Abstract
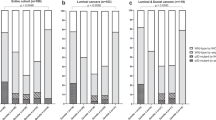
Constant denaturant gel electrophoresis (CDGE) was used to screen 179 breast carcinomas for mutations in the conserved regions of the TP53 gene (exons 5 through 8). Mutations were found in 35 of 163 primary tumours (21%) and in 5 of 16 metastases (31%) and resided predominantly in exon 7. The majority of the mutations were G:C-->A:T transitions. Immunohistochemistry demonstrated nuclear accumulation of p53 protein in 35 of 162 primary tumours (22%) and in four of 15 metastases (27%). TP53 mutation was strongly associated with nuclear accumulation of p53 protein. In total 42 of 163 primary tumours (26%) and 5 of 16 metastases (31%) were demonstrated to contain TP53 alterations (mutation and/or nuclear protein accumulation). TP53 alteration in primary tumour was significantly associated with the following parameters: positive node status, T status > 1, negative oestrogen receptor status, negative progesterone receptor status, presence of ERBB2 gene amplification, and invasive ductal histology. Furthermore, there were statistically significant associations, independent of other prognostic factors, between TP53 alterations in primary tumour and disease-free and overall survival.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Andersen, T., Holm, R., Nesland, J. et al. Prognostic significance of TP53 alterations in breast carcinoma. Br J Cancer 68, 540–548 (1993). https://doi.org/10.1038/bjc.1993.383
Issue Date:
DOI: https://doi.org/10.1038/bjc.1993.383
This article is cited by
-
Breast cancer genomes from CHEK2 c.1100delC mutation carriers lack somatic TP53 mutations and display a unique structural variant size distribution profile
Breast Cancer Research (2023)
-
Mutational Status of CDKN2A and TP53 Genes in Laryngeal Squamous Cell Carcinoma
Pathology & Oncology Research (2015)
-
The Clinical Value of LRRC3B Gene Expression and Promoter Hypermethylation in Breast Carcinomas
Cell Biochemistry and Biophysics (2014)
-
Number of rare germline CNVs and TP53 mutation types
Orphanet Journal of Rare Diseases (2012)
-
p53 mutations in classic and pleomorphic invasive lobular carcinoma of the breast
Cellular Oncology (2012)