimulate pro-survival signals and stimulate tumour cell proliferation (Benitez et al, 2011).
Shedding of MICA/B is mediated by endoplasmic reticulum protein 5 (ERp5), which binds to a conserved six amino acid motif in the α3-domain of MICA and MICB and induces a conformational change that allows protease cleavage, for example, by ADAM10 or ADAM17 (Kaiser et al, 2007; Waldhauer et al, 2008; Boutet et al, 2009; Wang et al, 2009).
Regulation of NKG2D ligand expression has been reported to occur at the transcriptional, post-transcriptional and post-translational levels, but is still not well understood (reviewed in Raulet et al (2013)). MicroRNAs can regulate MICA/B protein expression (Stern-Ginossar et al, 2008), while ubiquitination, palmitoylation and glycosylation have been shown to regulate intracellular retention and cell surface expression and distribution (Eleme et al, 2004; Thomas et al, 2008; Aguera-Gonzalez et al, 2011; Mellergaard et al, 2014). Moreover, MICA/B does not always reach the cell surface and can instead be released from cells through exosomes (Clayton and Tabi, 2005; Ashiru et al, 2010).
Most studies investigating expression and localisation of MICA/B have been performed using cell lines, often ones that were engineered to over-express MICA or MICB. Therefore, expression and, in particular, intracellular distribution of MICA/B in normal and tumour tissue is not well understood. Here we describe a systematic analysis of MICA/B expression in situ in a wide range of normal and tumour tissues, using a well-characterised antibody specific for the α3-domain of MICA/B that recognises a wide range of MICA and MICB alleles. MICA/B expression and sub-cellular localisation was characterised by immunohistochemistry (IHC) and confocal microscopy, which revealed intracellular expression of MICA/B in many epithelial cell types. Moreover, MICA/B was also found to be expressed by the majority of cancer samples analysed; however, we did not observe qualitative differences in cell surface expression of MICA/B between tumour cells and MICA/B-expressing epithelial cells.
Materials and methods
Antibodies
The anti-MIC mouse IgG2a antibody B1-F2A4 was a provided by Dr T Spies from the Fred Hutchinson Cancer Research Center, Seattle, WA, USA), NIP228 mouse IgG2a isotype control antibody has been previously described (Percival-Alwyn et al, 2015).
The following additional antibodies were used: mouse anti-MICA/B (clone 6D4) (BioLegend, San Diego, CA, USA), rabbit anti-CD63 pAb (ab118307) (Abcam, Cambridge, UK), rabbit anti-CD107a (LAMP1) pAb (#9091) (Cell Signaling, Danvers, MA, USA) and rabbit immunoglobulin fraction isotype control (X0936) (Dako, Ely, UK).
The following secondary conjugated antibodies were used; Rabbit anti-Mouse IgG Alexa488 (A11059) and Goat anti-Rabbit IgG Alexa594 (A11037) (Invitrogen, Thermo Fisher, Basingstoke, UK), Goat anti-Mouse IgG PE (Jackson Laboratories, Bar Harbor, ME, USA).
Cells
All cells were cultured at 37 °C in 5% CO2 in medium as recommended by the supplier. The HeLa cell line was purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA), HT29 was purchased from the Leibniz Institute DSMZ – German Collection of Microorganisms (DSMZ, Braunschweig, Germany), MDA-MB-231 and Pan02 were from the National Cancer Institute (NCI, Bethesda, MD, USA) and HCT116 was from the European Collection of Authenticated Cell Cultures (ECACC) (Porton Down, UK).
C1R MICA*004 and C1R MICB*001 cell lines were a kind gift from Dr T Spies and were cultured as described (Groh et al, 1996, 1998; Steinle et al, 2001).
Cell lines utilised for IHC were embedded in optimum cutting temperature compound (OCT, Sakura Finetek, Torrance, CA, USA) and snap frozen into pellets ready for cryo-sectioning.
Tissue
All tissue samples from both surgical and post-mortem procedures were obtained with full informed consent.
Human frozen tumour tissue micro-array (TMA) slides were purchased from Asterand Bioscience (Royston, Hertfordshire, UK) and contained duplicate samples from 10 donors in duplicate from eight tumour types; colorectal, non-small cell lung cancer (NSCLC), breast, prostatic, pancreatic, hepatocellular carcinoma (HCC), ovarian and gastric. Additional TMA slides (Tristar Technology Group LLC, (Rockville, MD, USA) comprised 10 single donors each of colorectal, ovarian, breast, lung and prostate; six donors of pancreatic; four donors of gastric; and two donors each of representative normal tissue for each tumour type.
Matching serum and tumour tissue from four gastric, two colorectal and three breast cancer donors were obtained from Tissue Solutions Ltd (Glasgow, UK).
Normal tissues were obtained from MedImmune’s (MedImmune, Cambridge, UK) internal tissue bank (sourced through Hammersmith Hospital, London, UK and Peterborough Hospital, Peterborough, UK), which included a total of eight stomach samples from two donors, 10 small intestine samples from nine donors, 11 colon samples from eight donors and an internal MedImmune normal TMA containing samples from two donors from each of 30 normal tissue types.
Normal human tissue cross reactivity TMA slides were also purchased from Asterand Bioscience (Royston, Hertfordshire, UK) and contained three donors in duplicate from 36 normal tissue types.
Flow cytometry
Cells (1 × 105) were washed twice with Flow Cytometry Staining Buffer (eBioscience, San Diego, CA, USA) and incubated with the primary antibody for 60 min on ice. Cells were washed again and incubated with PE conjugated anti-mouse IgG (Jackson Laboratories) for 30 min, washed three times and fluorescence measured using a BD FACSCanto (BD Biosciences, Franklin Lakes, NJ, USA) and analysed using FlowJo 10 software (FlowJo LLC, Ashland, OR, USA).
Luminex assay
A LifeCodes LSA-MIC Luminex kit (Gen-Probe, LSA MIC 3TL265300) was used to determine binding of B1-F2A4 and 6D4 to 28 different alleles of MICA. Because the kit is designed to measure antibodies in human serum, the assay was reconfigured to determine the binding of the B1-F2A4 and 6D4 mouse IgG antibodies using an anti-mouse IgG R-Phycoerythrin-AffiniPure F(ab’)2 fragment (Jackson ImmunoResearch).
ELISA
Soluble MICA and MICB were detected in human serum using human Duo Set ELISAs from R&D Systems (Abingdon, UK) according to the manufacturer’s protocol.
Immunohistochemistry
Frozen tissue samples were supported in OCT and cryo-sectioned at 5 μm.
Sections were briefly fixed in 10% neutral buffered formalin for 15 s and endogenous peroxidase activity blocked by 0.03% hydrogen peroxide for 10 min. Before incubation in primary antibodies, sections were incubated with 2.5% horse serum (Vector Laboratories, Burlingame, CA, USA) for 20 min. Primary antibodies were titrated across MICA/B expressing and non-expressing cell lines and tissues (pancreas, small intestine and colon) to elucidate the optimal concentration for each antibody.
Mouse primary antibodies, B1-F2A4 or NIP228 were incubated at 0.75 μg ml−1 for 1 h at room temperature, detected with anti-mouse Ig ImmPRESS-HRP reagent (Vector Laboratories) and visualised using DAB+ substrate (Dako). Nuclei were counterstained with Gill I Haematoxylin (Leica Biosystems, Nussloch, Germany), slides were dehydrated and permanently mounted with a coverslip. Slides were digitally scanned using an Aperio Scanscope XT (Leica Biosystems).
Immunofluorescence
Frozen tissue samples were supported in OCT and cryo-sectioned at 5 μm.
Sections were briefly fixed in 10% neutral buffered formalin for 15 s. Single-stain immunofluorescence was performed at room temperature using a 5% rabbit or 5% goat serum blocking step followed by incubation in either 0.75 μg ml−1 mouse B1-F2A4, 0.75 μg ml−1 mouse NIP228, 2 μg ml−1 rabbit CD63, 1 : 1000 stock dilution of rabbit CD107a or 2 μg ml−1 rabbit immunofraction antibodies for 1 h. Primary antibodies were detected and visualised using either Rabbit anti-Mouse IgG Alexa488 or Goat anti-Rabbit Alexa594 antibodies. Slides were counterstained with DAPI (Biotium, Hayward, CA, USA) and permanently mounted.
Dual stain immunofluorescence was performed at room temperature using a 2.5% bovine serum albumin protein block for 20 min followed by incubation in either 2 μg ml−1 rabbit anti-human CD63, 1 : 1000 stock dilution of rabbit CD107a or 2 μg ml−1 rabbit immunofraction antibodies for 1 h. Goat anti-Rabbit Alexa594 antibody was used to detect and visualise the rabbit primary antibodies. Rabbit serum was used to saturate any nonspecific binding potential of the anti-rabbit secondary antibody before incubation for 1 h in 0.75 μg ml−1 mouse B1-F2A4 or 0.75 μg ml−1 mouse NIP228 antibodies. Rabbit anti-Mouse IgG Alexa488 antibody was used to detect and visualise the mouse primary antibodies. Slides were counterstained with DAPI and permanently mounted.
Confocal imaging
Slides were reviewed and images taken using an Olympus FV1000 Confocal microscope (Olympus, Southend-on-Sea, UK).
Results
Characterisation of B1-F2A4
To systematically evaluate expression of MICA/B in both tumour and healthy tissue, we characterised a novel antibody for binding specificity and suitability for use in IHC.
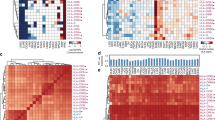
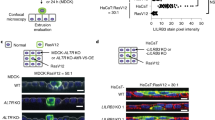
The B1-F2A4 antibody showed cell surface binding by flow cytometry to a selected panel of cell lines either transfected with MICA/B (CIR MICA*004, CIR MICB*001, CT26 MICA*004, 4T1 MICA*004, TrampC2 MICA*004) or reported to express MICA/B (HCT116, HeLa, HT29, MDA-MB-231) (Ashiru et al, 2010; Bedel et al, 2011; Chitadze et al, 2013; Schilling et al, 2015) but not to negative control mouse cell lines (Pan02, CT26, 4T1, TrampC2) (Figure 1A and Supplementary Figure 1A).
B1-F2A4 binding to cell lines and to a range of MICA alleles. (A) Indicated cell lines were stained with either B1-F2A4 or an isotype matched control antibody and analysed by flow cytometry (B) IHC staining of indicated cell lines with B1-F2A4 or 6D4 antibodies (brown=MICA/B staining/blue=cell nuclei). (C) MICA allelic coverage of B1-F2A4 and 6D4 antibodies using the Lifecodes LSATM-MIC Luminex assay. B1-F2A4 shows binding to all alleles, except MICA*043 and reduced binding to allele*46. 6D4 binds all alleles, except for MICA*036.
Antibody pull-down and subsequent western blotting experiments showed that B1-F2A4 indeed binds to MICA (Supplementary Figure 2 and Supplementary Materials and Methods). Biacore analysis confirmed that B1-F2A4 has high affinity for MICA and B, with KD values of 0.965 and 1.9 nM, respectively (Supplementary Figure 3 and Supplementary Materials and Methods).
The B1-F2A4 antibody was then validated for IHC using frozen cell pellets from these cell lines (Figure 1B and Supplementary Figure 1B) as described previously (Howat et al, 2014). The anti-human MICA antibody, 6D4, was also utilised as a comparator. Although B1-F2A4 stained all MICA/B expressing cell lines, 6D4 only showed strong staining of CIR MICA*004, CIR MICB*001, CT26 MICA*004, 4T1 MICA*004, TrampC2 MICA*004 and HeLa cells, weak staining of HT29, occasional staining of MDA-MB231 cells but no staining of HCT116 cells. No staining was observed in the negative Pan02, CT26, 4T1, and TrampC2 cell lines with either antibody.
The B1-F2A4 antibody was therefore chosen for use in studies to investigate the expression of MIC in frozen human tissue.
The MICA protein is highly polymorphic with 73 known alleles (Frigoul and Lefranc, 2005). Therefore an assay comprising recombinant MICA proteins, representing 28 of the most prevalent alleles, was used to characterise the binding of B1-F2A4 and 6D4. B1-F2A4 binds the α3 domain of MICA (data not shown), and when all 73 alleles of MICA are analysed based on their α3 domain sequence, they cluster into 15 unique α3-domain sequences (Supplementary Table 1). The assay used covers 8 of these α3 domain alleles; these are highly prevalent, covering 94–96% of the global population (Supplementary Table 1; Gonzalez-Galarza et al, 2015). As shown in Figure 1C, B1-F2A4 binds to all major alleles of MICA. Interestingly, B1-F2A4 did not bind to allele 043 and has reduced binding to allele 046, which both occur at a very low frequency in the population. Sequence comparison of the different alleles revealed that MICA*043 possesses a unique serine residue at position 256 in the α3 domain while all other alleles have an arginine residue at this position (Supplementary Table 1). This suggests that arginine 256 is critical for the binding of B1-F2A4 and has been confirmed by experiments showing reduced binding to MICA carrying mutations in this region (data not shown). The 6D4 antibody inhibits MIC-induced cytotoxicity (Das et al, 2001), therefore likely binds in the α1/2 domain that interacts with NKG2D. This antibody also shows broad MICA coverage in the same assay, binding all alleles except 036.
MICA/B is strongly expressed in major tumour types
MICA/B has been reported to be expressed in a wide range of tumour types (Zhang et al, 2015). MICA/B expression in major tumour types was analysed using both B1-F2A4 and 6D4 antibodies for IHC of TMA’s comprising samples from colorectal, NSCLC, breast, prostate, pancreatic, HCC, ovarian and gastric cancers.
B1-F2A4 staining demonstrated strong the MICA/B expression in most tumour types (Figure 2A). In contrast, no tumour staining was observed for the 6D4 antibody (Supplementary Figure 4). No specific staining was observed with the isotype control antibody (Supplementary Figure 5).
MICA/B expression in human tumour samples. Representative IHC and confocal images of eight tumour types stained with B1-F2A4 antibody. (A) Indicated tumour types were stained with the B1-F2A4 antibody and analysed by IHC. Scale bars, 100 μm. (B) IF Confocal images of selected tumour types. White arrow heads indicate occasional membrane localisation. Top images at ( × ) 400 magnification. (C) IF Confocal images of representative CRC samples from a patient with detectable soluble MICA/B (>200 pg ml−1) (left) and from a patient without detectable soluble MICA/B (right). Images at × 400 magnification. (Brown or green=MICA/B staining/blue=cell nuclei).
By IHC, 91 out of the 101 tumour samples stained with B1-F2A4 clearly demonstrated MICA/B expression (Table 1) specifically all colorectal, NSCLC, lung cancer (unknown type), breast and gastric cancers, as well as 11/12 prostate, 8/9 pancreatic, 3/6 HCC and 13/18 ovarian cancer samples demonstrated strong to moderate intracellular localisation. The amount of cell surface expression on tumour cells appeared to be occasional (⩽20%).
Because cell surface MICA/B is known to activate cytotoxic cells, including NK cells and CD8+T cells, we examined the localisation of MICA/B expression on tumour tissue using immunofluorescence (IF) with confocal microscopy. Interestingly, MICA/B expression appeared predominantly intracellular, as punctate granular staining with occasional (⩽20%) punctate cell surface expression (Figure 2B). This is in contrast to the cell surface expression of MICA/B observed on tumour cells lines using flow cytometry.
MICA and MICB have been reported to be shed from the surface of many tumours (Zhang et al, 2015). We therefore asked whether cell surface expression of MICA/B is associated with shedding. Serum samples from patients with breast, colorectal or gastric cancer were analysed for the presence of soluble MICA and MICB.
Samples with (n=5) or without (n=4) detectable levels of soluble MICA and/or MICB (>200 pg ml−1) were selected and corresponding tumour samples analysed by both IHC and IF confocal microscopy (Figure 2C and Supplementary Table 2). Interestingly, we did not detect any apparent differences in the pattern of expression, or intracellular localisation, of MICA/B between samples from patients with detectable soluble MICA/B levels and those from patients without soluble MICA/B in the serum.
MICA/B is expressed by normal tissue epithelia
While MICA/B is strongly expressed in many tumour types, it has been reported to be absent from most normal tissues, with the exception of the gastrointestinal tract (Groh et al, 1996). To confirm this, we systematically evaluated the expression of MICA/B in 36 normal tissue types utilising IHC with both the B1-F2A4 and 6D4 antibodies.
Surprisingly, B1-F2A4 antibody binding revealed strong MICA/B expression in epithelia of most tissues including: breast, colon, liver, pancreas, stomach, bronchus, bladder and ureter (Figure 3 and Table 2). Expression was also found in the epithelial cords of the thymus and squamous epithelium covering the tonsil (Supplementary Figure 6) and was detected occasionally in the Bowman’s capsule of kidney and in smooth muscle cells and/or myofibroblasts within stomach, small intestine, colon, bladder, cervix, fallopian tube, prostate and ureter. No expression was evident on other cell types. Moreover, no expression was detected in the adrenal gland, bone marrow, brain, peripheral nerves or spinal cord, eye, heart, skeletal/striated muscle, spleen or testis (Table 2). In contrast, no staining was demonstrated in normal tissue with the 6D4 antibody (Supplementary Figure 4). No specific staining was observed with the isotype control antibody (Supplementary Figure 5).
MICA/B expression in normal tissue. Representative IHC and confocal images of normal human tissue stained with B1-F2A4 antibody. (A–I) Human breast, colon, liver, pancreas, prostate, stomach, bronchus, bladder and ureter tissue stained B1-F2A4. Note: endogenous peroxidase activity present in IHC images of colon and stomach mucosal lamina propria. IHC images (top) and IF confocal images (bottom). (G–I) Occasional granular membrane localisation (white arrow heads) detected in bronchus, bladder and ureter. (J) Ureter staining with isotype control, IHC staining (top) IF Confocal staining (bottom). Scale bars indicate 100 μm, IF Confocal images at × 400 magnification. (Brown or green=MICA/B staining/blue=cell nuclei).
IF confocal microscopy was performed to determine the cellular localisation of MICA/B. Similar to what was observed in tumour tissue, the majority of MICA/B staining was found to be intracellular with only a small amount (⩽20%) of MICA/B protein detectable on the cell surface.
To confirm expression of MICA/B by epithelial cells, human mammary gland and bronchial epithelial cells were analysed by IF and showed strong, predominantly intracellular expression of MICA/B (Supplementary Figure 7A and Supplementary Materials and Methods). This was confirmed using flow cytometry and image flow cytometry using B1-F2A4 as well as 6D4. Both human mammary gland and bronchial epithelial cells showed weak expression of MICA/B on the cell surface. However, permeabilisation of the cells prior to staining with the anti-MICA/B antibody revealed strong intracellular expression of MICA/B (Supplementary Figure 7B and C and Supplementary Materials and Methods). Interestingly, while staining was observed with both test antibodies, the signal obtained with 6D4 was consistently lower than the one observed with B1-F2A4.
Ain addition, western blotting of immune-precipitates using commercially available antibodies confirmed expression of MICA/B in these epithelial cells (Supplementary Figure 7D and Supplementary Materials and Methods).
MICA/B expression co-localises with CD63 and CD107a
Because the majority of MICA/B expression does not appear to be localised on the cell surface but rather in distinct intracellular structures, and because MICA/B has been reported to be released in exosomes (Clayton and Tabi, 2005; Ashiru et al, 2010) we used an antibody to CD63 and IF confocal microscopy to investigate whether MICA/B is co-localised with CD63 in MICA/B expressing cells (Figure 4A).
MICA/B, CD63 and CD107a expression in human tumour and corresponding normal tissue. Representative IF confocal images of three tumour types and corresponding normal tissue types stained with (A) B1-F2A4 and CD63 or (B) B1-F2A4 and CD107a. White arrows indicate co-localisation of B1-F2A4 with either CD63 or CD107a. Top row images at × 400 magnification (green=MICA/B staining/red=CD63 or CD107a staining/blue=cell nuclei).
CD63 was found to be expressed in all samples of cancer and normal tissue and, as expected, was confined to discrete intracellular structures. Co-localisation with MICA/B was observed in most of the samples (7/9), however the majority of MICA/B in the cells was not co-localised with CD63 (Supplementary Table 3). Interestingly, co-localisation of CD63 and MICA/B appeared to be more frequent in healthy tissue than in cancer tissue, even though this was not statistically significant.
In additionally, co-localisation of CD63 and MICA/B was independent of whether soluble MICA/B was present in patient serum (data not shown).
We next asked whether MICA/B proteins co-localise with CD107a (LAMP-1), a marker for lysosomes. As expected, CD107a was found to be expressed in all samples of cancer and normal tissue and was confined to discrete intracellular structures (Figure 4B). Co-localisation with MICA/B was observed in all of the samples (9/9) with CD107a frequently co-localised with MICA/B (Supplementary Table 3).
Discussion
The expression of MICA and MICB proteins has been reported for many tumour types, with high expression associated with poor prognosis (Spear et al, 2013). In contrast expression of MICA and MICB has been reported to be restricted to only a few normal tissue types, including gastrointestinal tract epithelium (Groh et al, 1996; Allegretti et al, 2013), and adjacent normal mucosa from gastric cancer patients (Ribeiro et al, 2015). MICA/B is also reported to be shed from the surface of cancer cells (Salih et al, 2008; Zhang et al, 2015).
Because prolonged exposure to high concentrations of soluble MICA has been shown to blunt NKG2D activity on NK cells and CD8+ T cells (Groh et al, 2001; Salih et al, 2002; Coudert et al, 2005; Hanaoka et al, 2010), MICA/B shedding has been considered as a potential mechanism of tumour immune escape.
Soluble MICA has been proposed as a potential clinical biomarker in HCC (Kumar et al, 2012) and as target for cancer therapy (Spear et al, 2013). However, a systematic analysis of the expression of MICA and MICB in situ in healthy normal tissue as well as a wide variety of tumour types was missing due to the lack of well validated antibodies for IHC and IF confocal microscopy.
We therefore characterised a novel antibody specific for MICA/B which binds a wide range of MICA alleles at a unique epitope in the α3 domain.
Using this antibody we demonstrate for the first time that MICA/B is more broadly expressed across a wide range of normal tissue epithelia, and confirm expression by a broad range of cancer types. Importantly, we show that the majority of MICA/B staining is present in distinct intracellular structures with only occasional (<20%) membrane localisation in a few normal tissue types (bladder, bronchus, kidney and colon). Interestingly, while MICA/B is expressed by the majority of tumours from a range of indications, similar to normal tissue, MICA/B localisation was found to be predominantly intracellular with only occasionally detectable cell surface expression. While previous studies have reported strong intracellular expression of MICB in neuroblastoma (Raffaghello et al, 2004) and MICA/B in renal cell carcinoma (RCC) (Sconocchia et al, 2009) it was not clear what proportion of MICA/B proteins are cell surface expressed vs intracellular.
This pattern of expression is consistent with a recent report that showed low but detectable intracellular MICA/B expression in enterocytes in samples from healthy individuals and, importantly, large MICA/B containing aggregates oriented to the apical pole associated to the perinuclear region in mucosal samples with mild, moderate and severe enteropathy in Coeliac Disease (Allegretti et al, 2013). Additionally, Schrambach et al, have reported MICA/B expression on epithelial cells and fibroblasts in Rheumatoid Arthritis synovitis and Sjögren syndrome accessory salivary glands, respectively (Schrambach et al, 2007).
Furthermore, intracellular retention and low or absent cell surface expression of MICA has been described in several human melanoma cell lines, and this was reported to protect these cells from NKG2D mediated killing by NK cells (Fuertes et al, 2008).
The NKG2D/NKG2D ligand system is thought to have evolved as a mechanism by which cells alert the immune system to stress, transformation or infection. NKG2D ligands are expressed on the cell surface of stressed or damaged cells, and facilitate recognition and killing by NKG2D-expressing cytotoxic cells. Expression of NKG2D ligands is therefore tightly controlled on multiple levels. Epithelial cells that cover the exterior surfaces of organs are often the first site of contact with pathogens, irritants or transforming agents. They therefore need to be able to react rapidly to these stimuli and multiple mechanisms by which cells are able to do this have been described (Raulet et al, 2013). Storing pre-formed MICA/B molecules in vesicular structures intracellular would enable epithelial cells to rapidly transport MICA/B to the cell surface upon stress, infection or transformation.
The relatively low abundance of MICA/B on the cell surface was not expected, however this could be due to the relatively short half-life of MICA/B on the cell surface. MICB (Aguera-Gonzalez et al, 2009) and MICA (unpublished observation) have been shown to reside only briefly on the cell surface of overexpressing cells before being either shed from the membrane or endocytosed and recycled to internal departments. This is, however, unlikely to account entirely for the low abundance of MICA/B on the cell surface since a substantial amount of MICA/B should be present at any given time before it is recycled. MICA has also been reported to be organised into lipid rafts on the surface of MICA-expressing cell lines (Eleme et al, 2004; Aguera-Gonzalez et al, 2011), which is in line with our observation that MICA/B cell surface expression, where detectable, tends to be in discreet structures visualised as granular/punctate staining.
Since proteolytic cleavage of MICA/B from the plasma membrane is one of the major mechanisms by which tumour cells are thought to evade detection by NKG2D-expressing cytotoxic cells, we screened selected patient samples for the presence of soluble MICA or MICB in the serum. Surprisingly, using a limited number of samples, we did not see any obvious differences in the patterns of expression or cellular distribution of MICA/B between samples from patients in which we had detected soluble MICA/B and those where we did not, suggesting that shedding may not contribute significantly to the low abundance of MICA/B on the plasma membrane.
Another potential mechanism that has been proposed for tumour cell down-modulation of MICA/B expression is through release of MICA/B containing exosomes (Clayton and Tabi, 2005; Ashiru et al, 2010). CD63 is often used as a marker for exosomes, and CD63 and MICA have been detected in exosomes released from MICA-expressing cells (Ashiru et al, 2010). We observed co-localisation of CD63 with MICA/B in all tumour samples, raising the possibility that at least a small fraction of intracellular MICA/B may be released through exosomes. However, co-localisation with CD107a was more abundant showing that a good proportion of MICA/B is localised in lysosomes and could be targeted for degradation.
Interestingly, CD63 co-localisation with MICA/B appeared to be more frequent in normal tissue than in the corresponding cancer tissue. This is in line with the observation that CD63 expression is downregulated in later stages of colon and breast carcinoma and that low CD63 expression correlates with invasiveness and metastasis (Sordat et al, 2002; Sauer et al, 2003).
However, in addition to its presence in exosomes, CD63 is also abundantly present in late endosomes (Pols and Klumperman, 2009) and MICB has been reported to co-localise with CD63 in late endosomes in cell lines (Aguera-Gonzalez et al, 2009), therefore whether co-localisation of MICA/B and CD63 is indicative of release of MICA/B through exosomes remains to be determined.
Most studies have used cell lines to investigate cellular localisation and shedding of MICA/B. Similar to previous reports, we have detected MICA/B cell surface expression of all cell lines that showed intracellular MICA/B expression, suggesting that regulation of cell surface expression of MICA/B is controlled differently than in tissues. Schilling et al recently showed that hypoxia, which is common in solid tumours, induced a decrease in the membrane expression of MICA/B on some cell lines and is associated with a reduced sensitivity to NK cell-mediated lysis (Schilling et al, 2015). This suggests an alternative explanation for reduced cell surface MICA/B in tumours.
The anti-MICA antibody 6D4, used in other studies and used as a comparator in this study, may not be an optimal IHC reagent, as it stained only some of the cell lines that are known to express MICA/B in IHC (Figure 1B). Moreover, it only showed staining of some highly expressing tissue types, such as intestine. This could be due to differences in epitopes that are recognised by B1-F2A4, which has been shown to bind the α3 domain of MICA/B, and 6D4, which binds to the α1/2 domain of MICA/B (Groh et al, 1998), and which could be differentially affected by post-translational modifications such as glycosylation (Mellergaard et al, 2014).
The use of the novel B1-F2A4 antibody therefore allowed the first systematic IHC analysis of MICA/B expression in a wide range of tumour tissue as well as in healthy normal tissue, showing that MICA/B is strongly expressed in most tumour samples analysed, with predominantly intracellular localisation and only occasional cell surface localisation. Importantly, we report a similar profile for many epithelial cells in normal tissues.
References
Aguera-Gonzalez S, Boutet P, Reyburn HT, Vales-Gomez M (2009) Brief residence at the plasma membrane of the MHC class I-related chain B is due to clathrin-mediated cholesterol-dependent endocytosis and shedding. J Immunol 182: 4800–4808.
Aguera-Gonzalez S, Gross CC, Fernandez-Messina L, Ashiru O, Esteso G, Hang HC, Reyburn HT, Long EO, Vales-Gomez M (2011) Palmitoylation of MICA, a ligand for NKG2D, mediates its recruitment to membrane microdomains and promotes its shedding. Eur J Immunol 41: 3667–3676.
Allegretti YL, Bondar C, Guzman L, Cueto Rua E, Chopita N, Fuertes M, Zwirner NW, Chirdo FG (2013) Broad MICA/B expression in the small bowel mucosa: a link between cellular stress and celiac disease. PLoS One 8: e73658.
Ashiru O, Boutet P, Fernandez-Messina L, Aguera-Gonzalez S, Skepper JN, Vales-Gomez M, Reyburn HT (2010) Natural killer cell cytotoxicity is suppressed by exposure to the human NKG2D ligand MICA*008 that is shed by tumor cells in exosomes. Cancer Res 70: 481–489.
Bedel R, Thiery-Vuillemin A, Grandclement C, Balland J, Remy-Martin JP, Kantelip B, Pallandre JR, Pivot X, Ferrand C, Tiberghien P, Borg C (2011) Novel role for STAT3 in transcriptional regulation of NK immune cell targeting receptor MICA on cancer cells. Cancer Res 71: 1615–1626.
Benitez AC, Dai Z, Mann HH, Reeves RS, Margineantu DH, Gooley TA, Groh V, Spies T (2011) Expression, signaling proficiency, and stimulatory function of the NKG2D lymphocyte receptor in human cancer cells. Proc Natl Acad Sci USA 108: 4081–4086.
Boutet P, Aguera-Gonzalez S, Atkinson S, Pennington CJ, Edwards DR, Murphy G, Reyburn HT, Vales-Gomez M (2009) Cutting edge: the metalloproteinase ADAM17/TNF-alpha-converting enzyme regulates proteolytic shedding of the MHC class I-related chain B protein. J Immunol 182: 49–53.
Chitadze G, Lettau M, Bhat J, Wesch D, Steinle A, Furst D, Mytilineos J, Kalthoff H, Janssen O, Oberg HH, Kabelitz D (2013) Shedding of endogenous MHC class I-related chain molecules A and B from different human tumor entities: heterogeneous involvement of the ‘a disintegrin and metalloproteases’ 10 and 17. Int J Cancer 133: 1557–1566.
Clayton A, Tabi Z (2005) Exosomes and the MICA-NKG2D system in cancer. Blood Cells Mol Dis 34: 206–213.
Coudert JD, Zimmer J, Tomasello E, Cebecauer M, Colonna M, Vivier E, Held W (2005) Altered NKG2D function in NK cells induced by chronic exposure to NKG2D ligand-expressing tumor cells. Blood 106: 1711–1717.
Das H, Groh V, Kuijl C, Sugita M, Morita CT, Spies T, Bukowski JF (2001) MICA engagement by human Vgamma2Vdelta2 T cells enhances their antigen-dependent effector function. Immunity 15: 83–93.
Eleme K, Taner SB, Onfelt B, Collinson LM, McCann FE, Chalupny NJ, Cosman D, Hopkins C, Magee AI, Davis DM (2004) Cell surface organization of stress-inducible proteins ULBP and MICA that stimulate human NK cells and T cells via NKG2D. J Exp Med 199: 1005–1010.
Frigoul A, Lefranc MP (2005) MICA: standardized IMGT allele nomenclature, polymorphisms and diseases. Recent Res Devel Human Genet 3: 95–145.
Fuertes MB, Girart MV, Molinero LL, Domaica CI, Rossi LE, Barrio MM, Mordoh J, Rabinovich GA, Zwirner NW (2008) Intracellular retention of the NKG2D ligand MHC class I chain-related gene A in human melanomas confers immune privilege and prevents NK cell-mediated cytotoxicity. J Immunol 180: 4606–4614.
Gonzalez-Galarza FF, Takeshita LY, Santos EJ, Kempson F, Maia MH, da Silva AL, Teles e Silva AL, Ghattaoraya GS, Alfirevic A, Jones AR, Middleton D (2015) Allele frequency net 2015 update: new features for HLA epitopes, KIR and disease and HLA adverse drug reaction associations. Nucleic Acids Res 43: D784–D788.
Groh V, Bahram S, Bauer S, Herman A, Beauchamp M, Spies T (1996) Cell stress-regulated human major histocompatibility complex class I gene expressed in gastrointestinal epithelium. Proc Natl Acad Sci USA 93: 12445–12450.
Groh V, Rhinehart R, Randolph-Habecker J, Topp MS, Riddell SR, Spies T (2001) Costimulation of CD8alphabeta T cells by NKG2D via engagement by MIC induced on virus-infected cells. Nat Immunol 2: 255–260.
Groh V, Steinle A, Bauer S, Spies T (1998) Recognition of stress-induced MHC molecules by intestinal epithelial gammadelta T cells. Science 279: 1737–1740.
Hanaoka N, Jabri B, Dai Z, Ciszewski C, Stevens AM, Yee C, Nakakuma H, Spies T, Groh V (2010) NKG2D initiates caspase-mediated CD3zeta degradation and lymphocyte receptor impairments associated with human cancer and autoimmune disease. J Immunol 185: 5732–5742.
Howat WJ, Lewis A, Jones P, Kampf C, Ponten F, van der Loos CM, Gray N, Womack C, Warford A (2014) Antibody validation of immunohistochemistry for biomarker discovery: recommendations of a consortium of academic and pharmaceutical based histopathology researchers. Methods 70: 34–38.
Kaiser BK, Yim D, Chow IT, Gonzalez S, Dai Z, Mann HH, Strong RK, Groh V, Spies T (2007) Disulphide-isomerase-enabled shedding of tumour-associated NKG2D ligands. Nature 447: 482–486.
Kumar V, Yi Lo PH, Sawai H, Kato N, Takahashi A, Deng Z, Urabe Y, Mbarek H, Tokunaga K, Tanaka Y, Sugiyama M, Mizokami M, Muroyama R, Tateishi R, Omata M, Koike K, Tanikawa C, Kamatani N, Kubo M, Nakamura Y, Matsuda K (2012) Soluble MICA and a MICA variation as possible prognostic biomarkers for HBV-induced hepatocellular carcinoma. PLoS One 7: e44743.
Mellergaard M, Skovbakke SL, Schneider CL, Lauridsen F, Andresen L, Jensen H, Skov S (2014) N-glycosylation of asparagine 8 regulates surface expression of major histocompatibility complex class I chain-related protein A (MICA) alleles dependent on threonine 24. J Biol Chem 289: 20078–20091.
Percival-Alwyn JL, England E, Kemp B, Rapley L, Davis NH, McCarthy GR, Majithiya JB, Corkill DJ, Welsted S, Minton K, Cohen ES, Robinson MJ, Dobson C, Wilkinson TC, Vaughan TJ, Groves MA, Tigue NJ (2015) Generation of potent mouse monoclonal antibodies to self-proteins using T-cell epitope ‘tags’. MAbs 7: 129–137.
Pols MS, Klumperman J (2009) Trafficking and function of the tetraspanin CD63. Exp Cell Res 315: 1584–1592.
Raffaghello L, Prigione I, Airoldi I, Camoriano M, Levreri I, Gambini C, Pende D, Steinle A, Ferrone S, Pistoia V (2004) Downregulation and/or release of NKG2D ligands as immune evasion strategy of human neuroblastoma. Neoplasia 6: 558–568.
Raulet DH, Gasser S, Gowen BG, Deng W, Jung H (2013) Regulation of ligands for the NKG2D activating receptor. Annu Rev Immunol 31: 413–441.
Ribeiro CH, Kramm K, Galvez-Jiron F, Pola V, Bustamante M, Contreras HR, Sabag A, Garrido-Tapia M, Hernandez CJ, Zuniga R, Collazo N, Sotelo PH, Morales C, Mercado L, Catalan D, Aguillon JC, Molina MC (2015) Clinical significance of tumor expression of major histocompatibility complex class I-related chains A and B (MICA/B) in gastric cancer patients. Oncol Rep 35: 1309–1317.
Salih HR, Holdenrieder S, Steinle A (2008) Soluble NKG2D ligands: prevalence, release, and functional impact. Front Biosci 13: 3448–3456.
Salih HR, Rammensee HG, Steinle A (2002) Cutting edge: down-regulation of MICA on human tumors by proteolytic shedding. J Immunol 169: 4098–4102.
Sauer G, Kurzeder C, Grundmann R, Kreienberg R, Zeillinger R, Deissler H (2003) Expression of tetraspanin adaptor proteins below defined threshold values is associated with in vitro invasiveness of mammary carcinoma cells. Oncol Rep 10: 405–410.
Schilling D, Tetzlaff F, Konrad S, Li W, Multhoff G (2015) A hypoxia-induced decrease of either MICA/B or Hsp70 on the membrane of tumor cells mediates immune escape from NK cells. Cell Stress Chaperones 20: 139–147.
Schrambach S, Ardizzone M, Leymarie V, Sibilia J, Bahram S (2007) In vivo expression pattern of MICA and MICB and its relevance to auto-immunity and cancer. PLoS One 2: e518.
Sconocchia G, Spagnoli GC, Del Principe D, Ferrone S, Anselmi M, Wongsena W, Cervelli V, Schultz-Thater E, Wyler S, Carafa V, Moch H, Terracciano L, Tornillo L (2009) Defective infiltration of natural killer cells in MICA/B-positive renal cell carcinoma involves beta(2)-integrin-mediated interaction. Neoplasia 11: 662–671.
Sordat I, Decraene C, Silvestre T, Petermann O, Auffray C, Pietu G, Sordat B (2002) Complementary DNA arrays identify CD63 tetraspanin and alpha3 integrin chain as differentially expressed in low and high metastatic human colon carcinoma cells. Lab Invest 82: 1715–1724.
Spear P, Wu MR, Sentman ML, Sentman CL (2013) NKG2D ligands as therapeutic targets. Cancer Immun 13: 8.
Steinle A, Li P, Morris DL, Groh V, Lanier LL, Strong RK, Spies T (2001) Interactions of human NKG2D with its ligands MICA, MICB, and homologs of the mouse RAE-1 protein family. Immunogenetics 53: 279–287.
Stern-Ginossar N, Gur C, Biton M, Horwitz E, Elboim M, Stanietsky N, Mandelboim M, Mandelboim O (2008) Human microRNAs regulate stress-induced immune responses mediated by the receptor NKG2D. Nat Immunol 9: 1065–1073.
Thomas M, Wills M, Lehner PJ (2008) Natural killer cell evasion by an E3 ubiquitin ligase from Kaposi's sarcoma-associated herpesvirus. Biochem Soc Trans 36: 459–463.
Ullrich E, Koch J, Cerwenka A, Steinle A (2013) New prospects on the NKG2D/NKG2DL system for oncology. Oncoimmunology 2: e26097.
Waldhauer I, Goehlsdorf D, Gieseke F, Weinschenk T, Wittenbrink M, Ludwig A, Stevanovic S, Rammensee HG, Steinle A (2008) Tumor-associated MICA is shed by ADAM proteases. Cancer Res 68: 6368–6376.
Wang X, Lundgren AD, Singh P, Goodlett DR, Plymate SR, Wu JD (2009) An six-amino acid motif in the alpha3 domain of MICA is the cancer therapeutic target to inhibit shedding. Biochem Biophys Res Commun 387: 476–481.
Zhang J, Basher F, Wu JD (2015) NKG2D ligands in tumor immunity: two sides of a coin. Front Immunol 6: 97.
Acknowledgements
We thank Meggan Czapiga and John Travers for their expert technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
HG is a full-time employee of MedImmune Ltd. LB is a full-time employee of MedImmune Ltd. Chris Lloyd is a full-time employee of MedImmune Ltd. Leeanne Lewis is a full-time employee of MedImmune Ltd. Arthur Lewis is a full-time employee of MedImmune Ltd. Janette Dillon is a full-time employee of MedImmune Ltd. Richard Sainson is a full-time employee of MedImmune Ltd. Jelena Jovanovic is a full-time employee of MedImmune Ltd. Natalie J. Tigue is a full-time employee of MedImmune Ltd. David Bannister is a full-time employee of MedImmune Ltd. Lisa Bamber is a full-time employee of MedImmune Ltd. Viia Valge-Archer is a full-time employee of AstraZeneca Ltd. Robert W. Wilkinson is a full-time employee of MedImmune Ltd.
Additional information
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-Non-Commercial-Share Alike 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Ghadially, H., Brown, L., Lloyd, C. et al. MHC class I chain-related protein A and B (MICA and MICB) are predominantly expressed intracellularly in tumour and normal tissue. Br J Cancer 116, 1208–1217 (2017). https://doi.org/10.1038/bjc.2017.79
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.79
Keywords
This article is cited by
-
Bystander activated CD8+ T cells mediate neuropathology during viral infection via antigen-independent cytotoxicity
Nature Communications (2024)
-
Novel role of immune-related non-coding RNAs as potential biomarkers regulating tumour immunoresponse via MICA/NKG2D pathway
Biomarker Research (2023)
-
αVEGFR2-MICA fusion antibodies enhance immunotherapy effect and synergize with PD-1 blockade
Cancer Immunology, Immunotherapy (2023)
-
The past, present, and future of immunotherapy for colorectal cancer
Medical Oncology (2023)
-
Major histocompatibility complex (MHC) associations with diseases in ethnic groups of the Arabian Peninsula
Immunogenetics (2021)