Abstract
Background:
Prolyl hydroxylation is a post-translational modification that affects the structure, stability and function of proteins including collagen by catalysing hydroxylation of proline to hydroxyproline through action of collagen prolyl hydroxylases3 (C-P3H) and 4 (C-P4H). Three C-P3Hs (nomenclature was amended according to approval by the HGNC symbols and names at http://www.genenames.org/ and Entrez database at http://www.ncbi.nlm.nih.gov/gene) leucineproline-enriched proteoglycan (leprecan) 1 (Lepre1), leprecan-like 1 (Leprel1), leprecan-like 2 (Leprel2) and two paralogs Cartilage-Related Protein (CRTAP) and leprecan-like 4 (Leprel4) are found in humans. The C-P4Hs are tetrameric proteins comprising a variable α subunit, encoded by the P4HA1, P4HA2 and P4HA3 genes and a constant β subunit encoded by P4HB.
Methods:
We used RT–PCR, qPCR, pyrosequencing, methylation-specific PCR, western blotting and immunohistochemistry to investigate expression and regulation of the C-P3H and C-P4H genes in B lymphomas and normal bone marrow.
Results:
C-P3H and C-P4H are downregulated in lymphoma. Down-regulation is associated with methylation in the CpG islands and is detected in almost all common types of B-cell lymphoma, but the CpG islands are unmethylated or methylated at lower levels in DNA isolated from normal bone marrow and lymphoblastoid cell lines. Methylation of multiple C-P3H and C-P4H genes is present in some lymphomas, particularly Burkitt’s lymphoma.
Conclusions:
Methylation of C-P3H and C-P4H is common in B lymphomas and may have utility in differentiating disease subtypes.
Similar content being viewed by others
Main
Non-Hodgkin lymphomas (NHLs) are a heterogeneous group of lymphoid neoplasms that display distinct cytogenetic and molecular genetic features (Piva et al, 2010; Gutierrez-Garcia et al, 2011; Salaverria and Siebert, 2011; Visco et al, 2012). Studies of recurrent chromosome aberrations in NHL have provided significant insight into genetics of lymphomagenesis. Most common translocations are t(14;18) in follicular lymphoma (FL), t(11;14) in mantle cell lymphoma (MCL), t(3;14) in diffuse large B-cell lymphoma (DLBCL) and t(8;14) in Burkitt’s lymphoma (BL) (Ong and Le Beau, 1998; Chaganti et al, 2000). In addition to structural changes, transcriptional silencing is another major mechanism of tumour suppressor gene inactivation in human cancer, including haematological malignancies (Lee et al, 2010; Eberle et al, 2011). A number of mechanistically important genes have been identified as targets for methylation-dependent transcriptional silencing in lymphomas, including the cyclin-dependent kinase inhibitors CDKN2A and CDKN2B, the p53 homologue TP73, the DNA damage-activated protein kinase DAPK and other genes with putative tumour suppressor functions (Belaud-Rotureau et al, 2008; Guan et al, 2010; Murray et al, 2010). Using suppression subtraction PCR, we previously identified the Polo-like kinase PLK2 to be transcriptionally silenced in NHL, with a particularly high frequency of methylation in BL (Syed et al, 2006). However, epigenetic changes in some NHL types, for example FL and MCL remain to be elucidated.
Prolyl hydroxylation is an important post-translational modification that affects the structure and function of proteins. A major substrate of prolyl hydroxylation is collagen and appropriate post-translational processing of collagen requires both C-P3H and C-P4H, respectively. Three C-P3H enzymes have been identified in humans: leucineproline-enriched proteoglycan (leprecan) 1 (Lepre1), leprecan-like 1 (Leprel1) and leprecan-like 2 (Leprel2) (Wassenhove-McCarthy and McCarthy, 1999); Vranka et al, 2004). Prolyl 3-hydroxylation typically occurs in the Gly–3Hyp–4Hyp sequence of collagen and P3H modified residues are abundant in basement membrane collagens (Gryder et al, 1975). Initially, Leprel1 was shown to localise predominantly in the ER and Golgi, but more recently has been detected in tissues rich in basement membranes and shown to participate in the hydroxylation of type IV collagen (Jarnum et al, 2004; Tiainen et al, 2008). We have previously shown that Leprel1 and Leprel2, but not Lepre1, are subject to methylation-dependent transcriptional silencing in breast cancer. Further, whereas methylation in Leprel2 was also present in multiple tumour types, methylation in Leprel1 appeared specific to breast cancer as the CpGisland was unmethylated in several other carcinomas (Shah et al, 2009). The human genome contains two paralogs of the P3H genes, namely Cartilage-Related Protein (CRTAP) and leprecan-like 4 (Leprel4). Although CRTAP is not enzymatically active, it is required in vivo for collagen prolyl 3-hydroxylation (Morello et al, 2006). The biological importance of prolyl hydroxylation in the post-translational modification of collagen is demonstrated by the observation that germ-line mutations in CRTAP and Lepre1 cause recessive forms of osteogenesis imperfecta and loss or decrease of type I collagen prolyl 3-hydroxylation (Cabral et al, 2007; Baldridge et al, 2008) and mutation in Leprel1 is associated with inherited myopia (Mordechai et al, 2011). Prolyl 4-hydroxylation is essential for the biosynthesis of collagens because 4-hydroxyproline residues are critical for stability of the collagen triple helix. The human C-P4Hs are tetrameric isoenzymes composed of two β subunits (encoded by P4HB) and two (catalytic) α subunits. There are three different α subunits encoded by P4HA1, P4HA2 and P4HA3. The β subunit is identical to the protein duslphideisomerase. In contrast to the C-P3Hs, no disorders of collagen synthesis have thus far been associated with heritable change in genes encoding the C-P4Hs. As well as collagen, both C-P3Hs and C-P4Hs may have additional substrates that contain collagen-like domains.
Here, we have investigated expression and regulation of the collagen prolyl hydroxylase genes in lymphoma.
Materials and methods
Cell lines and tumours
NHL cell lines and EBV-immortalised B lymphoblastoid cell lines (LCL) were maintained in RPMI 1640 (Gibco, Invitrogen, Carlsbad, CA, USA) medium containing 10% fetal bovine serum. The following cell lines were used in our study:
Mantle cell lymphoma cell line: JVM2; Transformed FL cell lines: DoHH-1, DoHH-2 and SU-DHL-4. DLBCL cell lines: CRL, DHL-4, DHL-7, LK6; EBV−ve BL: BL2, BL41, DG75, Louckes and Ramos; EBV+ve BL: AW Ramos, Akata 2000, MAK-I, Rael, Namalwa, Raji, Wewack 1, P3HR1 c16 and Jijoye; EBV−ve BL because of loss of EBV: Akata 31; LCLs from peripheral blood: B301, LCL3, X50-7, LCL-K, BM+Akata, PF+B95-8 and CR+B95-8. LCL from cord blood: C2+BL16.
The study received approval from the Ethics Committee of Ioannina University Hospital, Ioannina, Greece. Formalin fixed, paraffin-embedded tissue samples were retrieved from the archives of the Pathology Department, University Hospital of Ioannina, Ioannina, Greece. As control tissues, we used bone marrow (BM) aspirates from individuals with non-malignant disease that contained no neoplastic cells. In all cases, the diagnosis was confirmed following review by an expert haematopathologist. Genomic DNA was isolated by digestion in proteinase K following xylene treatment of the tissue sections. Primary breast carcinomas (from the Tayside Tissue Bank) and breast and ovarian carcinoma cell lines were described previously (Shah et al, 2009).
Methylation reversal
Exponentially growing cells were treated with azacytidine (AZA; Sigma, St Louis, MO, USA) for 5 days. Cells were split every 2 days with the addition of fresh drug. After drug treatment, mRNA was isolated using the RNeasy kit (Qiagen Ltd., West Sussex, UK).
Analysis of gene expression
Total RNA was extracted from exponentially growing cultured cells using RNAzolB or Trizol. In all, 500–1000 ng of total RNA was converted to cDNA and used for analysis of gene expression by RT–PCR. GAPDH was used as a control gene. Primers were designed using Primer3 software. Primer sequences for RT–PCR were

Reactions were resolved on 1.2% agarose gels and visualised on a transilluminator after staining with ethidium bromide. GAPDH was co-amplified as a control gene using primers previously described (Syed et al, 2006).
Total RNA was extracted from 10 μm paraffin sections of archival lymphoma cases using commercial systems (RecoverAll Total Nucleic Acid Isolation kit, Ambion, Carlsbad, CA, USA). For qPCR analysis, 25 μl PCR reactions were performed using 50 ng of cDNA obtained by reverse transcription. Amplification and analysis were done according to the manufacturer’s protocol in 96-well plates in an ABI PRISM 7000 Sequence Detection System (Applied Biosystems, Foster City, CA, USA) and the pre-cast ‘TaqMan Gene Expression Assays’ (Applera, https://products.appliedbiosystems.com/). Quantification of target transcripts was performed in comparison to the reference transcript β2 microglobulin (Hs99999907_m1), using the ‘delta–delta Ct method for comparing relative expression results in real-time PCR as outlined by PE Applied Biosystems. Western blotting for Leprel1 and Leprel2 was done as described previously (Shah et al, 2009).
Methylation analysis
CpG islands are located in the 5′ end of the Lepre1, Leprel1 and Leprel2 genes (Shah et al, 2009). CpG islands were also identified in the 5′sequences of Leprel4, CRTAP and the P4H genes (http://www.ebi.ac.uk/Tools/emboss/cpgplot/index.html). Genomic DNA (1 μg) was modified by sodium bisulphite as described previously (Syed et al, 2006; Shah et al, 2009). Methylation-specific PCR (MSP) primers for the Lepre1, Leprel1 and Leprel2 CpG islands were as described previously (Shah et al, 2009). Primer sequences for MSP analysis of the remaining genes were as follows:

Methylation was also analysed quantitatively by pyrosequencing. Primer sequences were as follows:

PCR conditions were as follows: Initial activation step at 95°C for 10 min, denaturation 95°C for 30 s, annealing 55°C (Lepre1)/52°C (Leprel1)/54°C (Leprel2)/52°C (Leprel4)/51°C (CRTAP)/55°C (P4HA1)/54°C (P4HA2)/49°C (P4HA3) for 30 s for 30 s, extension 72°C for 40 s and final extension 72°C for 7 min. PCR was performed for 40 cycles after which PCR products were electrophoresed through 2% agarose gel, and visualised using a transilluminator. The PCR products were then analysed by pyrosequencing using the Biotage Sample Prep kit (Diatech, Jesi, Italy). Analysis of percentage methylation at each CpG dinucleotide was performed using Pyromark ID Software (Biotage). Placental DNA (Sigma) was used as negative control for methylation (0% average methylation) and a commercial methylated DNA (Millipore, Billerica, MA, USA) was used as positive control (98% average methylation).
Immunohistochemistry
Expression of Leprel1 and Leprel2 was analysed by immunohistochemistry (IHC) performed using standard techniques (with antigen retrieval) on archival lymphoma cases from the archives of the Pathology Department, University Hospital of Ioannina, Ioannina, Greece as described above in cell lines and tumours.
Statistics
We used descriptive statistics and Spearman’s rank correlation test to assess the association between methylation and down-regulation of expression. All statistical analyses were performed using Prism 5 (GraphPad software, Inc., La Jolla, CA, USA).
Results
Methylation-dependent transcriptional silencing of Leprel 4 in NHL
We first analysed expression of Leprel4 in a panel of NHL cell lines. Leprel4 transcript was expressed in LK6, JVM2 and DHL-7 and all LCL analysed but was undetectable in the remaining cell lines (Figure 1A and data not shown). Inspection of Leprel4 revealed the presence of a CpG island in the 5′ sequences of the gene (http://www.ebi.ac.uk/Tools/emboss/cpgplot/index.html), suggesting the possibility that expression is epigenetically regulated and we analysed the CpG island of Leprel4 in each cell line using MSP. In the NHL cell lines lacking expression of Leprel4 mRNA, there was methylation in the CpG island analysed but the CpG island was unmethylated in each cell line which expressed Leprel4 mRNA (Figure 1). To validate the MSP analysis, we further characterised methylation in the Leprel4 CpG island using a quantitative technique, pyrosequencing. Representative pyrograms for genes analysed in this study are shown in Supplementary Figures 1 and 2. There was dense methylation in the CpG island in CRL, DoHH-2 and DHL-4 (all cell lines positive by MSP), but only a low level of methylation in JVM2 (negative by MSP) (Figure 1). Next, we analysed methylation in a series of BL cell lines. There was frequent methylation in the Leprel4 CpGisland, with 13/16 cell lines showing dense methylation by pyrosequencing (Figure 1; Tables 1A and b). Exposure to AZA reactivated expression in MAK-I and AK31, consistent with methylation-dependent silencing (Figure 1). Normal BM control DNA was negative for Leprel4 methylation in 4/4 cases (Tables 1A). Leprel4 mRNA was expressed and the CpG island uniformly unmethylated in breast and ovarian cancer cell lines and primary breast carcinomas (Supplementary Figure 3).
Leprel4 and CRTAP are down-regulated by methylation in NHL. (A) RT–PCR and MSP analysis of expression and CpG island methylation of Leprel4 and CRTAP in NHL cell lines. Expression of Leprel4, CRTAP and GAPDH was determined as described in Materials and Methods. The MSP figure shows U and M specific reactions for the indicated NHL cell lines. Also shown is control methylated (CM) and control unmethylated (CU) genomic DNA modified in parallel with cell line DNA samples. (B) Pyrosequencing analysis of Leprel4 and CRTAP CpG islands in NHL cell lines. The % methylation at each analysed CpG dinucleotide is represented by the intensity of shading in the circles as shown in the figure, together with the mean % methylation for each cell line. (C) Demethylation reactivates expression of Leprel4 and CRTAP. MAK-I and AK31 cells (Leprel4 and CRTAP CpG islands methylated) were grown without AZA (−) or in the presence of AZA for 24 or 48 h as indicated and expression of Leprel4 and CRTAP determined by qPCR.
CRTAP methylation in NHL
These results prompted us to analyse expression of the Leprel4 homologue CRTAP in the same cell line panel. CRTAP mRNA was abundantly expressed in LCL and the majority of NHL cell lines analysed, but at lower levels in Jijoye, AK31, AK2000, Elijah, MAK-I and BL2 (Figure 1). Like Leprel4 the 5′ sequences of CRTAP contain a CpG island. Methylation-specific PCR analysis of the CpG island revealed dense methylation in five of the seven cell lines with down-regulated expression, but methylation was not detected by MSP in BL2 and Jijoye (Figure 1). To validate the MSP analysis, we further characterised methylation in the CRTAP CpG island using pyrosequencing. Methylation in LCL was uniformly low, typically 2–5% at each analysed CG (see for example BM Akata, Figure 1). We did not find evidence of methylation in CRTAP in the other NHL cell lines (DLBCL, FL and MCL) by either MSP or pyrosequencing (Figure 1; Table 1A). Exposure to AZA reactivated expression in MAK-I and AK31, consistent with methylation-dependent silencing (Figure 1). The CRTAP CpG island was unmethylated in breast carcinoma cell lines and ovarian carcinoma cell lines and primary breast carcinomas (Supplementary Figure 3).
Leprel1 and Leprel2 silencing in NHL cell lines
Leprel4 and CRTAP have three paralogs in the human genome Lepre1, Leprel1 and Leprel2, each of which has a CpG island (Shah et al, 2009). RT–PCR revealed expression of Leprel1mRNA in each LCL, with undetectable expression in all BL analysed (Figure 2) and in each of the four non-BL NHL cell lines (data not shown). Western blotting was consistent with RT–PCR (Figure 2). Leprel2 mRNA was undetectable by RT–PCR in the majority of BL cell lines (Figure 2) but was detectable in the four non-BL NHL cell lines (data not shown). AZA reactivated expression of Leprel1 and Leprel2 mRNA in DG75 and Ramos, consistent with methylation-dependent silencing and western blotting confirmed increased Leprel2 in Ramos cells treated with AZA (Figure 2). We performed pyrosequencing in the panel of NHL cell lines to analyse methylation in each CpG island (Figure 2). The Leprel1 CpGisland was methylated to a high degree in all NHL cell lines analysed, whereas Leprel2 CpG island was methylated to a high degree in the majority of BL cell lines but to a lower extent in the non-BL NHL cell lines (Figure 2). Methylation in both CpG islands was low in LCLs (Figure 2). There was an inverse correlation between expression of both Leprel1 and Leprel2 and methylation of their CpG islands (Figure 2). As seen with Leprel4 and CRTAP, there was also a good correlation between methylation detected by pyrosequencing and MSP (Figure 2). Lepre1 was expressed and the CpG island unmethylated in all analysed lymphoma cell lines (Supplementary Figure 4).
Leprel1 and Leprel2 are frequently co-methylated in BL cell lines. (A) RT–PCR analysis of Leprel1 (upper panel), Leprel2 (middle panel) and GAPDH (lower panel) in BL cell lines and LCL. (B) Western blot analysis of Leprel1 and Leprel2 in BL cell lines. Ramos+A indicates Ramos cells treated with AZA. (C) Demethylation reactivates expression of Leprel1 and Leprel2 in BL cell lines. DG75 and Ramos cell lines were grown without AZA (−) or in the presence of AZA (+) for 48 h and expression of Leprel1 and Leprel2 determined by qPCR. (D) Quantitative analysis of methylation in Leprel1 and Leprel2 CpG islands in NHL cell lines. Cell lines were subjected to pyrosequencing analysis as described in Materials and Methods. The number denotes each individual CG dinucleotide in the amplified fragment analysed by pyrosequencing. Five levels of methylation are represented, the level of methylation increasing with increasing intensity of shading in the circles as indicated. (E) MSP analysis of Leprel1 and Leprel2 CpG island methylation in NHL cell lines. MSP was performed as described in Materials and Methods. The figure shows U and M specific MSP reactions for the indicated NHL (upper panels) and BL (lower panels) cell lines. Also shown is control unmethylated (CU) and control methylated (CM) genomic DNA modified in parallel with cell line DNA.
Specific C-P4H genes are silenced by methylation in NHL cell lines
The frequent methylation of the C-P3H genes in NHL prompted us to also analyse expression of the C-P4H genes in the same panel of cell lines. The mRNA of P4HA1, P4HA2 and P4HB was abundantly expressed in each of the LCL tested (Figure 3). P4HA1 and P4HB were expressed in all BL cell lines at levels comparable to the LCL but the mRNA of P4HA2 was down-regulated in many of the BL cell lines (Figure 3). P4HA3 was expressed at low levels in all BL cell lines but was detectable using qPCR and was down-regulated in all BL cell lines analysed (Figure 3). We tested using pyrosequencing whether absence of expression was associated with aberrant methylation in the CpG islands of each C-P4H gene. Methylation was low in the CpG island of each gene in LCLs (Figure 3). Consistent with expression results, the P4HA1 and P4HB CpG islands were uniformly unmethylated. The CpG island of P4HA2 was methylated to a high degree in all of the BL except DG75 and Louckes and there was a good correlation between methylation of and expression of P4HA2 (as assessed by RT–PCR) and methylation (Figure 3). The CpG island of P4HA3 was densely methylated in all cell lines (Figure 3). Together these results imply that expression of P4HA2 and P4HA3 is silenced by methylation in BL cell lines. In the non-BL NHL cell lines, the CpG islands of P4HB, P4HA1 and P4HA2 were unmethylated, but the P4HA3 CpG island was, as in BL, uniformly methylated (Figure 4).
Analysis of P4H genes in BL cell lines. (A) Pyrosequencing analysis of CpG island methylation in C-P4H genes in BL and LCL (B301, LCL-K, LCL3 and CR B95.8). Pyrosequencing was performed as described in Materials and Methods. The degree of methylation at each analysed CG dinucleotide is represented by the intensity of shading in the circles as shown in the figure. (B) RT–PCR analysis of P4H gene expression in BL cell lines. Expression of P4H genes and GAPDH was determined as described in Materials and Methods. (C) qPCR analysis of P4HA3 expression. Expression is shown relative to B301.
Analysis of methylation in C-P4H gene CpG islands in NHL cell lines. The figure shows pyrosequencing analysis of the CpG islands of each P4H gene in NHL cells lines and LCL. The degree of methylation at each analysed CG dinucleotide is represented by the intensity of shading in the circles as indicated in the figure. Also shown in the mean % methylation of the analysed fragment and the result of expression analysis (RT–PCR or qPCR).
Methylation of multiple P3H and P4H genes in clinical cases
Our cell line studies showed that some of the C-P3H and C-P4H genes are down-regulated by methylation in B lymphomas. We therefore analysed methylation in the CpG island of each gene in a clinical series comprising all common types of B lymphomas, including BL, DLBCL, FL, MCL and MZL (Table 1A). As control tissues, we used BM aspirates from individuals (with non-neoplastic diagnoses), confirmed on histopathology to contain no malignant cells. To fully validate our MSP assays, we initially analysed a small series of cases by both pyrosequencing and MSP and this showed an excellent correlation between the two methodologies (Figure 5A). We also confirmed that methylation is associated with transcriptional down-regulation and this was the case (see Figure 5B for representative scatter plots for Leprel1 and Leprel2).
Analysis of methylation in P3H and P4H genes in primary DLBCL. (A) Representative profiles of CpG island methylation for each C-P3H and C-P4H gene in clinical DLBCL cases. Pyrosequencing was performed as described in Materials and Methods. Five levels of methylation are represented, the level of methylation increasing with increasing intensity of shading in the circles as shown in the figure. The result of simultaneous MSP analysis is indicated. (B) CpG island methylation correlates with down-regulation of expression in clinical cases of DLBCL. Scatter plots are shown of mRNA vs mean % CG methylation for Leprel1 and Leprel2 in DLBCL.
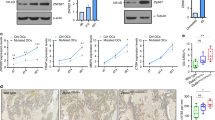
We then proceeded to analyse the clinical cases using the validated MSP assays (for representative assays see Figure 6; for summary of data see Table 1A and Supplementary Figure 5). Methylation was not detected in any of the nine analysed genes in the normal BM controls. Leprel4 was methylated exclusively in high-grade lymphomas, the highest frequency being in BL. Similarly, CRTAP was methylated predominantly in high-grade lymphomas with the highest frequency in DLBCL, although a single case of MCL showed methylation. Leprel1 was clearly the most frequently methylated of the C-P3H genes, with methylation in the Leprel1 CpG island detected in 86/100 (86%) of DLBCL and 9/11 (82%) of FL. Leprel2 methylation was infrequent and was detected only in DLBCL (at low frequency) and in BL (at high frequency) but not in other lymphoma types. Although methylation in the CpG island of Lepre1 was not detected in NHL cell lines, we nonetheless observed methylation in all lymphoma types, albeit at low frequencies, in the clinical series (Table 1A). In the case of the P4H genes, we did not detect methylation in P4HA1 or P4HB in any lymphoma analysed, consistent with analysis of cell lines. P4HA2 was methylated predominantly in BL, whereas P4HA3 was methylated to a high degree in all types of NHL and BL (Table 1A). To seek evidence that CpG island methylation does indeed result in reduced protein expression, we performed IHC for Leprel1 and Leprel2 (Figure 6B). In these studies, we observed down-regulation of both proteins in all common B lymphoma types, consistent with qPCR and methylation analyses, confirming that transcriptional silencing is associated with reduced or absent protein expression in lymphoma.
Analysis of the CpG islands in C-P3H and C-P4H genes in clinical lymphoma cases. (A) Examples of MSP analyses for C-P3H and C-P4H genes. For Leprel4, CRTAP, Leprel1 and Leprel2 reactions with unmethylated (U) and methylated (M) primer pairs are shown. For P4HA1, P4HA2 and P4HA3 cases are analysed with M primers only. For all genes, specificity and sensitivity of the MSP is demonstrated by the control DNA amplifications. Abbreviations: CM=control methylated DNA; CU=control unmethylated DNA. (B) Representativeimmunohistochemical analysis of Leprel1 and Leprel2 in B lymphomas. Cases with and without methylation in the respective CpG islands are shown.
Analysis of C-P3H and C-P4H genes discriminates DLBCL from BL
Inspection of the combined data set (Table 1A; Supplementary Figure 5) showed that Leprel1 and P4HA3 are methylated at high frequency in the majority of high-grade NHL, both in cell lines and in clinical cases of lymphoma, whereas P4HA2 and Leprel2 are methylated predominantly in BL and less frequently in DLBCL. We constructed ROC curves to address the possibility that methylation analysis of specific C-P3H and C-P4H genes facilitates discrimination between (i) NHL and LCL (ii) BL and DLBCL. In cell lines, methylation of Leprel1, Leprel2 and P4HA3 clearly discriminated between NHL and LCL with high sensitivity and specificity but for P4HA2 the sensitivity was lower, consistent with lower frequency of methylation in non-BL lymphomas (Figure 7). In clinical lymphomas, methylation of Leprel2 and P4HA2 was a sensitive and specific discriminator between BL and DLBCL (Figure 7). We further assessed MSP results to determine whether non-quantitative analysis of methylation can also distinguish BL from DLBCL. Using MSP, sensitivity and specificity for both Leprel2 and P4HA2 were similar to values obtained from pyrosequencing (Table 1B). Together, these results from two independent techniques suggest that methylation profiling of the CpG islands of Leprel2 and P4HA2 BL and DLBCL can be useful in distinguishing between BL and DLBCL. Finally, to further extend these observations, we asked whether immunohistochemical analysis of Leprel2 discriminates DLBCL from BL. Consistent with methylation analysis, expression of Leprel2 was absent or greatly down-regulated in BL at a significantly higher frequency than in DLBC (P<0.05).
Discussion
We show that expression of the C-P3H and C-P4H collagen prolyl hydroxylases is epigenetically downregulated in NHL cell lines and primary lymphomas. Our data is the first demonstration of epigenetic silencing of the C-P3H and C-P4H genes in lymphoid neoplasia and to the best of our knowledge the first study describing epigenetic changes in the C-P4H genes in human cancer. Our study of the C-P3H and C-P4H genes was prompted by initial identification of Leprel4 as a novel epigenetically regulated gene in lymphoma. This led us to investigate four paralogous C-P3H genes in the human genome (CRTAP, Lepre1, Leprel1 and Leprel2) and the C-P4H genes P4HB, P4HA1, P4HA2 and P4HA3, all of which encode proteins with functions in collagen biosynthesis. We demonstrate methylation-dependent transcriptional silencing of specific members of each gene family in B-cell lymphoma cell lines and primary lymphomas.
In the case of the C-P3H family members, Leprel1 was silenced at high frequency in all lymphoma types, with lower methylation frequencies in the CpG islands of each of the other C-P3H genes aside from BL in which we observed simultaneous silencing of Leprel1, Leprel2, Leprel4 and CRTAP in a high proportion of cases (both sporadic and HIV-associated). In previous work, we showed that Leprel2 is silenced in numerous epithelial tumour types, but silencing of Leprel1 was restricted to breast cancer with a strong preference for oestrogen receptor positive carcinomas (Shah et al, 2009). We did not detect methylation in the CpG islands of Leprel4 or CRTAP in breast and ovarian cancer cell lines or in primary breast cancers, suggesting that methylation in these genes is specific for haematological neoplasia. It is striking therefore that Leprel1 is transcriptionally silenced in the majority of B lymphomas irrespective of subtype. Imuunohistochemistry confirmed down-regulation of Leprel1 and Leprel2 (albeit with different frequencies) in clinical lymphoma cases.
In the case of the C-P4H family, we detected CpG island methylation only in P4HA2 and P4HA3, methylation never being seen either in P4HA1 or in P4HB (encoding the β subunit of the C-P4H tetramer). Of note, whereas the P4HA3 CpG island was methylated in a high proportion of lymphomas of all types, that of P4HA2 was methylated predominantly in BL and there was simultaneous methylation of P4HA2 and P4HA3 in many cases of BL. This parallels the situation with the C-P3H genes with Leprel1 and Leprel2, which are co-silenced at high frequency in BL. It remains to be determined whether this reflects differences between P4HA2 and P4HA3 in terms of substrate specificity or affinity.
Previous studies have linked germ-line mutations in Lepre1 and CRTAP to rare forms of the collagen disorder osteogenesis imperfecta (Morello et al, 2006; Cabral et al, 2007; Baldridge et al, 2008), mutations resulting in reduced 3-hydroxylation of the α chain of type I collagen. It is an interesting hypothesis that transcriptional silencing of the C-P3H and C-P4H genes may contribute to lymphomagenesis as a result of aberrant collagen biosynthesis and/or post-translational modification causing basement membrane malformation or other collagen abnormalities which promote neoplasia. Alternatively, failure of prolyl hydroxylation of one or more of the other identified substrates for these proteins could contribute to neoplasia in cells lacking specific C-P3H and C-P4H. A third possibility is that P3H and P4H have direct growth inhibitory properties. This has been directly demonstrated for Lepre1, Leprel1 and Leprel2 (Kaul et al, 2000; Shah et al, 2009). Further, mice lacking both CRTAP alleles exhibit increased proliferation in some tissues, also consistent with an anti-proliferative function (Baldridge et al, 2010). Resolution of these hypotheses will require detailed mechanistic studies.
Even with modern histopathology, immunoprofiling and molecular genetics, it is on occasion challenging to differentiate between BL and DLBCL (de Leval and Hasserjian, 2009). This is of major clinical importance because first-line chemotherapy for the two lymphoma subtypes is very different. Differences in methylation frequency between BL and DLBCL, especially in Leprel2 and P4HA2, led us to address the possible utility of these genes as biomarkers to differentiate between the lymphoma types. We show that analysis of CpG island methylation in both Leprel2 and P4HA2 discriminates between BL and DLBCL with high sensitivity and specificity. In preliminary studies, we showed using IHC that analysis of Leprel2 expression in BL and DLBCL may also have biomarker utility. Clearly, larger numbers of cases will require analysis in order to independently verify these observations.
In conclusion, we report the identification of C-P3H and C-P4H as novel transcriptionally silenced gene families in B lymphomas. Additional studies are required in order to validate the initial findings presented in this report and to further elucidate the possible role of transcriptional silencing of the C-P3H and C-P4H genes in lymphomagenesis.
Change history
04 October 2012
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Baldridge D, Lennington J, Weis M, Homan EP, Jiang MM, Munivez E, Keene DR, Hogue WR, Pyott S, Byers PH, Krakow D, Cohn DH, Eyre DR, Lee B, Morello R (2010) Generalized connective tissue disease in Crtap−/− mouse. PloS One 5 (5): e10560
Baldridge D, Schwarze U, Morello R, Lennington J, Bertin TK, Pace JM, Pepin MG, Weis M, Eyre DR, Walsh J, Lambert D, Green A, Robinson H, Michelson M, Houge G, Lindman C, Martin J, Ward J, Lemyre E, Mitchell JJ, Krakow D, Rimoin DL, Cohn DH, Byers PH, Lee B (2008) CRTAP and LEPRE1 mutations in recessive osteogenesis imperfecta. Human Mutation 29 (12): 1435–1442
Belaud-Rotureau MA, Marietta V, Vergier B, Mainhaguiet G, Turmo M, Idrissi Y, Ferrer J, Beylot-Barry M, Dubus P, Merlio JP (2008) Inactivation of p16INK4a/CDKN2A gene may be a diagnostic feature of large B cell lymphoma leg type among cutaneous B cell lymphomas. Virchows Archiv Int J Pathol 452 (6): 607–620
Cabral WA, Chang W, Barnes AM, Weis M, Scott MA, Leikin S, Makareeva E, Kuznetsova NV, Rosenbaum KN, Tifft CJ, Bulas DI, Kozma C, Smith PA, Eyre DR, Marini JC (2007) Prolyl 3-hydroxylase 1 deficiency causes a recessive metabolic bone disorder resembling lethal/severe osteogenesis imperfecta. Nat Genet 39 (3): 359–365
Chaganti RS, Nanjangud G, Schmidt H, Teruya-Feldstein J (2000) Recurring chromosomal abnormalities in non-Hodgkin’s lymphoma: biologic and clinical significance. Sem Hematol 37 (4): 396–411
de Leval L, Hasserjian RP (2009) Diffuse large B-cell lymphomas and burkitt lymphoma. Hem-oncol Clin N Am 23 (4): 791–827
Eberle FC, Rodriguez-Canales J, Wei L, Hanson JC, Killian JK, Sun HW, Adams LG, Hewitt SM, Wilson WH, Pittaluga S, Meltzer PS, Staudt LM, Emmert-Buck MR, Jaffe ES (2011) Methylation profiling of mediastinal gray zone lymphoma reveals a distinctive signature with elements shared by classical Hodgkin’s lymphoma and primary mediastinal large B-cell lymphoma. Haematologica 96 (4): 558–566
Gryder RM, Lamon M, Adams E (1975) Sequence position of 3-hydroxyproline in basement membrane collagen. Isolation of glycyl-3-hydroxyprolyl-4-hydroxyproline from swine kidney. J Biol Chem 250 (7): 2470–2474
Guan H, Xie L, Leithauser F, Flossbach L, Moller P, Wirth T, Ushmorov A (2010) KLF4 is a tumor suppressor in B-cell non-Hodgkin lymphoma and in classic Hodgkin lymphoma. Blood 116 (9): 1469–1478
Gutierrez-Garcia G, Cardesa-Salzmann T, Climent F, Gonzalez-Barca E, Mercadal S, Mate JL, Sancho JM, Arenillas L, Serrano S, Escoda L, Martinez S, Valera A, Martinez A, Jares P, Pinyol M, Garcia-Herrera A, Martinez-Trillos A, Gine E, Villamor N, Campo E, Colomo L, Lopez-Guillermo A (2011) Gene-expression profiling and not immunophenotypic algorithms predicts prognosis in patients with diffuse large B-cell lymphoma treated with immunochemotherapy. Blood 117 (18): 4836–4843
Jarnum S, Kjellman C, Darabi A, Nilsson I, Edvardsen K, Aman P (2004) LEPREL1, a novel ER and Golgi resident member of the Leprecan family. Biochem Biophys Res Commun 317 (2): 342–351
Kaul SC, Sugihara T, Yoshida A, Nomura H, Wadhwa R (2000) Gros1, a potential growth suppressor on chromosome 1: its identity to basement membrane-associated proteoglycan, leprecan. Oncogene 19 (32): 3576–3583
Lee S, Syed N, Taylor J, Smith P, Griffin B, Baens M, Bai M, Bourantas K, Stebbing J, Naresh K, Nelson M, Tuthill M, Bower M, Hatzimichael E, Crook T (2010) DUSP16 is an epigenetically regulated determinant of JNK signalling in Burkitt’s lymphoma. Br J Cancer 103 (2): 265–274
Mordechai S, Gradstein L, Pasanen A, Ofir R, El Amour K, Levy J, Belfair N, Lifshitz T, Joshua S, Narkis G, Elbedour K, Myllyharju J, Birk OS (2011) High myopia caused by a mutation in LEPREL1, encoding prolyl 3-hydroxylase 2. Am J Human Genet 89 (3): 438–445
Morello R, Bertin TK, Chen Y, Hicks J, Tonachini L, Monticone M, Castagnola P, Rauch F, Glorieux FH, Vranka J, Bachinger HP, Pace JM, Schwarze U, Byers PH, Weis M, Fernandes RJ, Eyre DR, Yao Z, Boyce BF, Lee B (2006) CRTAP is required for prolyl 3-hydroxylation and mutations cause recessive osteogenesis imperfecta. Cell 127 (2): 291–304
Murray PG, Fan Y, Davies G, Ying J, Geng H, Ng KM, Li H, Gao Z, Wei W, Bose S, Anderton J, Kapatai G, Reynolds G, Ito A, Marafioti T, Woodman CB, Ambinder R, Tao Q (2010) Epigenetic silencing of a proapoptotic cell adhesion molecule, the immunoglobulin superfamily member IGSF4, by promoter CpG methylation protects Hodgkin lymphoma cells from apoptosis. Am J Pathol 177 (3): 1480–1490
Ong ST, Le Beau MM (1998) Chromosomal abnormalities and molecular genetics of non-Hodgkin’s lymphoma. Sem Oncol 25 (4): 447–460
Piva R, Agnelli L, Pellegrino E, Todoerti K, Grosso V, Tamagno I, Fornari A, Martinoglio B, Medico E, Zamo A, Facchetti F, Ponzoni M, Geissinger E, Rosenwald A, Muller-Hermelink HK, De Wolf-Peeters C, Piccaluga PP, Pileri S, Neri A, Inghirami G (2010) Gene expression profiling uncovers molecular classifiers for the recognition of anaplastic large-cell lymphoma within peripheral T-cell neoplasms. J Clin Oncol 28 (9): 1583–1590
Salaverria I, Siebert R (2011) The gray zone between Burkitt’s lymphoma and diffuse large B-cell lymphoma from a genetics perspective. J Clin Oncol 29 (14): 1835–1843
Shah R, Smith P, Purdie C, Quinlan P, Baker L, Aman P, Thompson AM, Crook T (2009) The prolyl 3-hydroxylases P3H2 and P3H3 are novel targets for epigenetic silencing in breast cancer. Br J Cancer 100 (10): 1687–1696
Syed N, Smith P, Sullivan A, Spender LC, Dyer M, Karran L, O’Nions J, Allday M, Hoffmann I, Crawford D, Griffin B, Farrell PJ, Crook T (2006) Transcriptional silencing of Polo-like kinase 2 (SNK/PLK2) is a frequent event in B-cell malignancies. Blood 107 (1): 250–256
Tiainen P, Pasanen A, Sormunen R, Myllyharju J (2008) Characterization of recombinant human prolyl 3-hydroxylase isoenzyme 2, an enzyme modifying the basement membrane collagen IV. J Biol Chem 283 (28): 19432–19439
Visco C, Li Y, Xu-Monette ZY, Miranda RN, Green TM, Li Y, Tzankov A, Wen W, Liu WM, Kahl BS, d’Amore ES, Montes-Moreno S, Dybkaer K, Chiu A, Tam W, Orazi A, Zu Y, Bhagat G, Winter JN, Wang HY, O’Neill S, Dunphy CH, Hsi ED, Zhao XF, Go RS, Choi WW, Zhou F, Czader M, Tong J, Zhao X, van Krieken JH, Huang Q, Ai W, Etzell J, Ponzoni M, Ferreri AJ, Piris MA, Moller MB, Bueso-Ramos CE, Medeiros LJ, Wu L, Young KH (2012) Comprehensive gene expression profiling and immunohistochemical studies support application of immunophenotypic algorithm for molecular subtype classification in diffuse large B-cell lymphoma: A report from the International DLBCL Rituximab-CHOP consortium program study. Leukemia 29: 2103–2113
Vranka JA, Sakai LY, Bachinger HP (2004) Prolyl 3-hydroxylase 1, enzyme characterization and identification of a novel family of enzymes. J Biol Chem 279 (22): 23615–23621
Wassenhove-McCarthy DJ, McCarthy KJ (1999) Molecular characterization of a novel basement membrane-associated proteoglycan, leprecan. J Biol Chem 274 (35): 25004–25017
Acknowledgements
Breast Cancer Research Scotland and the Tayside Tissue Bank supported parts of the work reported herein. Eleftheria Hatzimichael received funding as a scholar from the Hellenic Society of Haematology Foundation. Tim Crook is a Scottish Senior Clinical Fellow in Medical Oncology.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies the paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Hatzimichael, E., Lo Nigro, C., Lattanzio, L. et al. The collagen prolyl hydroxylases are novel transcriptionally silenced genes in lymphoma. Br J Cancer 107, 1423–1432 (2012). https://doi.org/10.1038/bjc.2012.380
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2012.380
Keywords
This article is cited by
-
Network pharmacological analysis of corosolic acid reveals P4HA2 inhibits hepatocellular carcinoma progression
BMC Complementary Medicine and Therapies (2023)
-
TGF-β-dependent reprogramming of amino acid metabolism induces epithelial–mesenchymal transition in non-small cell lung cancers
Communications Biology (2021)
-
EZH2 oncogenic mutations drive epigenetic, transcriptional, and structural changes within chromatin domains
Nature Genetics (2019)
-
Protein arginine methyltransferase 5 regulates multiple signaling pathways to promote lung cancer cell proliferation
BMC Cancer (2016)