Abstract
Background:
Little is known about the factors that drive metastasis formation in colorectal cancer (CRC). Here, we set out to identify genes and proteins in patients with colorectal liver metastases that correlate with early disease recurrence. Such factors may predict a propensity for metastasis in earlier stages of CRC.
Methods:
Gene expression profiling and proteomics were used to identify differentially expressed genes/proteins in resected liver metastases that recurred within 6 months following liver surgery vs those that did not recur for >24 months. Expression of the identified genes/proteins in stage II (n=243) and III (n=176) tumours was analysed by immunohistochemistry on tissue microarrays. Correlation of protein levels with stage-specific outcome was assessed by uni- and multivariable analyses.
Results:
Both gene expression profiling and proteomics identified Maspin to be differentially expressed in colorectal liver metastases with early (<6 months) and prolonged (>24 months) time to recurrence. Immunohistochemical analysis of Maspin expression on tumour sections revealed that it was an independent predictor of time to recurrence (log-rank P=0.004) and CRC-specific survival (P=0.000) in stage III CRC. High Maspin expression was also correlated with mucinous differentiation. In stage II CRC patients, high Maspin expression did not correlate with survival but was correlated with a right-sided tumour location.
Conclusion:
High Maspin expression correlates with poor outcome in CRC after spread to the local lymph nodes. Therefore, Maspin may have a stage-specific function possibly related to tumour cell dissemination and/or metastatic outgrowth.
Similar content being viewed by others
Main
Five-year survival rates in colorectal cancer (CRC) vary dramatically from ∼93% in patients with localised stage I tumours to ∼6% in patients with metastasised inoperable disease (Siegel et al, 2012). The decision to administer adjuvant chemotherapy in CRC is predominantly stage-dependent. All stage III and high-risk stage II patients are recommended to receive adjuvant treatment. In contrast, patients with stage I and low-risk II disease generally do not receive adjuvant chemotherapy after resection of the primary tumour, as 5-year survival rates are high and there is only minimal benefit in unselected patients (O’Connell et al, 2004; Edge and Compton, 2010). Recently, gene expression profiling has been used to identify high-risk stage II patients who can possibly benefit from adjuvant chemotherapy. Two published gene signatures have proven to be able to select stage II cancer patients for adjuvant treatment: the Oncotype DX Colon Cancer Assay (12 genes) and the ColoPrint (18 genes). Both signatures seem to perform better than most conventional risk factors (Salazar et al, 2010; Webber et al, 2010; Kelley et al, 2011) and are currently being further validated. The discovery of novel biomarkers for predicting tumour behaviour is not only valuable from a prognostic viewpoint, but can also help to gain insight into the mechanisms that cause disease progression and metastasis formation. Differences in tumour cell survival, migration, invasion and extracellular matrix remodelling are all likely to have a role (Hanahan and Weinberg, 2000).
To turn a locally invasive colorectal tumour into a metastasising one very few, if any, additional genetic changes are required (Jones et al, 2008). It is possible that the factors driving initial metastasis formation from the primary tumour also promote tumour recurrence at later stages of the disease, following surgical removal of liver metastases (Eberhard et al, 2012). After resection of colorectal liver metastases, the vast majority of patients experience tumour recurrence within 2 years. However, the patients that remain disease-free beyond 2 years have a good chance of 5-year survival without additional therapy (Nordlinger et al, 1996; Fong et al, 1999; Simmonds et al, 2006; de Jong et al, 2009). This is most likely due to favourable tumour characteristics.
The rationale of this study was to obtain further insight into the mechanisms causing tumour progression and metastases formation. Gene expression profiling and proteomics were used to detect differences between stage IV CRC patients that recur within 6 months after resection vs those that fail to recur for >2 years. Subsequently, we analysed whether expression of the identified factors also predicts metastasis formation in early stages of CRC.
Materials and methods
Patients and samples
Microarray and mass-spectrometry analysis
Biopsies of 30 liver metastases were used to identify genes and proteins correlating with early recurrence in stage IV CRC patients. Patient, tumour and surgical characteristics were derived from our prospectively collected database. Thirty frozen tumour biopsies were collected between July 2003 and August 2008 at the University Medical Centre in Utrecht, the Netherlands (Data set 1) (Figure 1). Samples were included if patients were aged >18 years and received curative resection for histologically confirmed liver metastases from CRC. Patients with a history of non-colorectal malignancies, extra hepatic disease or microscopic residual disease (R1) after surgery and patients who received local ablation therapy or chemoembolization alone or in combination with resection were excluded. Only those specimens were included that were snap-frozen in liquid nitrogen within 30 min after resection and stored at −80 °C. The study protocol was approved by The Medical Ethical Committee (MEC) of the University Medical Center Utrecht as recognised by article 16 of the WMO (Dutch Law on Medical Research with human subjects). Written informed consent was obtained from all patients in Data set 1.
Tissue microarray study
Between 1996 and 2005, 419 patients underwent surgical colon cancer resection at the Kennemer Gasthuis Hospital in Haarlem, the Netherlands, that were classified as stage II (T3–4, N0, M0) or stage III (T1–4, N1–2, M0) according to the fourth edition of the TNM classification system (Data set 2) (Figure 1). These tumour samples were used to examine any stage-specific role of the biomarkers retrieved from Data set 1. Patient, tumour and surgical characteristics were retrospectively drafted from clinical and pathology reports. Collection, storage and use of tissue and patient data were performed in agreement with the ‘Code for Proper Secondary Use of Human Tissue in The Netherlands’ (available at http://www.federa.org/codes-conduct) (Belt et al, 2011).
Gene expression profiling
RNA isolation, labelling and hybridisation to whole-genome oligonucleotide high-density microarrays were performed as previously described (Snoeren et al, 2012). In short, two expression profiles in dye-swap experiments were generated for each sample. The samples were compared against a commercial reference (Universal Human Reference RNA catalogue #740 000, Stratagene, La Jolla, CA, USA). The Human Array-Ready Oligo set (version 2.0) was purchased from Qiagen (Venlo, Netherlands) and spotted on Codelink slides (GE Healthcare, Little Chalfont, UK) in a dust-filtered and humidity controlled clean room. The microarrays contained 70-mer oligonucleotides representing 21.329 human genes and expressed sequence tags (ESTs), as well as 3871 additional spots for control purposes. We applied total RNA and cRNA quality control criteria in accordance with the Tumour Analysis Best Practices Working Group (2004). Gene expression in metastases showing recurrence within 6 months after resection was compared with metastases that did not show recurrence for >2 years by using ANOVA (Wu et al, 2003). In a fixed effect analysis, sample, array and dye effects were modelled. P-values were determined by a permutation F2 test in which residuals were shuffled 5000 times globally. Genes with P <0.05 after family-wise error correction were considered significant.
Mass-spectrometry analysis
Tissue homogenisation
Frozen tumour tissue sections of five stage IV patients with short time to recurrence (<6 months) and five patients with prolonged time to recurrence (>24 months) were used for mass-spectrometry analysis. All patients were selected from Data set 1. The frozen tumour tissue sections were cut into pieces of ∼20 mg, after which they were solubilized in 800 μl SDS sample buffer (containing 62.5 mm Tris-HCl, 2% w/v SDS, 10% v/v glycerol, 0.0025% bromphenol blue, 100 mm DTT, pH 6.8) using a micro grinder. Denaturation was done by heating samples for 10 min at 100 °C. All remaining insoluble parts were removed by centrifugation (15 min; 14 000 r.p.m.).
Fractionation using gel electrophoresis
Equal amounts of protein (50 μg) were separated on NuPAGE Novex Bis-Tris Mini Gels (Invitrogen, Bleiswijk, Netherlands). Gels were stained with Coomassie brilliant blue G-250 (Pierce, Rockford, IL, USA, Bleiswijk, Netherlands), washed and each lane was sliced into 10 bands using a band pattern to guide the slicing. The gel slicing and in-gel digestion was performed in a laminar flow under keratin-free conditions.
In-gel digestion, NanoLC-MS/MS analysis
In-gel digestion and NanoLC-MS/MS analysis was performed as described by van Houdt et al (2011).
Protein identification
MS/MS spectra were searched against IPI human 3.62 (83947 entries) using Sequest version 27, rev 12 (Thermo, San Jose, CA, USA). Cysteine carboxamidomethylation and methionine oxidation were treated as variable modifications. Peptides precursor ions were searched with a maximum mass deviation of 10 p.p.m. and fragment ions with a maximum mass deviation of 1 Da. Sequest output files were imported in Scaffold 2.06.1 (Proteome software, Portland, OR, USA) and search results of the 10 gel bands per biological sample were combined. A protein was considered identified when at least two unique peptides were identified in one of the samples. Peptides were identified with a PeptideProphet (Keller et al, 2002) probability score of >95% and a ProteinProphet (Nesvizhskii et al, 2003) probability score of >99%. Proteins were (label-free) quantified by spectral counting (Liu et al, 2004; Pham et al, 2010), that is, the sum of all MS/MS spectra for each identified protein. For each biological sample the spectral counts for each protein were normalised on the sum of the counts for that sample and multiplied by the average of the sums across samples. Subsequently, the beta-binominal test was applied to detect significantly different proteins between early and late recurrence samples.
Tissue microarray construction
Tissue microarrays (TMAs) were constructed from stage II and III colon cancer samples as described previously (Simon et al, 2004; Belt et al, 2011). In summary, formalin-fixed, paraffin-embedded specimens of resected colon cancer tumours were used as donor blocks. Three 0.6-mm cores were taken from the centre of the tumour and three cores from the periphery of the tumour and transferred into recipient TMA paraffin blocks, resulting in six cores per tumour. The maximum number of samples that was transferred to a single TMA was 264.
Immunohistochemistry
Formalin-fixed paraffin-embedded cores were deparaffinized with xylene and rehydrated in decreasing ethanol dilutions. Endogenous peroxidase activity was blocked with 3% hydrogen peroxide. Antigen retrieval was achieved by boiling slides in citrate buffer (pH 6) for 20 min. Slides were then incubated (overnight 4 °C) with the monoclonal anti-Maspin antibody (clone G167-70; Pharmingen, San Jose, CA, USA) at a 1 : 1000 dilution. For detection, goat anti mouse poly-HRP (Powervision, Immunologic, Immunovision Technologies, Brisbane, CA, USA) was used. All slides were developed with diaminobenzidine (DAB). Slides were counterstained with filtered hematoxylin, dehydrated through a graded series of ethanol, immersed in xylene and mounted.
Immunoreactivity for Maspin was assessed and scored by two independent investigators who were blinded to clinical, pathological or survival data. The presence of nuclear and cytoplasmic staining in the tumour cells was assessed separately in each sample and divided into high and low scores. The immunohistochemistry scoring system used is based on scoring systems used in previous studies (Bettstetter et al, 2005; Dietmaier et al, 2006). A combination of intensity (negative, 1+, 2+ or 3+) and percentage of stained cells was used to score each core biopsy. Intensity score of 2+ in >50% of cells or 3+ in >10% of cells was considered high (Figure 4). In case of discrepancies between the scoring by the two investigators, the slides were reviewed and a consensus was reached. For analytical purposes protein expression was divided into high and low scores combining cytoplasmic and nuclear scores (see Table 1).
Western blotting
The homogenised tumour tissue from all five patients with short time to recurrence (<6 months), which was used for mass-spectrometry analysis, was used for western blot validation. Equal amounts of protein was run out on NuPAGE Novex Tris-Acetate Mini Gel (Invitrogen) and was analysed by western blotting using primary antibodies directed against Maspin (clone G167-70; Pharmingen) and β-Actin (AC-15, Novus Biologicals, Littleton, CO, USA) in combination with secondary antibody peroxidase-conjugated anti-mouse IgG (Dako, Glostrup, Denmark). For detection of antibodies Amersham ECL Western Blotting Detection Reagent was used (GE Healthcare Life Sciences).
Follow-up and survival
All patients were subjected to routine follow-up. The follow-up data were updated by letters and telephone calls to referring physicians and general practitioners. The duration of the follow-up and the time between surgery and the detection of recurrence were obtained, in addition to CRC-specific survival.
Statistical analysis
Time-to-recurrence and CRC-specific survival data were calculated from the day of surgery to the day of the first recurrence or the day of death caused by CRC. Median time to recurrence and CRC-specific survival were estimated by the Kaplan–Meier method. To determine the influence of possible risk factors on time to recurrence and CRC-specific survival, a univariable Cox regression analysis was performed. A multivariable Cox proportional hazards model, containing the factors that displayed P-values <0.2 in univariable analysis, was used to determine the independent prognostic impact of all variables on time to recurrence and CRC-specific survival. Statistical significance was assumed for P-values <0.05. Statistical analyses were performed using SPSS for Windows version 15.0 (SPSS, Chicago, IL, USA).
Results
Gene expression profiling identifies Maspin as a marker for recurrence in stage IV CRC
Forty-six patients from Data set 1 fulfilled the inclusion- and quality control criteria. To identify genes that are differentially expressed, patients with metastases showing early (<6 months, n=17) and prolonged (>24 months, n=13) time to recurrence were selected (Table 2). Gene expression profiling of these 30 tumours revealed that 14 genes were significantly differentially expressed between the two groups: SERPINB5, THEM2, GDF15, LYPLA1, AMPD1, ARL6IP5, LAPTM4A, C13ORF3, COLEC11, FYTTD1, BNIP3, SMYD2, GTF3c3 AND ASAP2. Of these genes, 11 were upregulated in the group with time to recurrence of <6 months (Table 3). SERPINB5, also known as Maspin, was the gene most upregulated in patients with a short time to recurrence. Maspin expression was ∼2.1 times higher in tumours of patients with a short time to recurrence vs tumours in patients with a prolonged time to recurrence (Figure 2A; P=0.01).
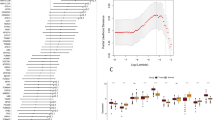
Microarray analysis of patients with short and prolonged time to recurrence identifies Maspin to be differentially expressed in stage IV CRC. (A) Median expression levels of Maspin in the early (<6 months, n=17) and prolonged (>24 months, n=13) time-to-recurrence groups (P=0.012). (B) Kaplan–Meier curves of 46 patients in Data set 1, illustrating time to recurrence in stage IV CRC liver metastasis patients with high Maspin expression (>median) and low Maspin expression (<median) (P=0.005, log-rank test). Median Maspin expression was used as a cutoff (168.6). (C) Kaplan–Meier curves illustrating time to recurrence in primary stage IV CRC patients with high Maspin expression (>median, n=27) and low Maspin expression (<median, n=26) using a previosuly published data set (Smith et al, 2010) (P=0.005). Median Maspin expression was used as the cutoff value (259.6). (D) Kaplan–Meier curves illustrating overall survival in 53 primary stage IV CRC patients with high Maspin expression (>median) and low Maspin expression (<median) using the study by Smith et al (2010) (P=0.029). Median Maspin expression was used as the cutoff value (259.6).
The Kaplan–Meier survival curves of all 46 patients from Data set 1 (including the 30 patients used for gene expression profiling) (Figure 2B) show that the 2-year recurrence-free survival probability of patients with Maspin-high tumours (>median (168.6) is an estimated 0.10 (95% CI=0.00–0.20) compared with 0.39 (95% CI=0.18–0.60) in patients with Maspin-low tumours (<median (168.6) (Cox regression analysis P=0.007, HR=2.65, 95% CI=1.307–5.380). Multivariable analysis shows that Maspin is an independent predictor of early recurrence (P=0.02, HR=2.971, 95% CI=1.168–7558) in our training set. Adjuvant treatment was the only other independent predictor of early recurrence in multivariable analysis (P=0.05, HR=0.329, 95% CI=0.109–0.994) (Supplementary Table 1).
This result was validated using an independent microarray data set from a cohort of 53 patients with stage IV primary CRC (Smith et al, 2010). Two Kaplan–Meier graphs, generated by the R2 microarray analysis and visualisation platform (http://r2.amc.nl) show that high levels (>median (259.6)) of Maspin are significantly correlated with disease-specific (Figure 2C; P=0.005) and overall survival (Figure 2D; P=0.029) in primary stage IV CRC patients.
Mass-spectrometry analysis confirms Maspin as marker for early recurrence in stage IV CRC
In parallel to gene expression profiling, we also analysed differentially expressed proteins in patients, from Data set 1, showing early (<6 months) and prolonged (>24 months) time to recurrence. To this end, mass-spectrometry analysis was performed on five tumours from each group (Supplementary Table 2). Protein-containing lysates of these tumour tissue samples were fractionated on an SDS–PAGE gel, followed by in-gel tryptic digestion (Figure 3A). Analysis of the extracted peptides was performed by Nano-LC-MS/MS, followed by database searching. In total, 2097 unique proteins were identified in all patient samples. Forty-six proteins were present in significantly different amounts in the two groups, of which 26 were overrepresented and 20 were underrepresented in the tumours of patients with early recurrence (Table 4). The sets of 14 genes and 46 proteins that showed a significant association with short or prolonged recurrence times contained only 1 overlapping factor: Maspin (Figure 3B). Validation of the mass-spectrometry results by western blotting showed that the number of spectral counts detected by mass-spectrometry analysis correlates very well with Maspin protein levels (Figure 3C). These results demonstrate that Maspin is differentially expressed on both mRNA and protein level in stage IV CRC patients with early and late recurrence.
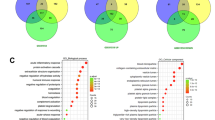
Mass-spectrometry based analysis of proteome differences between patients with early and prolonged time to recurrence in stage IV CRC. (A) Coomassie-stained protein gradient gel loaded with protein samples from tumours form patients with short (<6 months) and prolonged (>24 months) time to recurrence. This gel was used for mass-spectrometry analysis. (B) Venn diagram showing overlap between differently expressed genes and proteins found by micro-array and mass-spectrometry analysis. (C) Western blot analysis of Maspin levels in patients with early time to recurrence (<6 months) and the corresponding amount of spectral counts detected by mass-spectrometry analysis.
Tissue microarray analysis reveals that Maspin staining in the tumour centre is a stage-specific marker in CRC
The above results indicate that Maspin could be a prognostic marker for early recurrence in stage IV CRC patients. This prompted us to assess whether this marker had a similar prognostic power in stage II and III CRC patients. To this end, a total of 243 stage II and 166 stage III tumours on TMAs containing FFPE tissue cores were stained with specific monoclonal Maspin antibody (Bettstetter et al, 2005; Dietmaier et al, 2006; Fung et al, 2010). The presence of nuclear and cytoplasmic staining in both central tumour cores and peripheral tumour cores was assessed in each sample and divided in high and low scores. A high score was defined as an intensity score of 2+ in >50% of cells or 3+ in >10% of cells (Table 1, Figure 4).
Representative immunohistochemical Maspin staining in CRC specimens. (A) Cytoplasmic immunoreactivity with Maspin. Upper panel representing low scores: negative (i) intensity 1+ in >50% of cells (ii). Lower panel representing high scores: intensity 2+ (iii) and 3+ (iv) in >50% of cells. (B) Nuclear immunoreactivity of Maspin. Upper panel representing low scores: intensity 2+ in<50% of cells (i/ii). Lower panel representing high scores: intensity 3+ in>10% of cells (iii/iv).
In stage II CRC tumours (n=243), Maspin staining in the central tumour cores showed no significant differences in time-to-recurrence and CRC-specific survival times (Table 5). Kaplan–Meier curves of Maspin are depicted in Figure 5A and B. High Maspin expression was found to be correlated with a right-sided tumour location (P=0.001) (Supplementary Table 3).
In stage III CRC high Maspin expression in central cores is correlated with early time to recurrence and colorectal-specific death. (A) Kaplan–Meier curves illustrating the effects of different Maspin expression levels on time to recurrence in stage II CRC patients (P=0.98, log-rank test). (B) Kaplan–Meier curves illustrating the effect of different Maspin expression levels on colorectal-specific survival in stage II CRC patients (P=0.95, log-rank test). (C) Kaplan–Meier curves illustrating the effects of different Maspin expression levels on time to recurrence in stage III CRC patients (P=0.011, log-rank test). (D) Kaplan–Meier curves illustrating the effect of different Maspin expression levels on colorectal specific survival in stage III CRC patients (P=0.000, log-rank test). *The low cytoplasmic and high nuclear group were left out of the analysis due to the low number of tumours in this group (n=4 in stage II, n=2 in stage III).
In stage III CRC patients (n=166), Maspin staining in the central tumour cores was an independent predictor of time-to-recurrence and CRC-specific survival (Table 6, Figure 5C and D). Patients with combined low cytoplasmic and nuclear staining score had estimated 2-year recurrence-free and CRC-specific survival probabilities of 0.74 (95% CI=0.64–0.84) and 0.83 (95% CI=0.74–0.92; log-rank P=0.011), respectively. Patients with a combined high cytoplasmic and nuclear score did markedly worse with estimated 2-year recurrence-free and CRC-specific survival probabilities of 0.42 (95% CI=0.25–0.59) and 0.55 (0.38–0.72; log-rank P=0.000), respectively. The low-cytoplasmic and high-nuclear group was left out of the analysis due to the low number of tumours in this group (n=4 in stage II, n=2 in stage III).
High Maspin expression in the central tumour cores was also associated with poor histological grade, mucinous differentiation and MSI-high status in stage III CRC patients (Supplementary Table 4). In multivariable analysis, mucinous differentiation was the only independent factor associated with high Maspin expression.
Maspin staining in peripheral cores showed no correlation with time to recurrence or CRC-specific survival in stage II patients or stage III patients (Tables 5 and 6 and Supplementary Tables 5 and 6). The low-cytoplasmic and high-nuclear group was left out of the analysis due to the low number of tumours in this group (n=4 in stage II and n=2 in stage III).
We next validated these results using the microarray data from a cohort of 75 patients with stage II CRC and 78 patients with stage III CRC (Jorissen et al, 2009; Smith et al, 2010). Kaplan–Meier graphs, generated by the R2 microarray analysis and visualisation platform (http://r2.amc.nl), show that high levels of Maspin (>median 486.2) are significantly (P=0.02) correlated with early recurrence in stage III but not in stage II CRC patients (Figure 6).
External validation, using the SieberSmith database, confirms that Maspin is a stage dependent prognostic marker. (A) Kaplan–Meier curves illustrating that high (>median) and low (<median) Maspin expression have no significant effect on time to recurrence in stage II CRC patients (n=70) (P=0.5). Median Maspin expression was used as the cutoff value (426.3). (Jorissen et al, 2009; Smith et al, 2010). (B) Kaplan–Meier curves illustrating that stage III CRC patients (n=75) with high (>median) Maspin expression have significant shorter time to recurrence than patients with low (<median) Maspin expression (P=0.02). Median Maspin expression was used as the cutoff value (486.2). (Jorissen et al, 2009; Smith et al, 2010).
Discussion
In this report we identified Maspin as a marker for early recurrence after surgery for colorectal liver metastases. Maspin is a prognostic factor for early recurrence in stage III and IV CRC patients but not in stage II patients. Maspin is a member of the serine protease inhibitor (serpin) family. In breast cancer, Maspin acts as a tumour suppressor by inhibiting tumour cell motility, invasion and tumour growth (Zou et al, 1994). In breast, ovarian and lung cancer Maspin expression is correlated with a relatively good prognosis (Marioni et al, 2005; Zheng et al, 2008). By contrast, studies in lung, breast, gastric and pancreatic cancer show that Maspin expression is associated with more aggressive disease (Umekita et al, 2002; Cao et al, 2007; Woenckhaus et al, 2007; Yu et al, 2007). The reasons for these discrepancies are presently unknown.
Mixed-stage studies in CRC have failed to reach consensus whether Maspin is associated with good or poor prognosis (Song et al, 2002; Bettstetter et al, 2005; Markl et al, 2010). A study in patients with only stage III disease has shown that high nuclear Maspin is significantly correlated with poor overall survival (Dietmaier et al, 2006), which is in line with the results presented in this report. We used a combined nuclear and cytoplasmic expression score. The group with high-cytoplasmic and high-nuclear staining is associated with early recurrence. By contrast, the groups with low-nuclear and either high- or low-cytoplasmic staining are associated with late recurrence. This suggests that high-nuclear Maspin staining is the major variable determining the association with early recurrence. This is in line with earlier reports (Dietmaier et al, 2006; Markl et al, 2010). In contrast to the results presented here, a previous study failed to detect a significant association of nuclear or cytoplasmic Maspin staining with outcome in a large cohort of stage III CRC patients (Fung et al, 2010). As all three studies made use of the same antibody, the different outcome of the study of Fung et al (2010) may be related to the use of a different scoring system. The cutoffs used in this study resulted in a very unevenly distributed number of samples per group, in which the vast majority (75%) of samples were allocated to one group. Moreover, this study did not take the intensity of the staining into account, which has previously been associated with the amount of Maspin expression and clinical outcome (Dietmaier et al, 2006). However, when we apply the Fung scoring system to our own data set, we still find that nuclear Maspin expression is associated with disease-free and CRC-specific survival in stage III CRC patients (P=0.031 and P=0.001; data not shown). The same significant association between high levels of Maspin and early-recurrence and colorectal-specific survival was seen when applying the Dietmaier et al (2006) scoring system (P=0.040 and P=0.019; data not shown).
Tumours with high Maspin expression respond significantly better to adjuvant 5-FU chemotherapy in stage III CRC (Dietmaier et al, 2006). This trend was also observed in our study, although patient numbers were too low for this association to reach significance (Data not shown). Maspin may therefore not only have value as a prognostic marker but possibly also as a marker for predicting response to adjuvant treatment in advanced stage CRC. Whether Maspin itself could be a drug target depends on whether it is causally involved in facilitating the metastatic process.
In this study, Maspin was highly upregulated using two different high-throughput screens suggestive of a key role in predicting time to recurrence after surgery of colorectalliver metastasis. To our knowledge, this is the first study reporting a possible predictive role for Maspin in stage IV CRC. However, it must be noted that identification of Maspin and validation was performed in a relatively small cohort.
Experimental studies show that high Maspin expression is correlated with increased apoptosis resistance in CRC cells (Payne et al, 2011). Furthermore, it has been shown that circulating tumour cells in peripheral blood of CRC patients express high levels of Maspin (Findeisen et al, 2008). We did not find a difference in Maspin expression at the periphery of the tumour compared with that at the centre; however, Bettstetter et al (2005) did find that Maspin expression levels were higher at the invasive tumour font when compared with that at the more central parts of CRC tumours. This discrepancy can probably be explained by the fact that we scored peripheral cores containing mainly tumour bulk, but not necessarily cells invading the adjacent stroma, whereas Bettstetter et al (2005) only scored Maspin expression in cell clusters detaching from the peripheral tumour bulk.
These results combined with the results of our study, demonstrating that high Maspin expression is associated with poor prognosis in CRC that has metastasised to local lymph nodes or beyond, point to a potential role for Maspin in facilitating the metastatic process. Future work should therefore address whether (and how) high Maspin levels promote metastasis in advanced-stage CRC.
Change history
17 September 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Belt EJ, Fijneman RJ, van den Berg EG, Bril H, Delis-van Diemen PM, Tijssen M, van Essen HF, de Lange-de Klerk ES, Belien JA, Stockmann HB, Meijer S, Meijer GA (2011) Loss of lamin A/C expression in stage II and III colon cancer is associated with disease recurrence. Eur J Cancer 47: 1837–1845.
Bettstetter M, Woenckhaus M, Wild PJ, Rummele P, Blaszyk H, Hartmann A, Hofstadter F, Dietmaier W (2005) Elevated nuclear maspin expression is associated with microsatellite instability and high tumour grade in colorectal cancer. J Pathol 205: 606–614.
Cao D, Zhang Q, Wu LS, Salaria SN, Winter JW, Hruban RH, Goggins MS, Abbruzzese JL, Maitra A, Ho L (2007) Prognostic significance of maspin in pancreatic ductal adenocarcinoma: tissue microarray analysis of 223 surgically resected cases. Mod Pathol 20: 570–578.
de Jong MC, Pulitano C, Ribero D, Strub J, Mentha G, Schulick RD, Choti MA, Aldrighetti L, Capussotti L, Pawlik TM (2009) Rates and patterns of recurrence following curative intent surgery for colorectal liver metastasis: an international multi-institutional analysis of 1669 patients. Ann Surg 250: 440–448.
Dietmaier W, Bettstetter M, Wild PJ, Woenckhaus M, Rummele P, Hartmann A, Dechant S, Blaszyk H, Pauer A, Klinkhammer-Schalke M, Hofstadter F (2006) Nuclear Maspin expression is associated with response to adjuvant 5-fluorouracil based chemotherapy in patients with stage III colon cancer. Int J Cancer 118: 2247–2254.
Eberhard J, Gaber A, Wangefjord S, Nodin B, Uhlen M, Ericson LK, Jirstrom K (2012) A cohort study of the prognostic and treatment predictive value of SATB2 expression in colorectal cancer. Br J Cancer 106: 931–938.
Edge SB, Compton CC (2010) The American Joint Committee on Cancer: the 7th edn. of the AJCC cancer staging manual and the future of TNM. Ann Surg Oncol 17: 1471–1474.
Findeisen P, Rockel M, Nees M, Roder C, Kienle P, Von Knebel DM, Kalthoff H, Neumaier M (2008) Systematic identification and validation of candidate genes for detection of circulating tumour cells in peripheral blood specimens of colorectal cancer patients. Int J Oncol 33: 1001–1010.
Fong Y, Fortner J, Sun RL, Brennan MF, Blumgart LH (1999) Clinical score for predicting recurrence after hepatic resection for metastatic colorectal cancer: Analysis of 1001 consecutive cases. Ann Surg 230: 309–318.
Fung CL, Chan C, Jankova L, Dent OF, Robertson G, Molloy M, Bokey L, Chapuis PH, Lin BP, Clarke SJ (2010) Clinicopathological correlates and prognostic significance of maspin expression in 450 patients after potentially curative resection of node-positive colonic cancer. Histopathology 56: 319–330.
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100: 57–70.
Jones S, Chen WD, Parmigiani G, Diehl F, Beerenwinkel N, Antal T, Traulsen A, Nowak MA, Siegel C, Velculescu VE, Kinzler KW, Vogelstein B, Willis J, Markowitz SD (2008) Comparative lesion sequencing provides insights into tumour evolution. Proc Natl Acad Sci USA 105: 4283–4288.
Jorissen RN, Gibbs P, Christie M, Prakash S, Lipton L, Desai J, Kerr D, Aaltonen LA, Arango D, Kruhoffer M, Orntoft TF, Andersen CL, Gruidl M, Kamath VP, Eschrich S, Yeatman TJ, Sieber OM (2009) Metastasis-associated gene expression changes predict poor outcomes in patients with Dukes stage B and C colorectal cancer. Clin Cancer Res 15: 7642–7651.
Keller A, Nesvizhskii AI, Kolker E, Aebersold R (2002) Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 74: 5383–5392.
Kelley RK, Van Bebber SL, Phillips KA, Venook AP (2011) Personalized medicine and oncology practice guidelines: a case study of contemporary biomarkers in colorectal cancer. J Natl Compr Canc Netw 9: 13–25.
Liu H, Sadygov RG, Yates JR III (2004) A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal Chem 76: 4193–4201.
Marioni G, Blandamura S, Giacomelli L, Calgaro N, Segato P, Leo G, Fischetto D, Staffieri A, De FC (2005) Nuclear expression of maspin is associated with a lower recurrence rate and a longer disease-free interval after surgery for squamous cell carcinoma of the larynx. Histopathology 46: 576–582.
Markl B, Arnholdt HM, Jahnig H, Schenkirsch G, Herrmann RA, Haude K, Spatz H, Anthuber M, Schlimok G, Oruzio D (2010) Shift from cytoplasmic to nuclear maspin expression correlates with shorter overall survival in node-negative colorectal cancer. Hum Pathol 41: 1024–1033.
Nesvizhskii AI, Keller A, Kolker E, Aebersold R (2003) A statistical model for identifying proteins by tandem mass spectrometry. Anal Chem 75: 4646–4658.
Nordlinger B, Guiguet M, Vaillant JC, Balladur P, Boudjema K, Bachellier P, Jaeck D (1996) Surgical resection of colorectal carcinoma metastases to the liver. A prognostic scoring system to improve case selection, based on 1568 patients. Association Francaise de Chirurgie. Cancer 77: 1254–1262.
O’Connell JB, Maggard MA, Ko CY (2004) Colon cancer survival rates with the new American Joint Committee on Cancer sixth edition staging. J Natl Cancer Inst 96: 1420–1425.
Payne CM, Holubec H, Crowley-Skillicorn C, Nguyen H, Bernstein H, Wilcox G, Bernstein C (2011) Maspin is a deoxycholate-inducible, anti-apoptotic stress-response protein differentially expressed during colon carcinogenesis. Clin Exp Gastroenterol 4: 239–253.
Pham TV, Piersma SR, Warmoes M, Jimenez CR (2010) On the beta-binomial model for analysis of spectral count data in label-free tandem mass spectrometry-based proteomics. Bioinformatics 26: 363–369.
Salazar R, Roepman P, Capella G, Moreno V, Simon I, Dreezen C, Lopez-Doriga A, Santos C, Marijnen C, Westerga J, Bruin S, Kerr D, Kuppen P, van d V, Morreau H, Van VL, Glas AM, Van’t Veer LJ, Tollenaar R (2010) Gene expression signature to improve prognosis prediction of stage II and III colorectal cancer. J Clin Oncol 29 (1): 17–24.
Siegel R, Naishadham D, Jemal A (2012) Cancer statistics 2012. CA Cancer J Clin 62: 10–29.
Simmonds PC, Primrose JN, Colquitt JL, Garden OJ, Poston GJ, Rees M (2006) Surgical resection of hepatic metastases from colorectal cancer: a systematic review of published studies. Br J Cancer 94: 982–999.
Simon R, Mirlacher M, Sauter G (2004) Tissue microarrays. Biotechniques 36: 98–105.
Smith JJ, Deane NG, Wu F, Merchant NB, Zhang B, Jiang A, Lu P, Johnson JC, Schmidt C, Bailey CE, Eschrich S, Kis C, Levy S, Washington MK, Heslin MJ, Coffey RJ, Yeatman TJ, Shyr Y, Beauchamp RD (2010) Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138: 958–968.
Snoeren N, van H Sr, Adam R, van HR, Voest EE, Guettier C, van Diest PJ, Nijkamp MW, Brok MO, van LD, Koerkamp MJ, Holstege FC, Rinkes IH (2012) Exploring gene expression signatures for predicting disease free survival after resection of colorectal cancer liver metastases. PLoS One 7: e49442.
Song SY, Lee SK, Kim DH, Son HJ, Kim HJ, Lim YJ, Lee WY, Chun HK, Rhee JC (2002) Expression of maspin in colon cancers: its relationship with p53 expression and microvessel density. Dig Dis Sci 47: 1831–1835.
Tumour Analysis Best Practices Working Group (2004) Expression profiling–best practices for data generation and interpretation in clinical trials. Nat Rev Genet 5: 229–237.
Umekita Y, Ohi Y, Sagara Y, Yoshida H (2002) Expression of maspin predicts poor prognosis in breast-cancer patients. Int J Cancer 100: 452–455.
van Houdt WJ, Emmink BL, Pham TV, Piersma SR, Verheem A, Vries RG, Fratantoni SA, Pronk A, Clevers H, Borel RI, Jimenez CR, Kranenburg O (2011) Comparative proteomics of colon cancer stem cells and differentiated tumour cells identifies BIRC6 as a potential therapeutic target. Mol Cell Proteomics 10: M111.
Webber EM, Lin JS, Evelyn PW (2010) Oncotype DX tumour gene expression profiling in stage II colon cancer. Application: prognostic, risk prediction. PLoS Curr 2: RRN1177.
Woenckhaus M, Bubendorf L, Dalquen P, Foerster J, Blaszyk H, Mirlacher M, Soler M, Dietmaier W, Sauter G, Hartmann A, Wild PJ (2007) Nuclear and cytoplasmic Maspin expression in primary non-small cell lung cancer. J Clin Pathol 60: 483–486.
Wu H, Kerr M.K, Cui, Churchill G. A. (2003) MAANOVA: A software package for the analysis of spotted cDNA microarray experiments. The analysis of gene expression data: Methods and software. Ref type: generic.
Yu M, Zheng H, Tsuneyama K, Takahashi H, Nomoto K, Xu H, Takano Y (2007) Paradoxical expression of maspin in gastric carcinomas: correlation with carcinogenesis and progression. Hum Pathol 38: 1248–1255.
Zheng HC, Saito H, Masuda S, Wang ZG, Takano Y (2008) Cytoplasmic and nuclear maspin expression in lung carcinomas: an immunohistochemical study using tissue microarrays. Appl Immunohistochem Mol Morphol 16: 459–465.
Zou Z, Anisowicz A, Hendrix MJ, Thor A, Neveu M, Sheng S, Rafidi K, Seftor E, Sager R (1994) Maspin, a serpin with tumour-suppressing activity in human mammary epithelial cells. Science 263: 526–529.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Snoeren, N., Emmink, B., Koerkamp, M. et al. Maspin is a marker for early recurrence in primary stage III and IV colorectal cancer. Br J Cancer 109, 1636–1647 (2013). https://doi.org/10.1038/bjc.2013.489
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.489
Keywords
This article is cited by
-
Serum leucine-rich alpha-2-glycoprotein-1 with fucosylated triantennary N-glycan: a novel colorectal cancer marker
BMC Cancer (2018)
-
The integrated pathway of TGFβ/Snail with TNFα/NFκB may facilitate the tumor-stroma interaction in the EMT process and colorectal cancer prognosis
Scientific Reports (2017)
-
Active nuclear IKK correlates with metastatic risk in cutaneous squamous cell carcinoma
Archives of Dermatological Research (2015)