Abstract
Background:
Lung cancer has the highest mortality of all cancers. The aim of this study was to examine DNA hypermethylation in sputum and validate its diagnostic accuracy for lung cancer.
Methods:
DNA hypermethylation of RASSF1A, APC, cytoglobin, 3OST2, PRDM14, FAM19A4 and PHACTR3 was analysed in sputum samples from symptomatic lung cancer patients and controls (learning set: 73 cases, 86 controls; validation set: 159 cases, 154 controls) by quantitative methylation-specific PCR. Three statistical models were used: (i) cutoff based on Youden’s J index, (ii) cutoff based on fixed specificity per marker of 96% and (iii) risk classification of post-test probabilities.
Results:
In the learning set, approach (i) showed that RASSF1A was best able to distinguish cases from controls (sensitivity 42.5%, specificity 96.5%). RASSF1A, 3OST2 and PRDM14 combined demonstrated a sensitivity of 82.2% with a specificity of 66.3%. Approach (ii) yielded a combination rule of RASSF1A, 3OST2 and PHACTR3 (sensitivity 67.1%, specificity 89.5%). The risk model (approach iii) distributed the cases over all risk categories. All methods displayed similar and consistent results in the validation set.
Conclusions:
Our findings underscore the impact of DNA methylation markers in symptomatic lung cancer diagnosis. RASSF1A is validated as diagnostic marker in lung cancer.
Similar content being viewed by others
Main
Lung cancer has the highest mortality rate of all cancers, because of the presence of metastases at time of presentation (Siegel et al, 2012). Since the 1970s, the average overall five-year survival rate hovers at 15%, despite new insights in therapeutic strategies (Siegel et al, 2012). For late-stage disease, treatment options remain limited and of palliative intent. However, prognosis improves considerably when lung cancer is detected at stage I or II, where patients are treated with curative intent (Patz et al, 2000).
Currently, lung cancer is detected and staged by imaging techniques. Ideally, the diagnosis is pathologically confirmed. Therefore, tumour tissue needs to be obtained through invasive methods, such as bronchoscopy or transthoracic needle aspiration. In daily practice, this is not always possible, due to for instance, localisation of the tumour or physical stress for the patient.
Thus, there is need for a novel diagnostic method. The use of sputum is of interest, as procurement is non-invasive, inexpensive and simple. Tumour cells and tumour DNA are shed in the respiratory epithelial lining fluid and usually make up <1% of sputum composition. Sputum cytology has a low sensitivity of 66% (range 42–97%) for lung cancer diagnosis (Rivera et al, 2013). More promising is the application of molecular techniques that are able to detect minimal amounts of aberrant tumour DNA in sputum (Honorio et al, 2003; Shivapurkar et al, 2007).
DNA promoter hypermethylation of tumour-suppressor genes leads to transcriptional silencing (Esteller, 2011). Previous research has shown that various genes are hypermethylated in lung cancer patients as opposed to controls and can be detected in sputum (Belinsky et al, 2005; Cirincione et al, 2006; Shivapurkar et al, 2007). In a preliminary study, we investigated DNA hypermethylation of biomarkers RASSF1A, APC and cytoglobin (CYGB) in sputum of lung cancer patients (Hubers et al, 2014). A novel classification system for lung cancer prediction was introduced, which proved to be reproducible in two independent sets of subjects. In particular, hypermethylated RASSF1A demonstrated to have potential as a diagnostic marker.
Here, we report on an independent validation of these and additional novel discovered biomarkers (Shivapurkar et al, 2007; Steenbergen et al, 2013) in an external cohort of prospectively collected sputum obtained from lung cancer patients and cancer-free controls. In addition, the diagnostic value of the molecular sputum analysis was compared with sputum cytology.
Methods
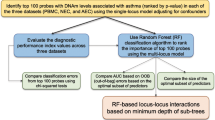
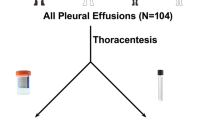
Subjects were included between June 2009 and February 2013 by pulmonologists in the regions of Amsterdam and Nieuwegein, the Netherlands (Figure 1). Cases were patients diagnosed with lung cancer. Their sputum was collected before lung cancer treatment, or when patients showed lung cancer progression while on treatment. Staging was performed according to the 7th edition of UICC TNM system (Sobin and Gospodarowicz, 2009). Controls were cancer-free subjects, mainly diagnosed with chronic obstructive disease (COPD), classified according to the GOLD criteria (Gold, 2009; Table 1). Patients who were cancer-free for a period of at least 3 years after curative treatment for lung cancer were also considered as controls. From the initially included controls (without symptoms at time of sputum collection), six patients developed lung cancer within a period of 6 months and were placed in the lung cancer group at time of analyses. Controls who developed lung cancer >6 months after sputum collection (n=14) were excluded from the main analyses and analysed separately.
The study was approved by the institutional review boards of the participating hospitals. All subjects provided signed informed consent. Oral and written information was provided to all subjects. Sociodemographic details and smoking habits were assessed by a questionnaire, completed by the subjects (Wood et al, 2005; Field et al, 2009). Clinical data were retrieved from medical records, blinded to outcome of methylation analysis.
Collection, recoding and processing of sputum with dithiothreitol, DNA isolation and hypermethylation analysis were performed as described before (Hubers et al, 2012). Based upon previous research of our group, DNA hypermethylation of the promoter regions of the following biomarkers were tested using multiplex quantitative methylation-specific PCRs: RASSF1A, CYGB, APC (Shivapurkar et al, 2007; Hubers et al, 2012) and recently discovered PRDM14, FAM19A4 and PHACTR3 (Snellenberg et al, 2012; Steenbergen et al, 2013). 3OST2 was tested in a singleplex quantitative methylation-specific PCR assay (Shivapurkar et al, 2007). Samples were tested in a blinded manner.
Cytological analysis
Following dithiothreitol processing, 0.2 ml of each sample was used for cytological analysis. Single layer slides were prepared using the Hettich Cyto-System. Cytological analysis was blinded for molecular analysis results and case–control status. Sputum cytology was scored for the following parameters: cell abundance; amount of neutrophilic granulocytes; cellular debris; squamous and/or cylinder cells; squamous metaplasia; and (suspicious) cancer cells. Sputum samples were considered representative for the respiratory tract if alveolar macrophages or respiratory epithelial cells were present. Cytology was defined ‘positive’ when cancer cells or cells suspicious for cancer (atypia) were identified.
Data and statistical analysis
A sputum bank was composed from the prospectively collected sputum samples. Only the first sputum canister (days 1–3) of subjects on which all biomarkers and cytology were assessed was included for analyses. An independent learning and validation set were randomly assigned from the sputum bank using a 1:2 ratio of both cases and controls, respectively (Figure 1).
To evaluate the diagnostic value of methylation for lung cancer in sputum, three different approaches were used, as described previously (Hubers et al, 2014). Receiver operating characteristic curves were composed with the ratio values of each marker. The second statistical model is based on a recent review (Hubers et al, 2013) that assesses the ‘diagnostic’ value of a biomarker (that is, minimal number of false-positive test results). In this review, we developed a rationale to determine the true diagnostic capacity of the methylation markers, substantiating that undiagnosed lung cancer is present in maximally 4% of the control population (based on combination of prevalence, risk and time). This resulted in a threshold setting for all markers at a fixed specificity per marker of 96%. Positive and negative predictive values (PPV and NPV, respectively) and diagnostic odds ratios (DORs) of markers were calculated for both the first and second approach with 95% confidence intervals (95% CI). In addition, multivariate logistic regression (markers as categorical variable) with a forward selection procedure was performed with the biomarkers in the learning set, leading to a combination rule with the highest sensitivity. This combination of markers with same thresholds was subsequently tested in the validation set. Biomarkers with a P-value ≤0.05 entered the logistic regression model. Results between the two sets were compared with the χ2 test or Fisher’s exact test. Chi-square tests were used to examine differences in DNA hypermethylation frequency between COPD patients without lung cancer and lung cancer patients without COPD. Moreover, to investigate whether COPD could be a possible confounder in the association between methylation and lung cancer, stratified DNA hypermethylation analysis and COPD-corrected logistic regression analysis were performed.
Furthermore, the complementary effect of DNA hypermethylation to cytology for lung cancer diagnosis was evaluated for the whole set with the McNemar test. To assess the additive value of sampling sputum during a prolonged time of 4–9 days, cumulative hypermethylation analysis was performed as described before using the cutoff obtained by Youden’s J index (Hubers et al, 2012). To examine the learning effect in time over canisters I to III, generalised estimating equations were used for each biomarker. Repeated measures for each subject were defined as the outcome of biomarker (positive or negative; using the cutoff obtained via the first approach) of one to three different canisters. An exchangeable structure was chosen for the correlation matrix, the logit-function was used as link function between the true status (case or control) of the subject and the outcome of biomarker, the number of the canister and their two-way interaction.
All statistical tests were two-sided with a significance level at 0.05 (P≤0.05). SPSS version 20.0 was used (IBM Corp., Armonk, NY, USA).
Results
Characteristics of subjects
Figure 1 shows the enrolment and follow-up of study subjects. Of 472 subjects, information on DNA hypermethylation analysis and cytology was available from the first sputum canister. Samples were randomised in a learning set (n=73 cases, n=86 controls) and validation set (n=159 cases, n=154 controls). Median duration of follow-up was 23 months in controls (range 0–43) and 8 months in cases (range 0–43).
Sociodemographic and clinical characteristics of cases, with sputum collection <6 months before diagnosis and controls in learning and validation sets are described in Table 1. In the validation set, mean age of controls was higher than of cases, and controls had smoked less pack years (P=0.011 and P=0.014, respectively). COPD was more prevalent in controls in both learning and validation sets (P=0.009 and P<0.001, respectively). Similar distributions were observed between the sets for the other variables.
Fifty per cent of lung cancer patients were diagnosed with stage IIIB and IV lung cancer. Adenocarcinoma (40%) and squamous cell carcinoma (35%) were the most prevalent histological types.
Cytology
Cytological analysis was performed for all subjects of learning and validation set combined, showing a positive result in 13.8% (95% CI: 9.6%–18.9%) of lung cancer cases with a specificity of 99.6% (95% CI: 97.7%–99.99%). Sensitivity marginally improved when cases had collected sputum during 9 days; 10 of 169 lung cancer patients, who had a negative cytology result in canister I, were detected in either canister II or III (5.9%; 95% CI: 2.9%–10.6%).
DNA hypermethylation analysis
Approach (i): discrimination capability of biomarkers between lung cancer patients and controls
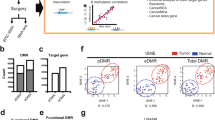
DNA hypermethylation analysis of RASSF1A, APC, CYGB, 3OST2, PHACTR3, FAM19A4 and PRDM14 was performed for all samples in learning and validation sets. Receiver operating characteristic curves for each marker are shown in Figure 2, for learning and validation set, respectively. Cutoff values were calculated based on Youden’s J index. Univariate analyses with 95% CIs of all biomarkers in both learning and validation sets are shown in Table 2. Regarding high specificity, RASSF1A showed the best diagnostic performance in both learning and validation sets (sensitivity and specificity were 42.5% and 96.5%, 36.5% and 88.3%, respectively). PPV was 91.2% (95% CI: 76.3%–98.1%; Supplementary Table 1a), NPV was 66.4% (95% CI: 57.4%–74.6%) and DOR was 20.4 (95% CI: 5.9–70.7). The combination rule of biomarkers RASSF1A, 3OST2 and PRDM14 was selected by multivariate logistic regression from the learning set for independent evaluation in the validation set. Both 3OST2 and PRDM14 showed individual high AUC scores (Table 2) with comparable results in the validation set. Positive DNA hypermethylation in one or more of these three markers demonstrated a sensitivity for lung cancer diagnosis of 82.2% (95% CI: 71.5%–90.2%) with a specificity of 66.3% (95% CI: 55.3%–76.1%) in the learning set. Similar results were observed for this panel in the validation set: sensitivity of 79.2% (95% CI: 72.1%–85.3%; P=0.60) and specificity of 64.3% (95% CI: 56.2%–71.8%; P=0.76). Diagnostic efficiency of the biomarker panel remained more or less similar with the addition of cytology (sensitivity of 83.6% in learning set). Molecular sputum analysis is superior over sputum cytology (P<0.001).
Receiver operator characteristic (ROC) curves were composed with the ratio values of markers RASSF1A, APC, CYGB, FAM19A4, 3OST2, PHACTR3 and PRDM14 for (A) learning set and (B) validation set. The true positive rate (sensitivity) is plotted against the false-positive rate (1-specificity) for the different possible cutoff values.
No relation was observed between early (stage I–II) and advanced (stage III–IV) lung cancer and DNA hypermethylation (P-values >0.10). Regarding histology of the tumours (adenocarcinoma versus squamous cell carcinoma), PHACTR3 showed to be more hypermethylated in adenocarcinomas when compared with squamous cell carcinomas (P=0.001; Table 3). Although not significant, RASSF1A hypermethylation was more observed in squamous cell carcinomas.
In the group of never-smokers (22 cases and 17 controls), hypermethylation of most biomarkers was comparable for sensitivity and specificity in smokers with >15 pack years (P>0.04; data not shown). RASSF1A and 3OST2 demonstrated high specificity (95% and 91%, respectively) with a sensitivity of 47% and 53%, respectively. When smokers <15 pack years were combined with never smokers similar results were obtained.
For clinical parameters, such as age and smoking status, no association was observed with DNA hypermethylation. In comparing COPD patients without lung cancer with lung cancer patients without COPD, all tested methylation markers have a (significantly) higher fraction of positive cases in lung cancer (Supplementary Table 2). To examine whether COPD is a confounding factor, cases of learning and validation sets were combined and logistic regression analysis revealed after correcting for COPD that the regression coefficient changed less than 10% for all tested methylation markers (for example from b=1.798 to 1.793 for RASSF1A), excluding COPD as confounding factor. Furthermore, analyses of the association between methylation markers and lung cancer stratified by COPD status did not reveal relevant differences (Supplementary Table 3) neither did analyses of the association between methylation markers and COPD status stratified by group (lung cancer or control; Supplementary Table 4).
Fourteen subjects presented with lung cancer more than 6 months after sputum collection. Of these, four were positive for RASSF1A hypermethylation.
Approach (ii): diagnostic value of biomarkers
The diagnostic value of the methylation markers was examined starting with a fixed 96% specificity for each marker in the learning set (Table 2). Multivariate logistic regression analysis was performed and resulted in the combination of RASSF1A, 3OST2 and PHACTR3, yielding a sensitivity of 67.1% (95% CI: 55.1%–77.7%) and specificity of 89.5% (95% CI: 90.1%–99.3%) in the learning set, versus 64.8% (95% CI: 56.8%–72.2%) and 80.5% (95% CI: 73.4%–86.5%) in the validation set, respectively. No differences were observed between both sets for sensitivity and specificity (P=0.73 and 0.07, respectively). PPV, NPV and DOR for all methylation markers are shown in Supplementary Table 1b.
Approach (iii): risk classification model
The risk classification model was composed with samples of the learning set and subsequently evaluated on the validation set. Logistic regression analysis first included RASSF1A for identification of high-risk individuals and next 3OST2 and PRDM14 for lower-risk categories in the model (Table 4).
In the learning set, RASSF1A classified 39.7% of lung cancer patients in the high-risk group (≥60% chance on lung cancer) with few false-positive controls (2.3%). The risk factors 3OST2 and PRDM14 assigned half of the remaining lung cancer cases in the moderate lung cancer risk groups and 30% in the lowest risk group, whereas the majority of controls (81.4%) were allocated to the lowest risk group. Consistent results were demonstrated in the validation set with slightly more lung cancer patients in the moderate risk groups (18.2% and 21.4%, respectively), and with a marginally lower specificity for RASSF1A.
Prolonged sputum sampling
From 195 cases and 228 controls, a complete set of the three canisters (i.e., I, II and III) with sufficient DNA for hypermethylation analysis was available (Figure 1). Using cutoff values based on Youden’s J index, McNemar tests and Cochran Q tests did not show statistically significant differences in frequency of hypermethylation among the three canisters, except for CYGB, which demonstrated significantly more hypermethylation in the third canister, when compared with canisters I and II (50%, 51% and 58%, respectively; P=0.03). Therefore, sputum quality is comparable among the canisters.
The number of lung cancer patients who tested negative in canister I and with positive hypermethylation in canisters II and III is shown in Table 5. Individual marker analysis showed that proportion of additional positive cases is larger for the risk markers (mean 41%), compared with diagnostic marker RASSF1A (11%). Seventeen of thirty-seven cases were detected in addition when either canister II or III was tested for hypermethylation of biomarker panel RASSF1A, 3OST2 and PRDM14 (45.9%; 95% CI: 29.5%–63.1%). Generalised estimating equations did not show an interaction between canister number and the outcome of the biomarker, indicating that no learning effect for sputum sampling occurred over time.
Discussion
This study reports on a training-validation approach of DNA hypermethylation analysis of biomarkers in sputum for the diagnosis of lung cancer. All tested biomarkers were able to discriminate between lung cancer patients and controls. RASSF1A showed best performance with high positive predictive value and high DORs; not only in two different and independent sets, but also using three different statistical approaches. This confirms the value of RASSF1A hypermethylation as a diagnostic marker (according to previous definition, i.e., false-positive test results in <4% of controls; Hubers et al, 2013). In addition, novel biomarkers were examined, which have not been tested in sputum before (PRDM14, PHACTR3, FAM19A4; Steenbergen et al, 2013). We developed and evaluated two panels, each consisting of three biomarkers. Both panels had in common the inclusion of biomarkers RASSF1A and 3OST2, and dependent of the panel application (high sensitivity or high specificity), the third biomarker of the panel was either PRDM14 or PHACTR3, respectively. In both learning and validation sets, the first combination rule showed similar sensitivity of 80–82% and specificity of 65–66% for lung cancer diagnosis. In diagnostic setting, that is, with focus on high specificity, the second panel revealed 67% sensitivity and 90% specificity. Last, a risk model for lung cancer prediction was composed, incorporating RASSF1A, 3OST2 and PRDM14. Accuracy should be improved, but this model shows potential as clinical tool for application in a population at risk for lung cancer to categorise subjects in different risk groups.
Twenty-one controls tested false positive for RASSF1A hypermethylation in their first sputum canister. Of note, one of these controls died from liver metastases of an unknown primary source three years after sputum collection. Another control presented with weight loss and fatigue, showing a non-progressing infiltrate in the right upper lobe. No further diagnostic work up was performed. In addition, five patients presented with lung metastases from a primary tumour different from lung cancer. These were separately analysed for RASSF1A hypermethylation. Two showed RASSF1A hypermethylation in their sputum: one had a primary breast tumour, the other metastatic colon carcinoma. For both cancer types, RASSF1A hypermethylation has been reported in the literature (Pfeifer and Dammann, 2005). Thus, RASSF1A hypermethylation in subjects without a primary lung cancer should not be interpreted as a false-positive test at first glance, as it is not excluded that the positivity may be due to the presence of not established lung cancer (Hubers et al, 2013) or to lung metastases originating from another primary tumour.
Methylation frequencies of 3OST2 in our data (49.7–56.2%) are comparable as reported previously (Leng et al, 2012), although numbers on specificity are contrasting. Leng et al (2012) observed a lower specificity, whereas Shivapurkar et al (2007) found no false-positive controls. COPD increases the risk of lung cancer among smokers and COPD patients present with similar symptoms (Adcock et al, 2011). In our study, COPD was no confounding factor.
For diagnostic use of hypermethylation markers, for example, to confirm malignancy after imaging of a solid lesion or ground glass opacity, one should strive after high specificity of the markers. From this point of view, cutoff values for all methylation markers were set at a specificity of 96%, based on a rationale as previously published (Hubers et al, 2013). Logistic regression yielded a novel combination of RASSF1A, 3OST2 and PHACTR3 with a sensitivity of 67% and specificity of 90%. In this scenario, sensitivity is lower. As a consequence, the high specificity leads to reduction in false-positive results.
A risk classification model was constructed, based on the approach as introduced in the previous research (Belinsky et al, 2002; Zöchbauer-Müller et al, 2003; Shivapurkar et al, 2007; Baryshnikova et al, 2008; Leng et al, 2008; Van der Drift et al, 2008; Stidley et al, 2010; Hubers et al, 2014). Instead of the conventional way of interpreting dichotomised test results of biomarkers as the ‘absolute’ presence or absence of hypermethylation, this novel approach assumes that ratio values of biomarkers can be divided into categories with corresponding risk probabilities. This could be a practical tool for the clinician, both for use during the diagnostic process and for screening of lung cancer. In the current study, markers were examined in comparison with RASSF1A, APC and CYGB (which were included in the proposed risk model before) and we observed that RASSF1A, 3OST2 and PRDM14 were more accurate in predicting chance of lung cancer. The risk model showed consistent results between learning and validation sets.
Irrespective of the statistical model used, about 40% of the lung cancer patients will not be diagnosed with the current hypermethylation markers. This is explained by several reasons. First, none of the genes is hypermethylated in 100% of lung cancer, emphasising the need for complementary biomarkers. Second, previous research showed a mean concordance of 78% between matched primary hypermethylated lung cancer tissue and sputum (Hubers et al, 2013), which raises a question about the representativity: not all cases with hypermethylation in the primary tumour show detectable DNA methylation in their sputum samples.
Interestingly, we observed a high specificity of RASSF1A and 3OST2 in never-smokers as well as in smokers with limited number of pack years (<15). This may suggest to examine these markers in other subjects than those meeting the current inclusion criteria for lung cancer screening (i.e., heavy smokers).
When compared with sputum cytology, DNA hypermethylation analysis showed to be superior to sputum cytology in lung cancer detection. At the 99% specificity level (comparable to sputum cytology), sensitivity of hypermethylation RASSF1A is still 16%.
Spontaneous sputum is usually easily obtained from smokers. Although in former and non-smokers collection of sputum seems more difficult, careful instruction may still lead to representative sputum samples. Alternatively, induced sputum may be an option (Chanez et al, 2002; Anjuman et al, 2013). In this study, we only collected spontaneous sputum, because this is more easily accomplished as collection can be performed at home. Detailed instructions in the information brochure increases patient compliance. Induced sputum requires additional logistics, such as an extra visit to the hospital and efforts of the patients.
A limitation of the study is that the impact of sputum testing on clinical decision making, clinical outcomes of patients to whom testing is applied and costs are not assessed.
Future research is needed to optimise the marker panels. Given the heterogeneous nature of lung cancer and the numerous cellular pathways involved (Hansen et al, 2011), it is likely that a panel of biomarkers will yield a higher sensitivity compared with a single marker. Promising additional (diagnostic) markers such as microRNAs and tumour-specific proteins in sputum may further improve the efficiency for lung cancer diagnosis (Sun et al, 2009; Xing et al, 2010; Yu et al, 2010).
Overall, test characteristics of sputum methylation have been reproduced. RASSF1A hypermethylation in sputum is validated as diagnostic marker for lung cancer. The panel of RASSF1A, 3OST2 and PHACTR3 hypermethylation revealed a 67% sensitivity and high specificity (90%) in a diagnostic setting.
Change history
17 March 2015
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Adcock IM, Caramori G, Barnes PJ (2011) Chronic obstructive pulmonary disease and lung cancer: new molecular insights. Respiration 81: 265–284.
Anjuman N, Li N, Guarnera M, Stass SA, Jiang F (2013) Evaluation of lung flute in sputum samples for molecular analysis of lung cancer. Clin Transl Med 2: 15.
Baryshnikova E, Destro A, Infante MV, Cavuto S, Cariboni U, Alloisio M, Ceresoli GL, Lutman R, Brambilla G, Chiesa G, Ravasi G, Roncalli M (2008) Molecular alterations in spontaneous sputum of cancer-free heavy smokers: results from a large screening program. Clin Cancer Res 14: 1913–1919.
Belinsky SA, Klinge DM, Dekker JD, Smith MW, Bocklage TJ, Gilliland FD, Crowell RE, Karp DD, Stidley CA, Picchi MA (2005) Gene promoter methylation in plasma and sputum increases with lung cancer risk. Clin Cancer Res 11: 6505–6511.
Belinsky SA, Palmisano WA, Gilliland FD, Crooks LA, Divine KK, Winters SA, Grimes MJ, Harms HJ, Tellez CS, Smith TM, Moots PP, Lechner JF, Stidley CA, Crowell RE (2002) Aberrant promoter methylation in bronchial epithelium and sputum from current and former smokers. Cancer Res 62: 2370–2377.
Chanez P, Holz O, Ind PW, Djukanovic R, Maestrelli P, Sterk PJ (2002) Sputum induction. Eur Respir J 20: 3S–8S.
Cirincione R, Lintas C, Conte D, Mariani L, Roz L, Vignola AM, Pastorino U, Sozzi G (2006) Methylation profile in tumor and sputum samples of lung cancer patients detected by spiral computed tomography: a nested case-control study. Int J Cancer 118: 1248–1253.
Van der Drift MA, Prinsen CFM, Hol BEA, Bolijn AS, Jeunink MAF, Dekhuijzen PNR, Thunnissen FBJM (2008) Can free DNA be detected in sputum of lung cancer patients? Lung Cancer 61: 385–390.
Esteller M (2011) Epigenetic changes in cancer. F1000 Biol Rep 3: 9.
Field JK, Liloglou T, Niaz A, Bryan J, Gosney JR, Giles T, Brambilla C, Brambilla E, Vesin A, Timsit J-F, Hainaut P, Martinet Y, Vignaud JM, Thunnissen FB, Prinsen C, Snijders PJ, Smit EF, Sozzi G, Roz L, Risch A, Becker HD, Elborn JS, Magee ND, Montuenga LM, Pajares MJ, Lozano MD, O’Byrne KJ, Harrison DJ, Niklinski J, Cassidy A (2009) EUELC project: a multi-centre, multipurpose study to investigate early stage NSCLC, and to establish a biobank for ongoing collaboration. Eur Respir J 34: 1477–1486.
Gold PM (2009) The 2007 GOLD Guidelines: a comprehensive care framework. Respir Care 54: 1040–1049.
Hansen KD, Timp W, Bravo HC, Sabunciyan S, Langmead B, McDonald OG, Wen B, Wu H, Liu Y, Diep D, Briem E, Zhang K, Irizarry RA, Feinberg AP (2011) Increased methylation variation in epigenetic domains across cancer types. Nat Genet 43: 768–775.
Honorio S, Agathanggelou A, Schuermann M, Pankow W, Viacava P, Maher ER, Latif F (2003) Detection of RASSF1A aberrant promoter hypermethylation in sputum from chronic smokers and ductal carcinoma in situ from breast cancer patients. Oncogene 22: 147–150.
Hubers AJ, van der Drift MA, Prinsen CFM, Witte BI, Wang Y, Shivapurkar N, Stastny V, Bolijn AS, Hol BEA, Feng Z, Dekhuijzen PNR, Gazdar AF, Thunnissen E (2014) Methylation analysis in spontaneous sputum for lung cancer diagnosis. Lung Cancer 84: 127–133.
Hubers AJ, Heideman DAM, Herder GJM, Burgers SA, Sterk PJ, Kunst PW, Smit HJ, Postmus PE, Witte BI, Duin S, Snijders PJF, Smit EF, Thunnissen E (2012) Prolonged sampling of spontaneous sputum improves sensitivity of hypermethylation analysis for lung cancer. J Clin Pathol 65: 541–545.
Hubers AJ, Prinsen CFM, Sozzi G, Witte BI, Thunnissen E (2013) Molecular sputum analysis for the diagnosis of lung cancer. Br J Cancer 109: 530–537.
Leng S, Do K, Yingling CM, Picchi MA, Wolf HJ, Kennedy TC, Feser WJ, Baron AE, Franklin WA, Brock MV, Herman JG, Baylin SB, Byers T, Stidley CA, Belinsky SA (2012) Defining a gene promoter methylation signature in sputum for lung cancer risk assessment. Clin Cancer Res 18: 3387–3395.
Leng S, Stidley CA, Willink R, Bernauer A, Do K, Picchi MA, Sheng X, Frasco MA, Van Den Berg D, Gilliland FD, Zima C, Crowell RE, Belinsky SA (2008) Double-strand break damage and associated DNA repair genes predispose smokers to gene methylation. Cancer Res 68: 3049–3056.
Patz EF, Rossi S, Harpole DH, Herndon JE, Goodman PC (2000) Correlation of tumor size and survival in patients with stage IA non-small cell lung cancer. Chest 117: 1568–1571.
Pfeifer GP, Dammann R (2005) Methylation of the tumor suppressor gene RASSF1A in human tumors. Biochem 70: 576–583.
Rivera MP, Mehta AC, Wahidi MM (2013) Establishing the diagnosis of lung cancer: Diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 143: e142S–e165S.
Shivapurkar N, Stastny V, Suzuki M, Wistuba II, Li L, Zheng Y, Feng Z, Hol B, Prinsen C, Thunnissen FB, Gazdar AF (2007) Application of a methylation gene panel by quantitative PCR for lung cancers. Cancer Lett 247: 56–71.
Siegel R, Naishadham D, Jemal A (2012) Cancer statistics, 2012. CA Cancer J Clin 62: 10–29.
Snellenberg S, De Strooper LMA, Hesselink AT, CJLM Meijer, Snijders PJF, Heideman DAM, Steenbergen RDM (2012) Development of a multiplex methylation-specific PCR as candidate triage test for women with an HPV-positive cervical scrape. BMC Cancer 12: 551.
Sobin LH, Gospodarowicz MK WC (2009) International Union Against Cancer (UICC) TNM classification of malignant Tumours, 7th edn Wiley-Blackwell: Oxford.
Steenbergen RD, Ongenaert M, Snellenberg S, Trooskens G, van der Meide WF, Pandey D, Bloushtain-Qimron N, Polyak K, Meijer CJ, Snijders PJ, Van Criekinge W (2013) Methylation-specific digital karyotyping of HPV16E6E7-expressing human keratinocytes identifies novel methylation events in cervical carcinogenesis. J Pathol 231: 53–62.
Stidley CA, Picchi MA, Leng S, Willink R, Crowell RE, Flores KG, Kang H, Byers T, Gilliland FD, Belinsky SA (2010) Multivitamins, folate, and green vegetables protect against gene promoter methylation in the aerodigestive tract of smokers. Cancer Res 70: 568–574.
Sun B, Wang H, Wang X, Huang H, Ding W, Jing R, Shi G, Zhu L (2009) A proliferation-inducing ligand: a new biomarker for non-small cell lung cancer. Exp Lung Res 35: 486–500.
Wood DM, Mould MG, Ong SBY, Baker EH (2005) “Pack year” smoking histories: what about patients who use loose tobacco? Tob Control 14: 141–142.
Xing L, Todd NW, Yu L, Fang H, Jiang F (2010) Early detection of squamous cell lung cancer in sputum by a panel of microRNA markers. Mod Pathol 23: 1157–1164.
Yu L, Todd NW, Xing L, Xie Y, Zhang H, Liu Z, Fang H, Zhang J, Katz RL, Jiang F (2010) Early detection of lung adenocarcinoma in sputum by a panel of microRNA markers. Int J Cancer 127: 2870–2878.
Zöchbauer-Müller S, Lam S, Toyooka S, Virmani AK, Toyooka KO, Seidl S, Minna JD, Gazdar AF (2003) Aberrant methylation of multiple genes in the upper aerodigestive tract epithelium of heavy smokers. Int J Cancer 107: 612–616.
Acknowledgements
We thank all subjects who participated in this study. Furthermore, special thanks to Marleen Peterse-Van ‘t Schip, Mirjam Nauta, Stephan Jans, Remco Boksem, Gerrit-Jan Ilbrink, Albert Olijve and Geertje Houwaard for excellent assistance and efforts in patient inclusion. The help of members of pathology administration and cytology units (VUmc) for screening of slides is greatly appreciated.
This study receives funding from Dutch Cancer Society (grant VU2008-4220).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Pieter E. Postmus has board membership at Boehringer Ingelheim (advisory board on nintedanib), payment for lectures BMS (chair and lecture at symposium at WCLC 2013). The remaining authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Hubers, A., Heideman, D., Burgers, S. et al. DNA hypermethylation analysis in sputum for the diagnosis of lung cancer: training validation set approach. Br J Cancer 112, 1105–1113 (2015). https://doi.org/10.1038/bjc.2014.636
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.636
Keywords
This article is cited by
-
Liquid biopsies based on DNA methylation as biomarkers for the detection and prognosis of lung cancer
Clinical Epigenetics (2022)
-
Genome-wide analysis of cell-Free DNA methylation profiling with MeDIP-seq identified potential biomarkers for colorectal cancer
World Journal of Surgical Oncology (2022)
-
Cytoglobin attenuates pancreatic cancer growth via scavenging reactive oxygen species
Oncogenesis (2022)
-
MPDZ as a novel epigenetic silenced tumor suppressor inhibits growth and progression of lung cancer through the Hippo-YAP pathway
Oncogene (2021)
-
High-Yield Methylation Markers for Stool-Based Detection of Colorectal Cancer
Digestive Diseases and Sciences (2020)