Abstract
Background:
Prostate cancer (PCa) screening currently relies on prostate-specific antigen (PSA) testing and digital rectal examination. However, recent large-scale studies have questioned the long-term efficacy of these tests, and biomarkers that accurately identify PCa are needed.
Methods:
We analysed the levels of circulating microRNAs (miRNAs) in patients with elevated PSA who were diagnosed with either localised PCa (n=36) or benign prostatic hyperplasia (BPH, n=31) upon biopsy. Real-time RT–PCR with Taqman probes was used to measure plasma levels of miRNAs. To circumvent problems associated with circulating miRNA quantitation, we computed the expression ratios of upregulated and downregulated miRNAs.
Results:
The miR-106a/miR-130b and miR-106a/miR-223 ratios were significantly different between the biopsy-positive and BPH groups (P<0.0001), and yielded statistical power values that were >0.99. Both miRNA ratios were highly sensitive and more specific than PSA in discriminating localised PCa from BPH. Receiver operating characteristic curve analysis revealed area under curve values of 0.81 (miR-106a/miR-130b) and 0.77 (miR-106a/miR-223).
Conclusions:
Testing for circulating miR-106a/miR-130b and miR-106a/miR-223 ratios may reduce the costs and morbidity of unnecessary biopsies and is feasible for large-scale screening, as it requires measuring only three miRNAs.
Similar content being viewed by others
Main
Prostate cancer (PCa) is the most frequent cancer in males in Europe and North America. Digital rectal examination (DRE) and serum prostate-specific antigen (PSA) monitoring followed by serial biopsy and histological analysis is the cornerstone of PCa screening/diagnosis. However, although very high PSA values indicate a high probability of cancer detection by biopsy, lower values poorly discriminate PCa from inflammatory processes and benign hyperplasia (Ploussard et al, 2013; Hayes and Barry, 2014). On the basis of the results of two large epidemiological studies showing a very small or no effect of PSA testing on PCa mortality (Andriole et al, 2012; Schroder et al, 2012) and the negative consequences of PSA screening, which include overdiagnosis, overtreatment and treatment complications, the US Preventive Services Task Force recommended against the use of PSA testing for PCa screening (Aly et al, 2015; Johnson et al, 2015).
These considerations have engendered an interest in the identification of novel markers of PCa with improved specificity compared with PSA. Circulating microRNAs (miRNAs; Bartel, 2004; Waltering et al, 2011; Kim and Kim, 2013) may represent excellent biomarkers because of their ease of detection, stability in biological fluids and minimal invasiveness of the test. Different studies identified specific miRNA signatures associated with PCa (Mahn et al, 2011; Zhang et al, 2011; Chen et al, 2012; Kelly et al, 2015; Mihelich et al, 2015), although a clear consensus has not yet emerged. Discrepant findings may in part reflect differences in pre-analytical sample processing and data normalisation/analysis.
In the present study, we explored the efficacy of circulating miRNA as biomarkers for PCa. To circumvent problems linked to data normalisation, we calculated ratios of upregulated and downregulated miRNAs. Results indicated that the miR-106a/miR-130b and miR-106a/miR-223 ratios are highly sensitive and more specific than PSA in discriminating localised PCa from benign prostatic hyperplasia (BPH).
Materials and methods
Patients
A total of 67 patients were recruited; 36 were positive for PCa upon biopsy (BIO+) and 31 were negative (BIO−). BIO+ and BIO− patients were comparable in terms of mean age (65.2 and 66.7 years, respectively) and PSA (8.5 and 8.4 ng ml−1, respectively). All the BIO+ patients had localised PCa, and all of the BIO− patients were diagnosed with BPH. Additional clinical characteristics of the patients are described in the Supplementary Materials and Methods and Supplementary Table S1.
Quantitative RT–PCR
RNA was extracted from plasma as described in the Supplementary Materials and Methods. Five microlitres of total RNA were used for first-strand cDNA synthesis in a 15-μl reaction volume, using the TaqMan miRNA Reverse Transcription kit and miRNA-specific stem-loop primers (Thermo Fisher Scientific, Foster City, CA, USA). cDNAs (2.5 μl) were amplified for 45 cycles using TaqMan miRNA primers and probes (Thermo Fisher Scientific) and LightCycler 480 PCR Master Mix (Roche Diagnostics GmbH, Mannheim, Germany). No-RT and no-template negative controls were included. The amplification reactions were performed in a LightCycler 480 II (Roche). Signals were quantified using the second derivative maximum method (Software Version 1.5, Roche). All amplification curves were carefully examined, and products with poor curves were discarded. Ct values <40 were obtained for >99% of the reaction samples.
Data normalisation
As candidate normalisers we tested spiked-in cel-miR-39 as well as human miR-24, which exhibited low variability in our data set (data not shown) and was used as a normaliser in other studies (Bianchi et al, 2011).
Relative expression of target miRNAs was calculated as ΔCtmiR=CtmiR−Ctnormalizer. Differential expression of each miRNA was then evaluated as ΔΔCtmiR=ΔCtmiR(patient)− i.e., by subtracting the mean ΔCt value for the BIO− group from the ΔCt of each BIO+ patient.
i.e., by subtracting the mean ΔCt value for the BIO− group from the ΔCt of each BIO+ patient.
To circumvent biases linked to the choice of an appropriate reference, we also used the miRNA ratio approach (Boeri et al, 2011). The Ct value of each miRNA was converted to the corresponding expression level (2−Ct). Twenty-one different miRNA ratios were calculated as 
Statistical analysis
The two-tailed Wilcoxon rank-sum test was used to identify miRNAs significantly different between the BIO+ and BIO− groups, whereas the one-tailed Wilcoxon rank-sum test was used for miRNA ratios. Post hoc power analysis was performed using a nonparametric resampling technique (10 000 replications). A univariate logistic regression model was built to evaluate the ability of each miRNA ratio to predict the BIO+ outcome. The two most significant miRNA ratios were also used in a bivariate logistic regression model. Odds ratios and 95% confidence intervals were estimated for the fitted logistic regression models. Receiver operating characteristic (ROC) curves were plotted and the area under the ROC curve (AUC) was estimated to compare the two most significant miRNA ratios. Optimal thresholds were determined using the Youden’s J statistic. Sensitivity, specificity, negative predictive value, positive predictive value and accuracy were determined. Statistical analyses were performed in the R environment, using a customised code and the pROC package for ROC curve analysis.
Results
Circulating miRNA in PCa and BPH patients
We examined 36 patients with localised PCa confirmed by serial biopsy (BIO+) and 31 individuals diagnosed with BPH (BIO−). Clinical assessment of the patients included age, PSA, DRE, Gleason scoring and T stage (Supplementary Table S1). Twelve miRNAs were selected among those reported to be involved in PCa (Supplementary Table S2). To facilitate their application to large-scale testing, we focused on miRNAs that could be detected with a single round of qRT–PCR (i.e., without a pre-amplification step).
Several studies underscored the critical importance of data normalisation when measuring circulating miRNAs. Pilot tests with the cel-miR-39 spike-in reference showed a wide variability in Ct values (Supplementary Figure S1), possibly due to differences in plasma RNAase levels, as previously reported (Mahn et al, 2011). We thus normalised each miRNA using as internal reference miR-24, which was relatively constant in our samples and was used as a normaliser in other studies (Bianchi et al, 2011). Results showed that miR-106a, miR-20a, miR-223 and miR-21 were significantly different between BIO+ and BIO− individuals (Figure 1).
Boxplots of the distribution of –ΔΔCt values for circulating miR-20a, miR-21, miR-93, miR-106a, miR-130b and miR-223 in patients with localised PCa vs biopsy-negative individuals. Each dot represents a patient sample. –ΔΔCt values (see Materials and methods) were calculated using endogenous miR-24 as a reference. Differences between BIO+ and BIO− samples were analysed using two-tailed Wilcoxon rank-sum test. miR-106a, miR-223, miR-20a and miR-21 showed P-values<0.05.
miRNA ratios distinguish PCa (BIO+) from BPH (BIO−) patients
To overcome the bias linked to data normalisation using a calibrator RNA, we next calculated the ratios between the expression values of miRNA pairs listed in Supplementary Table S3. Fourteen ratios were found to be significantly different between the BIO+ and BIO− groups with P<0.05. We focused on the miR-106a/miR-130b and miR-106a/miR-223 ratios (Figure 2A), which ranked highest in terms of statistical significance (P-values were <0.0001) and showed a statistical power >0.99 (see Supplementary Table S3 for details). As shown in Figure 1, miR-106a was upregulated in BIO+ samples, whereas miR-130b and miR-223 were downregulated.
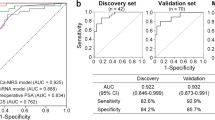
Circulating miR-106a/miR-130b and miR-106a/miR-223 ratios predict the presence of localised prostate cancer. (A) Relative expression levels are shown as 2−ΔCt (see Materials and methods). Both miRNA ratios were significantly higher (one-tailed Wilcoxon rank-sum test) in PCa patients compared with biopsy-negative patients (P=6.97 × 10−6 for miR-106a/miR-130b; P=8.5 × 10−5 for miR-106a/miR-223). (B) Receiver operating characteristic (ROC) curves generated for univariate (red and blue traces) and bivariate (black trace) logistic regression models against the ROC curve of PSA (green trace). The miRNA ratios were better than PSA for discriminating PCa from BIO− samples. Areas under ROC curves (AUC) are shown in parentheses. (C) Proposed miRNA-based diagnostic flowchart for PCa diagnosis.
These miRNA ratios were next evaluated by univariate logistic regression analysis. Resulting areas under ROC curves (AUC) were 0.81 and 0.77 for miR-106a/miR-130b and miR-106a/miR-223, respectively (Figure 2B); bivariate logistic regression analysis using both miRNA ratios as predictors showed an AUC of 0.84 (Figure 2B). All these values were considerably higher than the AUC of 0.56 obtained for PSA, a value consistent with other studies (Guzel et al, 2015). Odds ratios are reported in Supplementary Tables S4 and S5.
Determining threshold values for miRNA ratios
Statistical analysis indicated optimal 2−ΔCt threshold values of 10.413 for miR-106a/miR-130b and 0.176 for miR-106a/miR-223. To facilitate application in clinical practice, results of the bivariate logistic regression model were transformed into a miRNA score=0.1855 × (miR-106a/miR-130b)+2.3162 × (miR-106a/miR-223), where the miRNA ratios indicate 2−ΔCt values. The optimal threshold for this miRNA score was 2.142. With these cutoff values, specificity and sensitivity were 0.87 and 0.72 (miR-106a/130b), 0.65 and 0.81 (miR-106a/223), and 0.81 and 0.83 (miRNA score), respectively.
A diagnostic flowchart for PCa screening
On the basis of these findings, we propose a flowchart for the selection of patients to be subjected to biopsy (Figure 2C). Patients with PSA >4 ng ml−1 (right-hand portion of the chart) would first be tested for miR-106a/miR-130b; patients with a 2−ΔCt>10.413 would be eligible for biopsy, whereas patients below this threshold would be tested for the miR-106a/miR-223 ratio. Patients with a miR-106a/miR-223 2−ΔCt>0.176 would be biopsied, whereas patients below this threshold would not.
The left-hand portion of the flowchart refers to patients with PSA<4 ng ml−1, but DRE+. In our data set, among the six patients with these characteristics, the miR-106a/miR-130b ratio correctly distinguished all four BIO+ from the two BIO− patients, generating no false-positive or false-negative results. Table 1 compares the diagnostic performance of the proposed biomarkers with PSA.
Conclusions
Recent large-scale studies found no or very small benefit for periodic PSA and DRE screening after up to 13 years of follow-up (Andriole et al, 2012; Schroder et al, 2012); identification of other biomarkers to aid early diagnosis and biopsy decision is thus in high demand.
In this study, we explored the efficacy of circulating miRNA as biomarkers for PCa screening. Our findings suggest that a single miRNA might be poorly informative, especially considering the known biases linked to data normalisation. We thus propose to calculate ratios between one upregulated miRNA and one downregulated miRNA in the same patient. Results showed that the miR-106a/miR-130b and miR-106a/miR-223 ratios discriminated between localised PCa and BPH patients with a specificity much superior to PSA. A diagnostic flowchart that employs two miR ratios in combination with standard PSA (Figure 2C) should considerably increase the specificity and also improve the sensitivity compared with PSA testing alone (Table 1).
The miRNA ratio approach described in the present study may be applied to large-scale screening, as it requires measuring only three circulating miRNAs and overcomes the need for a normaliser miRNA. The increased specificity of this assay compared with PSA alone could help to reduce the costs and morbidity of unnecessary repeated biopsies. Future studies should test the validity of this approach in an ample prospective cohort of patients.
Change history
14 June 2016
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Aly M, Dyrdak R, Nordstrom T, Jalal S, Weibull CE, Giske CG, Gronberg H (2015) Rapid increase in multidrug-resistant enteric bacilli blood stream infection after prostate biopsy–A 10-year population-based cohort study. Prostate 75: 947–956.
Andriole GL, Crawford ED, Grubb RL 3rd, Buys SS, Chia D, Church TR, Fouad MN, Isaacs C, Kvale PA, Reding DJ, Weissfeld JL, Yokochi LA, O’Brien B, Ragard LR, Clapp JD, Rathmell JM, Riley TL, Hsing AW, Izmirlian G, Pinsky PF, Kramer BS, Miller AB, Gohagan JK, Prorok PC (2012) Prostate cancer screening in the randomized Prostate, Lung, Colorectal, and Ovarian Cancer Screening Trial: mortality results after 13 years of follow-up. J Natl Cancer Inst 104: 125–132.
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: 281–297.
Bianchi F, Nicassio F, Marzi M, Belloni E, Dall’olio V, Bernard L, Pelosi G, Maisonneuve P, Veronesi G, Di Fiore PP (2011) A serum circulating miRNA diagnostic test to identify asymptomatic high-risk individuals with early stage lung cancer. EMBO Mol Med 3: 495–503.
Boeri M, Verri C, Conte D, Roz L, Modena P, Facchinetti F, Calabro E, Croce CM, Pastorino U, Sozzi G (2011) MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc Natl Acad Sci USA 108: 3713–3718.
Chen ZH, Zhang GL, Li HR, Luo JD, Li ZX, Chen GM, Yang J (2012) A panel of five circulating microRNAs as potential biomarkers for prostate cancer. Prostate 72: 1443–1452.
Guzel E, Karatas OF, Semercioz A, Ekici S, Aykan S, Yentur S, Creighton CJ, Ittmann M, Ozen M (2015) Identification of microRNAs differentially expressed in prostatic secretions of patients with prostate cancer. Int J Cancer 136: 875–879.
Hayes JH, Barry MJ (2014) Screening for prostate cancer with the prostate-specific antigen test: a review of current evidence. JAMA 311: 1143–1149.
Johnson JR, Polgreen PM, Beekmann SE (2015) Transrectal prostate biopsy-associated prophylaxis and infectious complications: report of a query to the emerging infections network of the infectious diseases society of America. Open Forum Infect Dis 2: ofv002.
Kelly BD, Miller N, Sweeney KJ, Durkan GC, Rogers E, Walsh K, Kerin MJ (2015) A Circulating microRNA Signature as a Biomarker for Prostate Cancer in a High Risk Group. J Clin Med 4: 1369–1379.
Kim WT, Kim WJ (2013) MicroRNAs in prostate cancer. Prostate Int 1: 3–9.
Mahn R, Heukamp LC, Rogenhofer S, von Ruecker A, Muller SC, Ellinger J (2011) Circulating microRNAs (miRNA) in serum of patients with prostate cancer. Urology 77: 1265.E9–16.
Mihelich BL, Maranville JC, Nolley R, Peehl DM, Nonn L (2015) Elevated serum microRNA levels associate with absence of high-grade prostate cancer in a retrospective cohort. PLoS One 10 (4): e0124245.
Ploussard G, Nicolaiew N, Marchand C, Terry S, Allory Y, Vacherot F, Abbou CC, Salomon L, de la Taille A (2013) Risk of repeat biopsy and prostate cancer detection after an initial extended negative biopsy: longitudinal follow-up from a prospective trial. BJU Int 111: 988–996.
Schroder FH, Hugosson J, Roobol MJ, Tammela TL, Ciatto S, Nelen V, Kwiatkowski M, Lujan M, Lilja H, Zappa M, Denis LJ, Recker F, Paez A, Maattanen L, Bangma CH, Aus G, Carlsson S, Villers A, Rebillard X, van der Kwast T, Kujala PM, Blijenberg BG, Stenman UH, Huber A, Taari K, Hakama M, Moss SM, de Koning HJ, Auvinen A (2012) Prostate-cancer mortality at 11 years of follow-up. N Engl J Med 366: 981–990.
Waltering KK, Porkka KP, Jalava SE, Urbanucci A, Kohonen PJ, Latonen LM, Kallioniemi OP, Jenster G, Visakorpi T (2011) Androgen regulation of micro-RNAs in prostate cancer. Prostate 71: 604–614.
Zhang HL, Yang LF, Zhu Y, Yao XD, Zhang SL, Dai B, Zhu YP, Shen YJ, Shi GH, Ye DW (2011) Serum miRNA-21: elevated levels in patients with metastatic hormone-refractory prostate cancer and potential predictive factor for the efficacy of docetaxel-based chemotherapy. Prostate 71: 326–331.
Acknowledgements
This study was supported by funds from the Antonveneta Foundation and Istituto Oncologico Veneto (Translational Medicine projects). We thank Alessandra R. Brazzale for helpful discussions.
Author contributors
VC supervised the study. ES and KR performed the RT-PCR analysis. AG designed and performed the statistical analysis. AM, FP, PB, FZ and MI provided blood samples and were responsible for the clinical aspects of the project. PZ provided guidance and support for the study. VC, DMD, ES and AG prepared the manuscript. All the authors contributed to the study design and data interpretation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Sharova, E., Grassi, A., Marcer, A. et al. A circulating miRNA assay as a first-line test for prostate cancer screening. Br J Cancer 114, 1362–1366 (2016). https://doi.org/10.1038/bjc.2016.151
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2016.151
Keywords
This article is cited by
-
The promising role of new molecular biomarkers in prostate cancer: from coding and non-coding genes to artificial intelligence approaches
Prostate Cancer and Prostatic Diseases (2022)
-
Cell-free nucleic acid patterns in disease prediction and monitoring—hype or hope?
EPMA Journal (2020)
-
A coordinate deregulation of microRNAs expressed in mucosa adjacent to tumor predicts relapse after resection in localized colon cancer
Molecular Cancer (2018)
-
A Plasma Biomarker Panel of Four MicroRNAs for the Diagnosis of Prostate Cancer
Scientific Reports (2018)
-
Applications of Extracellular RNAs in Oncology
Molecular Diagnosis & Therapy (2017)