Abstract
Background:
Growing evidence suggests that miR-29a has an important role in regulating tumourigenesis and development of various types of cancer. However, the role and the underlying mechanism of miR-29a in colorectal cancer (CRC) remain largely unknown.
Methods:
MiR-29a targeted gene was identified by the luciferase assay and western blot. MiR-29a function was analysed by invasion assays and the orthotopic transplantation mouse model. The miR-29a pathway was assayed by real-time PCR, western blot and chip analysis.
Results:
KLF4 was identified as a direct target gene of miR-29a. MiR-29a promoted CRC cell invasion, which was blocked by re-expression of KLF4. In addition, MMP2 was identified as a novel direct target of KLF4. Both miR-29a overexpression and KLF4 knockdown promoted MMP2 expression but inhibited E-cadherin expression. Furthermore, clinical data indicated that both miR-29a high expression and KLF4 mRNA low expression were associated with metastasis and poor prognosis in CRC patients, and KLF4 protein expression was inversely correlated with MMP2 but positively correlated with E-cad protein expression.
Conclusion:
Increased expression of miR-29a promoted CRC metastasis by regulating MMP2/E-cad through direct targeting KLF4, which highlights the potential of the miR-29a inhibitor as a novel agent against CRC metastasis.
Similar content being viewed by others
Main
Colorectal cancer (CRC) is one of the leading causes of cancer-related deaths worldwide (Jemal et al, 2009). Although radical resection of non-metastatic CRC obviously increases the survival rate, metastasis and relapse remain the main cause of death in CRC patients (Mina and Sledge, 2011). It is essential to unveil molecular pathways of CRC metastasis before we could develop new molecular markers for the diagnosis and treatment of CRC.
Tumour metastasis includes a series of biological processes including primary tumour angiogenesis, tumour cell invasion, lymphatic/vascular intravasation, distant organ extravasation and colonisation and growth of metastatic tumour cells (Hanahan and Weinberg, 2011). Although many molecules related to tumour metastasis have been identified, they cannot fully explain the phenomenon of cancer metastasis. It was reported that microRNAs (miRNAs), functioning as endogenous regulatory RNA molecules, modulated many physiological and pathological processes through downregulating target genes (Bartel, 2009). Our previous data demonstrated that miR-29a was frequently upregulated in CRC tissues (Fu et al, 2012), implying that miR-29a is a potential cancer-promoting miRNA in CRC. The same observation was also confirmed by the research conducted by The Cancer Genome Atlas Network (2012). However, the specific role of miR-29a in CRC and its possible mechanism of the action remain elusive.
KLF4, together with Oct3/4, Sox2 and c-Myc, was identified as the four defined critical factors of inducing differentiated cells into pluripotent stem cells (Takahashi and Yamanaka, 2006). Previous studies showed that KLF4 had a tumour suppressive function in CRC (Zhao et al, 2004). Heterozygous deletion of Klf4 gene in an Apc Min mice model increased the intestinal tumour burden (Ghaleb et al, 2007). In addition, KLF4 overexpression inhibited colony formation, migration and invasion of colon cancer cells in vitro and tumourigenicity in vivo (Dang et al, 2003). Downregulation of KLF4 in colon adenomas contributed to cellular hyperproliferation and malignant transformation (Choi et al, 2006). However, the underlying molecular mechanism for KLF4 low expression in CRC needs to be further explored.
In the present study, miR-29a was identified as a novel metastasis-promoting factor through targeting KLF4, and miR-29a high expression increased CRC cell invasion in vitro and liver metastasis in vivo. Furthermore, mechanical research revealed that KLF4 directly bound to the promoter of MMP2 and E-cad to regulate their expression in CRC cells. In summary, a novel molecular link from high expression of miR-29a to cancer metastasis through upregulation of MMP2 and downregulation of E-cad via targeting KLF4 in CRC tumourigenesis was established.
Material and Methods
Human tissues and cell lines
Eighty-five human primary CRC tissue specimens were collected by surgery at Xinhua Hospital of Shanghai Jiaotong University (Shanghai, China) between Jan 2009 and Jun 2009, immediately snap-frozen in liquid nitrogen and stored at −80 °C for analysis. Additional 82 pairs of human primary CRC tissues and adjacent non-cancerous tissues (NCTs) collected at the same time were constructed for tissue microarray. All human materials were obtained with signed informed consent, and the study was approved by the Clinical Research Ethics Committee of Xinhua Hospital of Shanghai Jiaotong University.
The human CRC cell lines HCT-116 and LoVo were purchased from the American Type Culture Collection (ATCC, USA). LoVo was maintained in F12-K, and HCT-116 was maintained in McCoy’s 5a. Media were supplemented with 10% fetal bovine serum (FBS) at 37 °C and 5% CO2.
RNA extraction and quantitative real-time PCR
Total RNA was extracted using RNAiso Plus (TaKaRa, Tokyo, Japan). Complementary DNA was synthesised with PrimeScript RT–PCR Kit (TaKaRa). Quantitative RT–PCR (qRT–PCR) was performed using SYBR Premix Ex Taq (TaKaRa) on ABI 7500 PCR System (Applied Biosystems, Foster City, CA, USA). Specific Bulge-LoopTM miRNA Primers were synthesised by RiboBio (Guangzhou, China). GAPDH and U6 small nuclear RNA were used as internal control for detection of mRNA and miRNA, respectively. The primer sequence for mRNA qRT–PCR was displayed in Supplementary Table S1. For the study of correlations between miR-29a and KLF4 mRNA expression, and correlations between miR-29a/KLF4 mRNA expression and overall survival (OS), miR-29a expression and KLF4 mRNA expression were classified to low and high expression according to their respective median expression.
Oligonucleotide transfection
MicroRNA-29a mimics and anti-miR-29a were synthesised by RiboBio. KLF4 siRNA (KLF4 siRNA-1 5′-CCAGCCAGAAAGCACUACAAU-3′, KLF4 siRNA-2 5′-UGACCAGGCACUACCGUAA-3′) was synthesised by Genepharma (Shanghai, China). RNA oligonucleotides were transfected by lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA).
Vector construction and lentivirus production
The human mature miR-29a sequence 5′-UAGCACCAUCUGAAAUCGGUUA-3′ was synthesised and then cloned into the lentivirus expression vector pGLV3 to generate pGLV3-miR-29a. To generate virus particles, package, envelop and expressing constructs were co-transfected into HEK293T cells using lipofectamine 2000. For infection, the virus was supplemented with 8 μg ml−1 of polybrene (Sigma, St Louis, MO, USA).
The 3′UTR of KLF4 was amplified from human genomic DNA and cloned into the pGL3-control vector (Promega, Madison, WI, USA). Site-directed mutagenesis of the miR-29a target-site in the KLF4 3′UTR was carried out using site-directed mutagenesis kit (TaKaRa, Japan). The primer sequence for KLF4 3′UTR Mut was F 5′-CACAGTCTAGACCCAGCCAGAAAGCACTACAATC-3′ and R 5′-CACAGGAGCTCACTCACCATCGTGGTAGATTTAGGC-3′.
The MMP2 proximal promoter (−1256/+299) was amplified from human genomic DNA by PCR. Both PCR products and pGL3-cyclin D1-luciferase reporter plasmid (addgene 32726) were digested with KpnI and HindIII and ligated together to form the (−1256/+299) MMP2-Luc reporter plasmid. The primers for constructing vectors are displayed in Supplementary Table S1.
Luciferase assay
HEK293T cells (1 × 105 per 0.05 ml) were cultured in 24-well plates. For 3′UTR of KLF4 report system, HEK293T cells were co-transfected with 200 ng WT or Mutant reporter plasmid, 25 nmol miRNAor anti-miRNA and 20 ng Renil using lipofectamine 2000 (Invitrogen). After 24-h transfection, firefly and Renilla luciferase activities were measured using the dual-luciferase reporter assay system (Promega). For MMP2 promoter report system, HEK293T cells were co-transfected with 200 ng WT or Mutant reporter plasmid, 25 nmol siRNA and 20 ng Renil for 48 h.
Western blotting
Cells were lysed with 1 × SDS–PAGE loading buffer. An equal amount of protein was separated on 10% SDS–PAGE gel and then transferred to the nitrocellulose membrane. After blocking with 3% BSA, the membrane was probed with the following primary antibodies: anti-KLF4 (Abcam, Cambridge, UK; 1 : 1000), anti-MMP2 (Bioworld, St Louis, MN, USA; 1 : 1000), anti-E-cad (BD, Bedford, MA, USA; 1 : 1000) and anti-GAPDH (Santa, Dallas, TX, USA; 1 : 1000). Glyceraldehyde 3-phosphate dehydrogenase was used as loading control. The proteins were visualised using ECL reagents (Pierce, Rockford, IL, USA).
Invasion assay
Transwell champers (24-well insert; Corning, Horseheads, NY, USA) were used to analyse the ability of cell invasion. The chambers were precoated with 0.05 ml matrigel (BD Biosciences, 50 mg ml−1 1 : 8) for 4 h. After 48-h transfection, cells were suspended in culture medium with 1% FBS and added to the upper chambers (HCT-116, 12 × 104cells per well and LoVo, 8 × 104cells per well). At the end point of incubation (HCT-116, 48 h and LoVo, 22 h), cells on the upper membrane surface were removed. The lower membrane surface was fixed by 4% formaldehyde, stained with Hoechst 33 342 and counted under a fluorescence microscope.
Cell proliferation assay
HCT-116 cells (3000 per well) were planted into 96-well culture plate. Sixteen hours later, miR-Ctrl or miR-29a oligonucleotides were transfected by lipofectamine 2000 (Invitrogen). Cell proliferation was measured at 24 h, 48 h and 96 h post-transfection, respectively, by using the Cell Counting Kit-8 (Dojindo, Japan) according to the manufacture’s instruction.
Cell cycle assay
HCT-116 cells were harvested by 0.25% trypsin without EDTA 48 h after transfection with miR-Ctrl or miR-29a. Then cells were washed with ice-cold PBS and fixed with 70% ethanol overnight at 4 °C. For flow cytometric analysis, the fixed cells were suspended in PBS, incubated with RNase at 37 °C for 1 h and subjected to PI staining (Sigma).
In vivo models
Six-week-old male Nu/Nu mice (Shanghai SLAC Laboratory Animal Co., Ltd., Shanghai, China) were housed under specific pathogen-free conditions. Control or miR-29a overexpressed LoVo cells (2 × 106) were resuspended in 0.05 ml PBS and injected into the wall of the cecum (five mice for each group) to established the orthotopic transplantation model. After 8 weeks, mice were killed, and livers were isolated and examined for the number of metastatic tumours and calculated for the expression of KLF4 and miR-29a.
Chromatin immunoprecipitation
The chromatin immunoprecipitation (ChIP) assay was performed using Magna ChIP A/G Chromatin Immunoprecipitation Kit (Millipore Cat no.: 17–10085; Temecula, CA, USA). Briefly, 1 × 107 HCT-116 cells were fixed with 1% paraformaldehyde and lysed by SDS lysis buffer. Then cell lysate was sonicated and immunoprecipitated with antibodies specific to KLF4 (R&D Cat no: AF3640; Minneapolis, MA, USA) or control goat IgG (Bioworld). After protein/DNA complexes were eluted, reverse cross-linked to free DNA and purified, the specific DNA fragments were quantitated by qRT–PCR and normalised to input from the same cells. The qRT–PCR primers for MMP2 are listed in Supplementary Table S1.
Immunohistochemical staining
Tissue microarray composed of 82 pairs of human primary CRC tissues and NCTs was immunohistochemically stained to determine the expression of KLF4, MMP2 and E-cad. The staining score was evaluated blindly by two independent investigators and recorded by the staining proportion (the percentage of cells stained) and the staining intensity, as suggested by Remmele and Stegner (Remmele and Stegner, 1987). The proportion score was scored as 0 (0%), 1 (>0 to ⩽25%), 2 (>25 to ⩽50%), 3 (>51 to ⩽75%) and 4 (>75%). The intensity score was record as 0 (negative result), 1 (weakly positive), 2 (moderately positive) and 3 (strongly positive). The final staining scores were calculated by proportion score × intensity score. For the classification of staining, a staining result ⩽4 indicates a low level of expression, whereas a staining score >4 represents a high level of expression.
Statistical analysis
Statistical analysis was performed using SPSS15.0. Values were expressed as mean±standard deviation (s.d.) for parametric data. Difference between groups was calculated using the Student’s t-test and non-parametric test (Mann–Whitney U-test). Overall survival curves were deprived from the Kaplan–Meier estimate, and the survival differences between different groups were compared by the log-rank test. The statistical significance of correlations between the expression of miR-29a and KLF4 mRNA, and correlations between KLF4 expression and MMP2 and E-cad expression were calculated by the Pearson χ2 test and the Spearman’s rank correlation. A P-value <0.05 was considered statistically significant.
Results
MiR-29a directly targets KLF4 in CRC cells
KLF4 is a well-established tumour suppressor gene in CRC and has been implicated in cell proliferation, migration and invasion. Using miRanda (John et al, 2004) and TargetScan (Grimson et al, 2007) algorithms, KLF4 was predicted as a potential direct target of miR-29a. To further confirm whether or not KLF4 was a direct target of miR-29a, the 3′UTR of KLF4 was cloned into a luciferase reporter vector and then the predicted binding site of miR-29a in the KLF4 3′UTR was mutated to evaluate the effect of miR-29a and anti-miR-29a using luciferase assay (Figure 1A). It was found that ectopic overexpression of miR-29a significantly inhibited the luciferase activity of pGL3-KLF4 3′UTR WT, whereas miR-29a silencing increased the luciferase activity (Figure 1B and C). In addition, mutation of the binding site of miR-29a in the 3′UTR of KLF4 abolished both the effect of miR-29a and anti-miR-29a (Figure 1B and C). In concordance with these results, endogenous KLF4 protein expression was decreased in miR-29a-transfected CRC cells, which could be reversed by anti-miR-29a treatment (Figure 1D and E). Therefore, these results suggest that miR-29a directly regulated KLF4 in CRC cells.
MicroRNA-29a directly targets KLF4 in CRC cells. (A) Putative miR-29a-binding sequence in KLF4 3′UTR. KLF4 3′UTR mutation was generated in the complementary site for the seed region of miR-29a as indicated. (B and C) Analysis of the luciferase activity of pGL3-KLF4 3′UTR WT and Mut plasmid in HEK293T cells by miR-29a or anti-miR-29a. (D) Western blot analysis of HCT-116 cells transfected with indicated miR-29a or anti-miR-29a. (E) Western blot analysis of LoVo cells transfected with indicated miR-29a or anti-miR-29a. All numerical data are presented as mean±s.d. *P<0.05, n=3.
MiR-29a promotes CRC cell invasion in vitro and metastasis in vivo through regulation of the target gene KLF4
A matrigel invasion assay was performed to investigate the potential role of miR-29a in CRC cell invasion. The transfection efficiency was confirmed by qRT–PCR 48 h after transfection (Supplementary Figure S1A and B). The invasive activity in miR-29a-transfected CRC cells increased significantly, whereas cell invasion in CRC cells treated with anti-miR-29a decrease markedly (Figure 2A and B). In addition, miR-29a was found to have no effect on cell proliferation and cell cycle regulation in the HCT-116 cells (Supplementary Figure S1C and D), suggesting that the effect of miR-29a on cell invasion was independent of cell proliferation and cell cycle. To further examine whether miR-29a-induced CRC cell invasion was dependent on KLF4, HCT-116 cells were transfected with miR-29a with or without KLF4 in the transwell invasion assay. It was found that the effect of miR-29a overexpression on cell invasion was blocked by KLF4 re-expression (Figure 2C). Our reciprocal experiment showed that anti-miR-29a inhibited the invasion of LoVo cells, and this effect was partially reversed by KLF4 silencing (Figure 2D). Finally, to investigate the role of miR-29a in CRC metastasis in vivo, LoVo transfectants stably expressing miR-29a using lentivirus infection were established and then injected into the cecum of nude mice. It was found that miR-29a overexpression significantly increased the number of metastatic nodules in the liver 8 weeks after implantation (Figure 2E and F), and meanwhile the metastatic nodules produced by miR-29a stably expressing LoVo cells still presented the lower KLF4 expression (Figure 2G) compared with the control. Collectively, our data indicate that miR-29a exerted a metastasis-promoting function in human CRC via targeting KLF4.
MicroRNA-29a promotes CRC cell invasion in vitro and metastasis in vivo through regulation of KLF4 ( A and B ) HCT-116 or LoVo cells transfected with miR-29a and anti-miR-29a were assayed for invasion. (C and D) KLF4 mediates miR-29a-induced invasion in CRC cells. (C) Invasion assay of HCT-116 cells transfected with miR-29a and/or KLF4 cDNA. (D) Invasion assay of LoVo cells transfected with anti-miR-29a and/or siKLF4. (E and F) Gross and microscopic examination for liver metastasis in both control group and miR-29a overexpression group mice. Magnification: × 10 and × 100. Both the arrow and the letter M indicate metastasis. (G) Quantitative RT–PCR analysis of KLF4 expression in metastatic nodules in the liver of mice. Numerical data are presented as mean±s.d. *P<0.05, **P<0.01, ***P<0.001, n⩾3.
The regulatory effect of the molecular link of miR-29a-KLF4 on MMP2 and E-cad expressions in CRC cells
Subsequently, the underlying mechanism of the molecular link of miR-29a-KLF4 in potentiating CRC cell invasion and CRC metastasis was investigated. It was interestingly to find that miR-29a overexpression promoted MMP2 expression at both mRNA and protein levels, whereas anti-miR-29a inhibited MMP2 expression (Figure 3A and Supplementary Figure S2A). In addition to MMP2, miR-29a downregulated E-cad expression at both mRNA and protein levels, whereas miR-29a silencing promoted E-cad expression (Figure 3A and Supplementary Figure S2B). Furthermore, the regulatory effect of KLF4 on MMP2 and E-cad was analysed using qRT–PCR and western blot. Interestingly, reduced expression of KLF4 by siRNA led to upregulation of MMP2 and downregulation E-cad at both mRNA and protein levels (Figure 3B and Supplementary Figure S2C). In addition, three potential KLF4-binding sites in MMP2 promoter were identified after analysing the promoter of MMP2 (Figure 3C). It was found that KLF4 siRNA significantly increased the promoter activity of MMP2, implicating that the potential KLF4-binding sites was a negative regulating element in the MMP2 promoter (Figure 3D). Subsequent chromatin immunoprecipitation assay showed that the KLF4 ChIPs were enriched in the potential binding sites as compared with the control IgG ChIPs by qRT–PCR (Figure 3E). All these data indicated that the link of miR-29a-KLF4 had important roles in regulating MMP2 and E-cad expression.
The link of miR-29a-KLF4 regulates MMP2 and E-cad expression in CRC cells. (A) Western blot analysis of MMP2 and E-cad expression in HCT-116 cells transfected with miR-29a or miR-Ctrl and in LoVo cells transfected with anti-miR-29a or anti-miR-Ctrl. (B) Western blot analysis of MMP2 and E-cad expression in HCT-116 cells transfected with siKLF4 or siNC. (C) Schematic structure of the MMP2 promoter. Black boxes represent GC-boxes containing putative KLF4-binding sites. The number of −1256 and +299 depicts the location of PCR primers for amplification of MMP2 promoters. (D) KLF4 silencing increased the promoter activity of MMP2. The MMP2-Luc reporter plasmid or pGL3 basic plasmid was co-transfected with renilla and siKLF4/siNC into HEK293T cells. Luciferase values were normalised to renilla levels and expressed as fold change over siNC. (E) Chip analysis of KLF4 at the promoter of MMP2. Anti-KLF4 or control IgG was used to immunoprecipitate DNA–protein complex in HCT-116 cells. The precipitated DNA either by anti-KLF4 or control IgG was depicted as percentages of the total input genome DNA. Numerical data are presented as mean±s.d. *P<0.05, **P<0.01, n=3.
Both high expression of miR-29a and low expression of KLF4 mRNA correlate with metastasis and poor prognosis of CRC
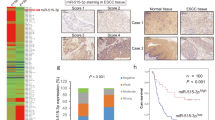
It was found in our previous study that miR-29a expression was increased and KLF4 mRNA expression was reduced in CRC tissues (Fu et al, 2012). To get insight into the clinical implication of miR-29a and KLF4, and the correlation between miR-29a and KLF4 in CRC tissues, the expression level of miR-29a and KLF4 mRNA was analysed in 85 cases of human primary CRC tissues using qRT–PCR. It was observed that the high expression of miR-29a was significantly correlated with metastasis (P=0.028, Mann–Whitney U-test) (Table 1 and Figure 4A), whereas low KLF4 mRNA expression was correlated with metastasis (P=0.029, Mann–Whitney U-test) (Figure 4B). Unsurprisingly, as the target gene of miR-29a, KLF4 mRNA expression inversely correlated with miR-29a expression (Spearman r=−0.271, P=0.013) (Figure 4C). In addition, the Kaplan–Meier analysis revealed that both miR-29a high expression and KLF4 low expression correlated with CRC metastasis and poor OS (Figure 4D and E). Furthermore, the patients with both high miR-29a expression and low KLF4 expression in the same tissue samples had the worst OS (Figure 4F). These data reveal that miR-29a-KLF4 inverse regulation was associated with CRC metastasis and poor survival.
MicroRNA-29a high expression and KLF4 low expression correlate with metastasis and poor survival of CRC patients. (A and B) Quantitative RT–PCR analysis of miR-29a and KLF4 mRNA expression in CRC patients with/without metastasis (A, P=0.028, B, P=0.029, Mann–Whitney U-test). (C) Inverse correlation of miR-29a expression with KLF4 mRNA expression in 85 cases of CRC patients (P=0.013, r=−0.271, χ2 test). (D–F) The Kaplan–Meier analysis of OS of CRC patients with miR-29a and KLF4 mRNA expression profiles (D and E), and subgroup analysis of CRC patients with miR-29a low expression and miR-29a high expression accompanied with KLF4 mRNA low expression and KLF4 mRNA high expression (F).
KLF4 protein low expression is associated with both MMP2 protein high expression and E-cad protein low expression in the CRC tissues
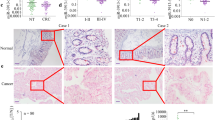
To confirm the regulatory relationship between the negative regulation of MMP2 expression and positive regulation of E-cad expression by KLF4, the 82 pairs of CRC tissues and NCTs were immunostained for KLF4, MMP2 and E-cad (Figure 5A). It was found that the KLF4 and E-cad protein expression levels were lower and MMP2 was higher in CRC tissues than those in NCTs (Figure 5B–D). More importantly, there was a significant inverse correlation between MMP2 and KLF4 expression levels, and a significant positive correlation between E-cad and KLF4 in CRC tissues (Figure 5E and F). These results indicate that MMP2 high expression and E-cad low expression were significantly associated with the loss of KLF4 expression in CRC tumourigenesis.
KLF4 expression is inversely associated with MMP2 expression and positively associated with E-cad expression. Three tissue microarrays of immunostained consecutive sections were performed through specific antibodies against MMP2, E-cad and KLF4. (A) Representative photographs of MMP2, E-cad and KLF4 protein expression levels in normal and cancer specimens (magnification: × 400 for the inserts, × 200 for all others). (B) Matrix metalloproteinase 2 expression level was significantly higher in CRC tissue species than that in the normal colorectal tissue species (P<0.001, χ2test), (C and D) whereas E-cad and KLF4 expression levels were significantly higher in normal colorectal tissue species than those in the CRC tissue species (C), P<0.001, (D), P<0.001, χ2test). (E and F) The KLF4 expression level was inversely correlated with the MMP2 expression level (E), P<0.001, r=−0.400, χ2test) and positively correlated with E-cad expression level (F, P<0.001, r=0.476, χ2test).
Discussion
Although aberrant expression of some miRNAs occurred in CRC metastasis, such as miR-17, miR-19a and miR-21 (Zhang et al, 2012), miR-31 (Wang et al, 2010), miR-103 and miR-107(Chen et al, 2012), miR-143 and miR-145 (Arndt et al, 2009) and miR-200 family (Hur et al, 2012), novel candidates with a definite molecular mechanism in promoting CRC metastasis need to be further explored. It was reported that the dysregulation of miR-29a performed a diverse function in different types of cancer (Gebeshuber et al, 2009; Muniyappa et al, 2009; Eyholzer et al, 2010; Santanam et al, 2010; Cui et al, 2011; Kong et al, 2011; Teichler et al, 2011). It was previously believed that high serum expression of miR-29a in CRC patients was correlated with liver metastasis, and was proposed as a novel biomarker for early and noninvasive diagnosis of CRC (Wang and Gu, 2012). In this study, both data from CRC cell line test and the tissue detection of CRC patients indicate that miR-29a promoted CRC metastasis. Our CRC cell line tests demonstrated that miR-29a directly targeted KLF4 to promote CRC cell invasion in vitro and metastasis in vivo, and MMP2 and E-cad were direct targets of KLF4. Although a previous study reported significant impact of the low miR-29a expression on the risk of recurrence in stage II patients by analysing the RNA from formalin-fixed paraffin-embedded CRC samples (Weissmann-Brenner et al, 2012), our and some other studies observed that miR-29a-KLF4 was associated with CRC metastasis and poor survival (The Cancer Genome Atlas Network (2012)). We also observed that KLF4 protein low expression was associated with both MMP2 protein high expression and E-cad protein low expression in CRC samples. Therefore, the miR-29a-KLF4-MMP2/E-cad axis, in which miR-29a promoted MMP2 and decreased E-cad expression through targeting KLF4, emerges as a novel mechanism for CRC metastasis.
It was reported that the expression of miR-29a was regulated by different signalling pathways in a cell context-dependent manner. The transforming growth factor -beta pathway, which was commonly inactivated at the late stage of CRC, was reported to suppress miR-29a expression in the fibrosis process (Qin et al, 2011). In addition, the canonical WNT signalling pathway induced miR-29a expression in osteoclast cells by the TCF4/LEF1-binding sites in the proximal promoter of miR-29a, whereas miR-29a directly suppressed the WNT antagonist expression, such as Dkk1, Kremen2 and sFRP2, to form a positive feedback loop for WNT signalling (Kapinas et al, 2010). Knowing that both TGF-beta and WNT signalling pathways were reported to be dysregulated in the CRC tumourigenesis (Fodde, 2002; Calon et al, 2012), it will be worthy to explore the correlation between dysregulation of the miR-29a and TGF-beta and WNT signalling pathway in CRC in future studies.
It was previously reported that KLF4 was a highly expressed transcription factor in post-mitotic and terminally differentiated epithelial cells of organs such as the skin, lung and gastrointestinal tract (Shields et al, 1996; Segre et al, 1999). Loss of KLF4 is critical for the pathogenesis and progression of digestive system tumours, including hepatic, gastric, colorectal and pancreatic cancer (Wei et al, 2005; Kanai et al, 2006; Wei et al, 2008; Li et al, 2011, 2012). It was proposed that the regulation of KLF4 expression was involved in both post-transcriptional and transcriptional regulation in CRC. On one hand, pVHL, as a facilitating factor in colorectal oncogenesis, has been demonstrated to physically interact with KLF4 and promote KLF4 protein degradation (Gamper et al, 2012). On the other hand, the decreased mRNA level of KLF4 seems to be the major cause of loss of KLF4 in CRC. Low mRNA expression level of KLF4 could be ascribed to hypermethylation of 5′-untranslated region, hemizygous deletion and point mutations of the KLF4 gene, APC mutation and β-catenin activation in CRC (Dang et al, 2001; Stone et al, 2002; Zhao et al, 2004). Here we demonstrated that miR-29a could directly target KLF4, whereas another group reported that KLF4 was regulated by miR-103/107 in CRC cells (Chen et al, 2012). Both studies reveal a novel miRNA-mediated modulation of KLF4 for the downregulation KLF4.
Another issue is to discover new target genes of KLF4. Although several target genes, such p21, Bmi1 and IFITM3, have been identified as having tumour suppressor function of KLF4 in CRC cells (Zhang et al, 2000; Li et al, 2011; Yu et al, 2012), the mechanism responsible for KLF4 regulating CRC metastasis remains to be elucidated. Matrix metalloproteinase 2 has been proved to promote tumour invasion and metastasis in various tumours (Giannelli et al, 1997; Deryugina and Quigley, 2006). Previous studies have shown that KLF4 inhibited Snail expression (Yori et al, 2011), and Snail significantly increased MMP2 expression (Miyoshi et al, 2004), implying an indirect way of regulation MMP2 by KLF4. In this study, we demonstrated that KLF4 could directly bind to the promoter of MMP2, playing as a transcriptional repressor of MMP2 gene. Thus, KLF4 could inhibit CRC tumour metastasis through suppressing MMP2 expression and promoting E-cad expression (Batlle et al, 2000; Yori et al, 2010) by both direct transcriptional repression and indirect repression through Snail.
In summary, this study has provided insight into the role of the miR-29a-KLF4-MMP2/E-cad axis in CRC metastasis. Our finding that increased expression of miR-29a promoted CRC metastasis by regulating MMP2/E-cad through direct targeting KLF4 highlights the potential of the miR-29a inhibitor as a novel agent against CRC metastasis.
Change history
21 January 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Arndt GM, Dossey L, Cullen LM, Lai A, Druker R, Eisbacher M, Zhang C, Tran N, Fan H, Retzlaff K, Bittner A, Raponi M (2009) Characterization of global microRNA expression reveals oncogenic potential of miR-145 in metastatic colorectal cancer. BMC Cancer 9: 374.
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136 (2): 215–233.
Batlle E, Sancho E, Franci C, Dominguez D, Monfar M, Baulida J, Garcia De Herreros A (2000) The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol 2 (2): 84–89.
Calon A, Espinet E, Palomo-Ponce S, Tauriello DV, Iglesias M, Cespedes MV, Sevillano M, Nadal C, Jung P, Zhang XH, Byrom D, Riera A, Rossell D, Mangues R, Massague J, Sancho E, Batlle E (2012) Dependency of colorectal cancer on a TGF-beta-driven program in stromal cells for metastasis initiation. Cancer Cell 22 (5): 571–584.
Chen HY, Lin YM, Chung HC, Lang YD, Lin CJ, Huang J, Wang WC, Lin FM, Chen Z, Huang HD, Shyy JY, Liang JT, Chen RH (2012) miR-103/107 promote metastasis of colorectal cancer by targeting the metastasis suppressors DAPK and KLF4. Cancer Res 72 (14): 3631–3641.
Choi BJ, Cho YG, Song JW, Kim CJ, Kim SY, Nam SW, Yoo NJ, Lee JY, Park WS (2006) Altered expression of the KLF4 in colorectal cancers. Pathol Res Pract 202 (8): 585–589.
Cui Y, Su WY, Xing J, Wang YC, Wang P, Chen XY, Shen ZY, Cao H, Lu YY, Fang JY (2011) MiR-29a inhibits cell proliferation and induces cell cycle arrest through the downregulation of p42.3 in human gastric cancer. PLoS One 6 (10): e25872.
Dang DT, Chen X, Feng J, Torbenson M, Dang LH, Yang VW (2003) Overexpression of Kruppel-like factor 4 in the human colon cancer cell line RKO leads to reduced tumorigenecity. Oncogene 22 (22): 3424–3430.
Dang DT, Mahatan CS, Dang LH, Agboola IA, Yang VW (2001) Expression of the gut-enriched Kruppel-like factor (Kruppel-like factor 4) gene in the human colon cancer cell line RKO is dependent on CDX2. Oncogene 20 (35): 4884–4890.
Deryugina EI, Quigley JP (2006) Matrix metalloproteinases and tumour metastasis. Cancer Metast Rev 25 (1): 9–34.
Eyholzer M, Schmid S, Wilkens L, Mueller BU, Pabst T (2010) The tumour-suppressive miR-29a/b1 cluster is regulated by CEBPA and blocked in human AML. Br J Cancer 103 (2): 275–284.
Fodde R (2002) The APC gene in colorectal cancer. Eu J Cancer 38 (7): 867–871.
Fu J, Tang W, Du P, Wang G, Chen W, Li J, Zhu Y, Gao J, Cui L (2012) Identifying microRNA-mRNA regulatory network in colorectal cancer by a combination of expression profile and bioinformatics analysis. BMC Systems Biol 6: 68.
Gamper AM, Qiao X, Kim J, Zhang L, DeSimone MC, Rathmell WK, Wan Y (2012) Regulation of KLF4 turnover reveals an unexpected tissue-specific role of pVHL in tumorigenesis. Mol Cell 45 (2): 233–243.
Gebeshuber CA, Zatloukal K, Martinez J (2009) miR-29a suppresses tristetraprolin, which is a regulator of epithelial polarity and metastasis. EMBO Rep 10 (4): 400–405.
Ghaleb AM, McConnell BB, Nandan MO, Katz JP, Kaestner KH, Yang VW (2007) Haploinsufficiency of Kruppel-like factor 4 promotes adenomatous polyposis coli dependent intestinal tumorigenesis. Cancer Res 67 (15): 7147–7154.
Giannelli G, Falk-Marzillier J, Schiraldi O, Stetler-Stevenson WG, Quaranta V (1997) Induction of cell migration by matrix metalloprotease-2 cleavage of laminin-5. Science 277 (5323): 225–228.
Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP (2007) MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27 (1): 91–105.
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144 (5): 646–674.
Hur K, Toiyama Y, Takahashi M, Balaguer F, Nagasaka T, Koike J, Hemmi H, Koi M, Boland CR, Goel A (2012) MicroRNA-200c modulates epithelial-to-mesenchymal transition (EMT) in human colorectal cancer metastasis. Gut 62 (9): 1315–1326.
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ (2009) Cancer statistics, 2009. CA Cancer J Clin 59 (4): 225–249.
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS (2004) Human MicroRNA targets. PLoS Biol 2 (11): e363.
Kanai M, Wei D, Li Q, Jia Z, Ajani J, Le X, Yao J, Xie K (2006) Loss of Kruppel-like factor 4 expression contributes to Sp1 overexpression and human gastric cancer development and progression. Clin Cancer Res 12 (21): 6395–6402.
Kapinas K, Kessler C, Ricks T, Gronowicz G, Delany AM (2010) miR-29 modulates Wnt signaling in human osteoblasts through a positive feedback loop. J Biol Chem 285 (33): 25221–25231.
Kong G, Zhang J, Zhang S, Shan C, Ye L, Zhang X (2011) Upregulated microRNA-29a by hepatitis B virus X protein enhances hepatoma cell migration by targeting PTEN in cell culture model. PLoS One 6 (5): e19518.
Li D, Peng Z, Tang H, Wei P, Kong X, Yan D, Huang F, Li Q, Le X, Xie K (2011) KLF4-mediated negative regulation of IFITM3 expression plays a critical role in colon cancer pathogenesis. Clin Cancer Res 17 (11): 3558–3568.
Li Q, Gao Y, Jia Z, Mishra L, Guo K, Li Z, Le X, Wei D, Huang S, Xie K (2012) Dysregulated Kruppel-like factor 4 and vitamin D receptor signaling contribute to progression of hepatocellular carcinoma. Gastroenterology 143 (3): 799–810 e1-2.
Mina LA, Sledge Jr. GW (2011) Rethinking the metastatic cascade as a therapeutic target. Nat Rev Clin Oncol 8 (6): 325–332.
Miyoshi A, Kitajima Y, Sumi K, Sato K, Hagiwara A, Koga Y, Miyazaki K (2004) Snail and SIP1 increase cancer invasion by upregulating MMP family in hepatocellular carcinoma cells. Br J Cancer 90 (6): 1265–1273.
Muniyappa MK, Dowling P, Henry M, Meleady P, Doolan P, Gammell P, Clynes M, Barron N (2009) MiRNA-29a regulates the expression of numerous proteins and reduces the invasiveness and proliferation of human carcinoma cell lines. Eur J Cancer 45 (17): 3104–3118.
Qin W, Chung AC, Huang XR, Meng XM, Hui DS, Yu CM, Sung JJ, Lan HY (2011) TGF-beta/Smad3 signaling promotes renal fibrosis by inhibiting miR-29. J Am Soc Nephrol 22 (8): 1462–1474.
Remmele W, Stegner HE (1987) [Recommendation for uniform definition of an immunoreactive score (IRS) for immunohistochemical estrogen receptor detection (ER-ICA) in breast cancer tissue]. Der Pathologe 8 (3): 138–140.
Santanam U, Zanesi N, Efanov A, Costinean S, Palamarchuk A, Hagan JP, Volinia S, Alder H, Rassenti L, Kipps T, Croce CM, Pekarsky Y (2010) Chronic lymphocytic leukemia modeled in mouse by targeted miR-29 expression. Proc Natl Acad Sci USA 107 (27): 12210–12215.
Segre JA, Bauer C, Fuchs E (1999) Klf4 is a transcription factor required for establishing the barrier function of the skin. Nature Genet 22 (4): 356–360.
Shields JM, Christy RJ, Yang VW (1996) Identification and characterization of a gene encoding a gut-enriched Kruppel-like factor expressed during growth arrest. J Biol Chem 271 (33): 20009–20017.
Stone CD, Chen ZY, Tseng CC (2002) Gut-enriched Kruppel-like factor regulates colonic cell growth through APC/beta-catenin pathway. FEBS Lett 530 (1-3): 147–152.
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126 (4): 663–676.
Teichler S, Illmer T, Roemhild J, Ovcharenko D, Stiewe T, Neubauer A (2011) MicroRNA29a regulates the expression of the nuclear oncogene Ski. Blood 118 (7): 1899–1902.
The Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature (2012) 487 (7407): 330–337.
Wang CJ, Stratmann J, Zhou ZG, Sun XF (2010) Suppression of microRNA-31 increases sensitivity to 5-FU at an early stage, and affects cell migration and invasion in HCT-116 colon cancer cells. BMC Cancer 10: 616.
Wang LG, Gu J (2012) Serum microRNA-29a is a promising novel marker for early detection of colorectal liver metastasis. Cancer Epidemiol 36 (1): e61–e67.
Wei D, Gong W, Kanai M, Schlunk C, Wang L, Yao JC, Wu TT, Huang S, Xie K (2005) Drastic down-regulation of Kruppel-like factor 4 expression is critical in human gastric cancer development and progression. Cancer Res 65 (7): 2746–2754.
Wei D, Kanai M, Jia Z, Le X, Xie K (2008) Kruppel-like factor 4 induces p27Kip1 expression in and suppresses the growth and metastasis of human pancreatic cancer cells. Cancer Res 68 (12): 4631–4639.
Weissmann-Brenner A, Kushnir M, Lithwick Yanai G, Aharonov R, Gibori H, Purim O, Kundel Y, Morgenstern S, Halperin M, Niv Y, Brenner B (2012) Tumour microRNA-29a expression and the risk of recurrence in stage II colon cancer. Int J Oncol 40 (6): 2097–2103.
Yori JL, Johnson E, Zhou G, Jain MK, Keri RA (2010) Kruppel-like factor 4 inhibits epithelial-to-mesenchymal transition through regulation of E-cadherin gene expression. J Biol Chem 285 (22): 16854–16863.
Yori JL, Seachrist DD, Johnson E, Lozada KL, Abdul-Karim FW, Chodosh LA, Schiemann WP, Keri RA (2011) Kruppel-like factor 4 inhibits tumorigenic progression and metastasis in a mouse model of breast cancer. Neoplasia 13 (7): 601–610.
Yu T, Chen X, Zhang W, Colon D, Shi J, Napier D, Rychahou P, Lu W, Lee EY, Weiss HL, Evers BM, Liu C (2012) Regulation of the potential marker for intestinal cells, Bmi1, by beta-catenin and the zinc finger protein KLF4: implications for colon cancer. J Biol Chem 287 (6): 3760–3768.
Zhang J, Xiao Z, Lai D, Sun J, He C, Chu Z, Ye H, Chen S, Wang J (2012) miR-21, miR-17 and miR-19a induced by phosphatase of regenerating liver-3 promote the proliferation and metastasis of colon cancer. Br J Cancer 107 (2): 352–359.
Zhang W, Geiman DE, Shields JM, Dang DT, Mahatan CS, Kaestner KH, Biggs JR, Kraft AS, Yang VW (2000) The gut-enriched Kruppel-like factor (Kruppel-like factor 4) mediates the transactivating effect of p53 on the p21WAF1/Cip1 promoter. J Biol Chem 275 (24): 18391–18398.
Zhao W, Hisamuddin IM, Nandan MO, Babbin BA, Lamb NE, Yang VW (2004) Identification of Kruppel-like factor 4 as a potential tumor suppressor gene in colorectal cancer. Oncogene 23 (2): 395–402.
Acknowledgements
This work was supported by grants from the Ministry of Science and Technology (2011CB965100, 2011CBA01100, 2011DFA30480, 2010CB944900, 2010CB945000 and 2012CB966603), the National Natural Science Foundation of China (91219305, 31101061, 31210103905, 31071306, 31000378, 31171432, 31201107 and 81170499), the Science and Technology Commission of Shanghai Municipality (10DJ1400500, 10DJ1400504, 11ZR1438500, 11XD1405300, 11DZ1922008 and 11DZ1922009), the Natural Science Foundation of Shanghai (11ZR1424200), IRT1168 and 20110072110039 from the Ministry of Education, 20114221 from Shanghai Municipal Health Bureau, and also supported by ‘Chen Guang’ project supported by Shanghai Municipal Education Commission and Shanghai Education Development Foundation (12CG19), the Fundamental Research Funds for the Central Universities and the Foundation of Shanghai JiaoTong University School of Medicine (11QYJ003).
Author information
Authors and Affiliations
Corresponding authors
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Tang, W., Zhu, Y., Gao, J. et al. MicroRNA-29a promotes colorectal cancer metastasis by regulating matrix metalloproteinase 2 and E-cadherin via KLF4. Br J Cancer 110, 450–458 (2014). https://doi.org/10.1038/bjc.2013.724
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.724
Keywords
This article is cited by
-
Integrated whole transcriptome profiling revealed a convoluted circular RNA-based competing endogenous RNAs regulatory network in colorectal cancer
Scientific Reports (2024)
-
Investigating genomic, proteomic, and post-transcriptional regulation profiles in colorectal cancer: a comparative study between primary tumors and associated metastases
Cancer Cell International (2023)
-
A three-plasma miRNA panel predicts the risk of colorectal cancer: a community-based nested case‒control study
Scientific Reports (2023)
-
Prognostic significance of KLF4 in solid tumours: an updated meta-analysis
BMC Cancer (2022)
-
MiR-29a-3p: a potential biomarker and therapeutic target in colorectal cancer
Clinical and Translational Oncology (2022)