Abstract
KIT exon 17 mutation is a poor prognostic factor in core-binding factor acute myeloid leukemia. However, the mutation detection method used for risk assessment is not assigned. It is necessary to verify the analytical and clinical performance before applying new methods. Herein, we firstly applied a highly sensitive allele-specific, real-time quantitative PCR (AS-qPCR) assay to analyze KIT mutations, which demonstrated excellent sensitivity and specificity. Much higher incidence of KIT mutations (62.2%, 69/111) and prevalence of multiple mutations (43.5%, 30/69) were observed using AS-qPCR, which meant the existence of multiple KIT mutant subclones. The relative KIT mutant level was variable (median, 0.3 per control allele 100 copies, 0.002–532.7) and was divided into two groups: high (⩾10, n=26) and low (<10) mutant level. Interestingly, rather than mutation positivity, mutant level was found to be associated with clinical outcome. High mutant level showed significantly inferior overall survival (P=0.005) and event-free survival (P=0.03), whereas low level did not influence the prognosis. The follow-up data showed that the mutant level were along with fusion transcripts in the majority (n=29), but moved separately in some cases, including the loss of mutations (n=5) and selective proliferation of minor clones (n=2) at relapse. This study highlighted that the KIT mutation should be analyzed using sensitive and quantitative techniques and set a cutoff level for identifying the risk group.
Similar content being viewed by others
Introduction
Core-binding factor acute myeloid leukemia (CBF AML) constitutes ~17% of AML1 and has a favorable prognosis because outcome is better than for other subtypes of AML. However, CBF AML is reclassified as the intermediate-risk category when accompanied with the KIT mutation. Various methods have been used to detect common mutations in the KIT gene including direct sequencing, PCR fragment analysis, denaturing high-performance liquid chromatography, pyrosequencing, melting-curve analysis and allele-specific PCR.2,3,4,5,6,7 KIT mutations are reported to occur in CBF AML patients at frequencies ranging from 17 to 46%.2, 3, 4, 5, 6, 7, 8, 9 A mutation-based PCR using a quenching probe provides greater sensitivity than direct sequencing does for detecting KIT mutations; detection rate of KIT mutations at initial presentation is reportedly boosted from 19.2 to 46.2%, and it is important for predicting prognosis for t(8;21).6 It is expected that sensitive detection methods result in an increased positive rate. The broad range of mutation positivity in previous reports reflects the different methods used to detect the mutations. Because KIT mutations already have relevance risk stratification of CBF AML, there is a concern regarding whether or not the low allele level detected by a highly sensitive technique has the same clinical significance as KIT mutations detected by previously used methods. If the prognosis depends on the mutant level, and not only on the presence or absence of mutations, the KIT mutation should be analyzed using sensitive and quantitative techniques to set a cutoff level to identify the real risk group.
Treatment responsiveness of leukemic cells with KIT mutations is another important point. Follow-up after treatments for CBF AML is accompanied by analyses of hematologic findings and monitoring of fusion transcripts such as RUNX1-RUNX1T1 and CBFB-MYH11.10, 11, 12 It is not recommended to use the KIT mutation as a monitoring marker because KIT mutations are heterogeneous and function as a second hit in accordance with fusion transcripts. However, the observation of the changes in KIT mutations after treatment is necessary for understanding the biological behavior of the leukemic cells with or without KIT mutations, especially in minor clones, and for preparing for advancements in therapeutic approaches.
In this study, we developed an allele-specific, real-time quantitative PCR (AS-qPCR) assay and evaluated its performance for detection of KIT mutations in CBF AML. We further tried to analyze the clinical significance of KIT mutations in CBF AML according to allele level and to understand the changes of mutations after treatment.
Patients and methods
Patients
A total of 111 CBF AML patients diagnosed between April 2009 and July 2013 were included in the study. Diagnosis of CBF AML was determined according to the 2008 World Health Organization classifications. We confirmed all CBF AML with chromosomal analysis and additional reverse transcriptase-PCR (RT-PCR) and/or qPCR. All patients were treated with remission induction chemotherapy according to our standard protocol, consisting of ‘3+7’ idarubicin plus BHAC (N4-behenoyl-1-β-d-arabinofuranosylcytosine) or cytosine arabinoside. After complete remission (CR) achievement, patients underwent one or two more cycles of standard consolidation chemotherapy that consisted of ‘3+5’ mitoxantrone (12 mg/m2 intravenously) plus intermediate dose cytosine arabinoside (1.0 g/m2 intravenously) or idarubicin (12 mg/m2) plus intermediate dose cytosine arabinoside alternatively up to two cycles. If an available donor was found during consolidation, patients were treated with allogeneic hematopoietic stem cell transplantation (HSCT). If the patient did not have an available allogeneic donor, we gave an autologous HSCT. There were 68 male and 43 female subjects. The median age was 44 years (range, 18–72 years) and median follow-up was 19.8 months from initial presentation (range, 0–63.2 months). Of these, 74 had t(8;21) AML and 37 had inv(16) AML. Paired DNA samples at disease relapse or follow-up bone marrow (BM) samples after treatment were available from 36 patients. According to the Declaration of Helsinki, all patients in this study provided written informed consent for genetic analyses, and the study protocol was approved by the Institutional Review Board of Seoul St Mary’s Hospital, The Catholic University of Korea.
Screening test for detection of KIT mutations
All DNA samples extracted from the BM were initially screened for D816V, D816H, D816Y and N822K mutations of KIT by allele-specific real-time PCR assay (Real-Q KIT Screening Kit; BioSewoom, Seoul, Korea). Positive samples were genotyped using the Real-Q KIT Genotyping Kit (BioSewoom) for discrimination of each mutation.
AS-qPCR for KIT mutations
We have developed a novel AS-qPCR assay for detecting KIT exon 17 mutations. Primers and probes for AS-qPCR of KIT exon 17 mutations were designed with a common reverse primer and a single probe for D816V, D816H, D816Y and N822K mutation reactions. The forward primer specific for the mutant allele was designed using the amplification refractory mutation system principle.13 The minor groove-binding non fluorescent quencher probe for KIT D816V, D816H, D816Y and N822K was fluorescently labeled with VIC on the 5′ end. An AS-qPCR was performed in the ABI 7500 (Applied Biosystems, Foster City, CA, USA) with an initial step at 50 °C for 2 min and 95 °C for 10 min, followed by 40 cycles of 15 s at 95 °C and 45 s at 63 °C. Each reaction mix contained a coamplified internal reference primer, which amplified a mutation-unspecific region of the KIT gene (exon 2). All DNA samples were analyzed in duplicate and negative and positive control samples were used for each run.
Standard curves were generated from calibrators constructed by cloning the target DNA into plasmids. Calibrators containing 102, 103, 104, 105 and 106 gene copies were prepared for D816V, D816H, N822K and control alleles (103, 104, 105, 106 and 107 gene copies for D816Y). For each run, mutant and control allele plasmid DNA-based standard curves were generated, and actual copy numbers were determined for both the mutant allele and the control allele. The relative mutant level was expressed as a ratio of the two populations (mutant allele copies/100 control allele copies).
Analytical performance
Specificity was evaluated using DNA samples derived from 100 normal donors14 and no false positivity was identified. Standard material was produced to evaluate the analytical performance of AS-qPCR of KIT mutation using a cloned plasmid DNA containing PCR amplification product from patient DNA bearing KIT exon 17 mutations. Analytical sensitivity was defined as the minimal proportion of mutant DNA that could consistently be detected with a 95% confidence interval using 10-fold serial dilutions of mutant plasmid DNA. Ten independent runs with triplicates of each sample were performed.The limit of detection was calculated by probit analysis at 95% detection level; D816V, 43.4 copies per μl; D816H, 13.8 copies per μl; D816Y, 168.9 copies per μl; and N822K, 1.7 copies per μl. A strict linear correlation was observed in dilutions ranging from 102 to 106 copies for D816V, D816H and N822K, and from 103 to 107 copies for D816Y. The correlation coefficient (R2) in this range was close to 1 (range 0.998–1.0; Supplementary Figure S1).
Clinical validation of AS-qPCR
The results from AS-qPCR assay was validated by melting-cure analysis, which is a sensitive method to detect KIT mutations with 5–10% sensitivity.15 Discrepant results were observed in samples with low mutant allele level; AS-qPCR-positive/melting-curve analysis-negative. In those samples, pyroseqeuncing and mutant enrichment with 3′-modified oligonucleotides PCR (MEMO-PCR) were used to confirm the KIT mutations.
Melting-curve analysis for the detection of exon 17 mutations
Melting-curve analysis based on real-time PCR was capable of detecting four mutation types (D816V, D816Y, D816H and N822K) using EvaGreen as the intercalating dye. The PCR program consisted of an initial preheating at 95 °C for 10 min to activate the Taq DNA polymerase, followed by 40 amplification cycles that consisted of a denaturation step at 95 °C for 15 s and an annealing step at 66 °C for 45 s. After PCR, melting-curve analysis was started at 60 °C up to 95 °C with slope of 0.1 °C/s and continuous detection of fluorescence at channel 530. Melting-curve analyses performed after PCR showed a peak at 83±1 °C corresponding to the internal control, and peaks at 74±1 and 76.5±1 °C corresponding to D816 and N822 mutations, respectively.
Pyrosequencing and MEMO-PCR
Pyrosequencing was performed using biotinylated PCR products. All primers were designed using the Pyrosequencing Assay Design Software (Biotage AB and Biosystems, Uppsala, Sweden; Supplementary Table S1). Products were sequenced on a PyroMark ID system (Biotage AB and Biosystems) using PyroMark Q96 reagents and protocols (Qiagen).
MEMO-PCR amplification was performed using two generic primers and one additional primer to enrich the mutant alleles of the KIT gene.16 The additional primer was designed to include the KIT mutation site to overlap with generic sense primer. The sequencing was performed on an ABI 3730xl DNA Analyzer (Applied Biosystems) with BigDye Terminator v.3.1 (Applied Biosystems) and generic primer.
Cloning
To investigate whether the two KIT mutations (D816 and N822) were on the same or different leukemic cells, we adopted an allele-specific cloning and sequencing approach in two patients carrying double KIT exon 17 mutations. PCR amplification of exon 17 containing both mutation sites was performed with specific oligonucleotide primers (Supplementary Table S1).17 The cloning was performed with the use of the TOPO-TA Cloning Kit (Invitrogen, Carlsbad, CA, USA) and direct sequencing was performed on plasmid DNA obtained from at least 20 isolated clones for each sample.
Other molecular analyses
Detection of KIT exon 8 mutations was performed by direct sequencing.4 Internal tandem duplication of FLT3 gene (FLT3-ITD) and mutations in tyrosine kinase domain (FLT3-TKD) were determined using conventional PCR according to the previous study.18 NPM1 exon 12 mutations were determined by Sanger sequencing. RUNX1-RUNX1T1 and CBFβ-MYH11 were detected by the multiplex RT-PCR assay and quantified by RT-qPCR. Results were expressed as a (fusiongene/Abelson gene (ABL1)) transcript ratio.
Definition of clinical end points
Achievement of CR was defined as a normocellular BM containing <5% leukemic blasts, and no evidence for circulating blasts and/or extramedullary leukemia (neutrophils 1500/μl and platelets >100 000/μl in peripheral blood). Relapse was defined as reoccurrence of >5% of leukemic blasts in BM, reappearance of circulating blasts or the development of extramedullary leukemia.19 Overall survival (OS) was defined as the time from diagnosis to death or date last known alive. Event-free survival (EFS) was defined as the time from diagnosis to relapse for patients who experienced a relapse, time to death for non-relapsed patients who did not survive or time to last follow-up for surviving patients who did not experience a relapse. Cumulative incidence of relapse was the incidence of relapse after CR with death in CR as a competing risk.
Statistical analyses
Differences in clinical variables according to mutation status were investigated using Fisher’s exact test for categorical variables and the Mann–Whitney U-test for continuous variables. Survival functions were calculated using the Kaplan–Meier survival analysis and the differences in survival curves were compared using a two-sided log-rank test. Cox proportional hazard models were used to estimate hazard ratios (HRs) for univariate and multivariate analyses for OS, EFS and cumulative incidence of relapse. All statistical analyses were performed using SPSS 12.0.1 for Windows (SPSS, Chicago, IL, USA) and EZR (Saitama Medical Center, Jichi Medical University, Saitama, Japan).20 A two-sided P-value <0.05 indicated statistical significance.
Results
Clinical utility of initial diagnosis: incidence and type of KIT exon 17 mutations
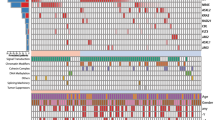
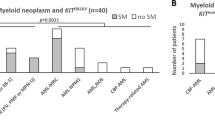
The AS-qPCR proved to be highly sensitive for detecting KIT exon 17 mutations of at least 200 copies per μl. We identified 69 cases (62.2%) with KIT mutations in this study, a higher incidence compared with what was reported in previous studies.2, 17 Among the patients with KIT mutations, 30 (43.5%) had multiple mutations according to this highly sensitive method: 23 with double mutations and 7 patients with triple mutations; this finding is novel (Figure 1).
In patients with t(8;21) AML, 63.5% (47/74) had KIT exon 17 mutations. Of the 47 patients, 28 had single mutation: D816 in 16 (D816V in 8, D816Y in 5 and D816H in 3) and N822K in 12 cases. Nineteen patients had multiple mutations: 16 patients with double mutations and 3 patients with triple mutations. In 37 patients with inv(16) AML, 22 (59.5%) had KIT exon 17 mutations. Of the 22 patients, 11 had single mutation: D816 in 9 (D816V in 6, D816Y in 2 and D816H in 1) and N822K in 2 cases. Eleven patients possessed multiple mutations: seven patients with double mutations and four patients with triple mutations.
The relative mutant level was also obtained using AS-qPCR, which was expressed as mutant allele copies per 100 KIT control allele copies. In cases with more than one KIT exon 17 mutations, all the calculated relative mutant levels were summed. The KIT mutant allele level was variable (median, 0.3; range, 0.002–532.7). The distribution of KIT allele level in all patients with KIT mutations showed a double peak as the fiducial point of 10 (Figure 2a). D816 mutations revealed higher mutant allele level compared with N822K (median, 0.7; range, 0.002–532.7 vs 0.1; 0.006–43.53, P=0.016; Figure 2b). The mutant allele levels in patients with t(8;21) AML were not different from those in patients with inv(16) AML (median, 0.5; range, 0.002–532.7 vs 0.1; 0.002–293.0, P=0.243; Figure 2c). Patients with multiple mutations did not reveal a higher mutant allele level when compared with those with a single mutation (median, 0.2; range, 0.002–490.9 vs 2.2; 0.002–532.7, P=0.077; Figure 2d). Eleven (36.7%) of 30 patients with multiple mutations had a single dominant mutation (allele level ⩾10) and other minor mutation(s), whereas 19 patients (63.3%) harbored mutations with a low allele level, <10. Cloning analysis in cases with double mutations revealed that D816V and N822K were on different alleles: 5 D816V clones and 6 N822K clones, respectively, in one, 10 D816H clones, 6 N822K clones, respectively, in another, indicating that the D816 and N822 mutations are on different alleles.
Comparison of results from AS-qPCR with melting-curve analysis was performed in 108 CBF AML patients. The concordance rate of the presence of KIT mutations was 74.1% (80/108). All mutations determined by melting-curve analysis could also be detected by AS-qPCR assay. AS-qPCR was able to detect additional mutations in 28 samples (19 with t(8;21) and 9 with inv(16); Supplementary Figure S2a). The positivity of KIT mutations in these 28 samples (AS-qPCR-positive and melting-curve analysis-negative) were confirmed by MEMO-PCR (n=18) and pyrosequencing (n=10). Mutant allele level of those samples was extremely low (median, 0.048; range, 0.002–6.8; Supplementary Figure S2b). The observations indicated that AS-qPCR used in this study has superior sensitivity when compared with melting-curve analysis, without decreasing specificity.
Clinical characteristics of patients with CBF AML according to KIT mutation status in exon 17
CBF AML patients with a KIT mutation showed a significantly higher BM blast percentage (77.0% vs 73.0%, P=0.032) compared with those without a mutation. There were no significant differences in the distribution of gender, age and peripheral blood cell count according to KIT exon 17 mutation statuses. When analyzed in t(8;21) AML patients, the KIT mutation was associated with high BM blast percentage and low CD34 expression. Differences were not observed in inv(16) AML.
All KIT exon 8 mutations were either small deletions or insertions or combination of deletions and insertions involving codon D419, causing its loss or replacement. KIT mutations in exon 8 were found in 9 (8.1%) of the 111 CBF AML patients (2 with t(8;21) AML and 7 with inv(16) AML) and 6 (5.4%) had concurrent mutations in exons 8 and 17.
FLT3-ITD and -TKD mutations were detected in 6 (5.5%) and 3 patients (2.7%), respectively. NPM1 mutations were detected in 5 patients (4.5%); all were NPM1 exon 12 mutation type A. FLT3-ITD and -TKD and NPM1 mutations were not associated with the status of KIT exon 17 mutations. These results are presented in Table 1.
Clinical significance of KIT mutation allele level determined by AS-qPCR
We evaluated the prognostic impact of KIT mutations detected by highly sensitive AS-qPCR. There was a tendency for lower OS and shorter EFS in patients with KIT mutations, although these were not statistically significant (P=0.20 and P=0.247, respectively). Next, we re-evaluated the prognostic impact of KIT mutations according to the mutant allele level. High mutant level (n=26) is defined when mutant level is ⩾10. The estimated 3-year OS was lowest in patients with high mutant level (35.6%, P=0.00496; Figure 3a), although there were no significant differences between patients with low mutant level (n=43) and unmutated cases (72.5% and 72.5%, respectively, P=0.943). The estimated 3-year EFS was also the lowest in patients with high mutant level (28.9%), when compared with those with low (61.2%) and unmutated (63.9%) patients (P=0.0252; high mutant level vs unmutated, P=0.0193; high vs low mutant level, P=0.0168; low mutant level vs unmutated, P=0.963; Figure 3b). In multivariate analyses, high mutant level was associated with a statistically significant adverse impact on both OS (HR, 2.927; confidence interval (CI): 1.487–5.758, P=0.002) and cumulative incidence of relapse (HR, 3.199; CI: 1.209–8.467, P=0.019).
Changes of KIT mutation status after treatment
We evaluated the changes of KIT mutation status after treatment to understand the biologic behavior of leukemic subclones. Follow-up BM samples were obtained from 27 patients carrying KIT exon 17 mutations at diagnosis and who had achieved hematologic CR. The KIT mutant allele levels decreased with the decrease of fusion transcript (Figures 4a and b). We evaluated KIT exon 17 mutations in both diagnosis and relapse samples in 13 cases (Supplementary Table S2). At the time of full relapse, six revealed detectable mutations and increased fusion transcripts (nos 1–6; Figure 4b). There was a wide distribution of mutant allele levels at the time of relapse (median 11.8, range 3.2–60.1). Three of six patients harboring multiple KIT mutations at diagnosis lost the minor mutations while retaining dominant mutation at relapse. In 2 patients (nos 5 and 6) with double KIT mutations at diagnosis, initially detected minor mutations had increased. The initial allele level of minor KIT mutations was low (0.2 and 0.01, respectively) and increased to 40.9 and 4.9 at relapse, respectively (Figure 4c). Five of 13 relapsed patients revealed undetectable KIT mutations, unlike increased fusion transcript level (Figure 4d). No cases demonstrated newly developed KIT mutations that had not been observed at diagnosis.
The kinetics of changes of relative KIT mutant and fusion transcripts (RUNX1-RUNX1T1) levels at diagnosis, after induction therapy and during follow-up in four representative patients with KIT exon 17 mutations. (a) One patient who achieved hematological remission without relapse: KIT mutations were not detectable during follow-up after induction therapy. (b) One patient with leukemia relapsed: the mutant allele level in KIT had a similar tendency to those of fusion transcripts. (c) One patient with double KIT mutations (D816Y and D816V) at diagnosis relapsed with the minor mutation (D816V). (d) Unlike a rapidly rising fusion transcript level, D816V mutation was detected at initial diagnosis but not at relapse.
Discussion
In this study, we developed a new AS-qPCR method to detect KIT exon 17 mutations in AML. The method is very sensitive; the overall incidence of KIT mutations at initial presentation was 62.2%. The sensitive melting-curve analysis revealed 35.2% incidence of KIT mutations, which was used to verify the sensitivity. A previous study showed similar results, with KIT mutations detected in 34% of adult CBF AML cases analyzed by melting-curve analysis.5 More sensitive mutation-based PCR reported higher detection rates (46.2%), when compared with direct sequencing (19.2%) in t(8;21) AML.6 We ruled out possibility of the false positivity of the low-level mutations using complementary assays such as MEMO-PCR and pyrosequencing. Therefore, we speculate that the melting-curve analysis did not detect the majority of mutations with low level, and the high KIT mutation rate in this study was caused by additional patients having a low mutant level within the leukemic cell population.
CBF AML is heterogeneous with regard to the KIT mutation.11 This study demonstrated that multiple KIT exon 17 mutations are not infrequent phenomena in CBF AML. A total of 43.5% of the patients with the KIT mutations contained multiple mutations. Most common were the dual mutations of D816H and N822K, followed by those of D816Y and N822K. This high rate of multiple mutations was detected because of the high sensitivity of AS-qPCR. There are a few reports on multiple KIT exon 17 mutations in the same individuals as the incidence is very low.2, 11, 21 The cloning analysis showed that D816 and N822 double mutations were located on a different allele or on different cells. D816 or S821 and N822 mutations have been reported on the same or different allele.2, 21 Mutant level in patients with multiple mutations was not different from those with single mutation. On the contrary, the sum of allele levels of multiple mutations was lower because the majority (63.3%) was composed of low allele level mutations (<10) without dominant mutations. Based on results from this study, we presume that the leukemic cells in CBF AML may harbor multiple genetically distinct and quantitatively small populations of subclones, in addition to the dominant KIT-negative leukemia clones at initial presentation. These tiny mutations are not detectable with Sanger sequencing or melting-curve analysis.
The prognostic impact of KIT mutations remains controversial, although KIT mutation is associated with poor prognosis in CBF AML patients, and alternative studies have recently quantified the KIT mutant allele level, which is measured by the pyrosequencing technique that is less sensitive than AS-qPCR is. The range of KIT mutant level was very wide in our study, similar to previous studies, which reported heterogeneity in mutant level (median 38%, range 5–78%).3 It has been reported that the mutant allele level, rather than the presence or absence of mutations in the FLT3, N-ras or p53 genes, is more significant prognostic factor in AML patients.22, 23, 24
Relatively few studies to date have evaluated the mutant level in KIT gene. Allen et al.3 reported that the high KIT mutant level was a significant adverse factor for relapse and OS was also lowest for the high mutant level group. A standardized analytical method with considerable sensitivity and accuracy is essential to minimize the unpredictable influence of KIT exon 17 mutation. It is also important to homogenize the calculation method for each heterogeneous mutation site using adequate reference genes and to determine a more significant cutoff level. We selected the relative mutant level calculated by a ratio of mutant allele copies per 100 control allele (mutation-unspecific region of the KIT gene) copies.
Mutant level in cases with multiple mutations was determined as the sum of each mutation. The sensitivity of the AS-qPCR was very high, thus the limit of detection was <200 copies per μl in both D816 and N822 mutations. We grouped the KIT exon 17 mutant level with a cutoff of 10 on the basis of mutation distribution. A high mutant level was an independent poor prognostic factor for OS, EFS and cumulative incidence of relapse, although the presence or absence of mutations as detected by AS-qPCR did not show significant difference in prognosis. These findings suggest that the prognostic impact of mutations cannot be explained simply by the presence or absence of molecular abnormality, when the KIT mutation-positive group included cases with low mutant levels detected by a highly sensitive method. On the other hand, a low mutant level (<10) did not reveal prognostic significance when compared with KIT wild type.
The follow-up of KIT mutation status after treatment demonstrated that KIT mutations became undetectable or gradually negative in patients during CR. These findings were consistent with those of an earlier study, which reported that the KIT mutation was undetected by allele-specific PCR in t(8;21) AML following CR.25 In addition, we performed a comparative analysis on paired diagnosis and relapse marrow samples. Patients harboring the KIT mutations at diagnosis retained the identical mutations at relapse, suggesting that KIT mutations have an important role in the leukemogenesis in a substantial number of CBF AML patients. However, five patients in this study had lost KIT mutations at relapse, which implies that the outcome of patients with CBF AML does not appear to be as homogeneous as their cytogenetic definition and KIT mutations alone are insufficient for transformation. There have been few studies on the molecular alterations associated with the relapse of CBF AML. We were also unable to find acquisition of new mutations involving KIT exon 17 at relapse. In one case with multiple mutations at diagnosis, minor leukemia clone with the KIT mutation detected by only AS-qPCR at diagnosis became dominant at relapse. This finding was similar to a previous report, in which a minor leukemia clone harboring KIT mutation at initial presentation developed resistance to treatment and became involved in relapse.6 However, the low-level leukemic cells with KIT mutations could not be detected with a less sensitive method, suggesting expansion of minor clones associated with drug resistance at relapse that could not be detected at diagnosis because of limited sensitivity of methods may be misinterpreted as clonal evolution.26 The results emphasize how accurate and quantitative mutation detection with a highly sensitive assay is necessary to understand the biologic behavior of KIT mutations at diagnosis and after treatment. Therefore, the use of KIT mutations as an independent marker for MRD monitoring seems to be limited in a subset of CBF AML patients because of the heterogeneity and instability of leukemic clone. The possible evolution of KIT mutations should be further explored and analyzed in future studies.27
In conclusion, we have successfully developed a highly sensitive detection method to identify common KIT mutations and quantify the relative mutant levels. The small populations of leukemic cells with KIT mutations could be detected and the linearity was excellent. The incidence of KIT exon 17 mutations in CBF AML patients is highest and the heterogeneous populations of leukemic cells exist as minor subclones along with the dominant leukemia clones. High mutant level (⩾10) was a poor prognostic factor rather than the KIT mutation positivity. The highly sensitive and quantitative measurement of the KIT exon 17 mutation is a valuable tool in evaluating prognosis at diagnosis and subsequent patient follow-up. Taken together, the study’s results show the necessity of a prospective therapeutic strategy to improve the clinical outcome for a specific group of patients with a high mutant allele level.28
References
Ashman LK, Griffith R . Therapeutic targeting of c-KIT in cancer. Expert Opin Investig Drugs 2013; 22: 103–115.
Paschka P, Marcucci G, Ruppert AS, Mrozek K, Chen H, Kittles RA et al. Adverse prognostic significance of KIT mutations in adult acute myeloid leukemia with inv(16) and t(8;21): a Cancer and Leukemia Group B Study. J Clin Oncol 2006; 24: 3904–391.
Allen C, Hills RK, Lamb K, Evans C, Tinsley S, Sellar R et al. The importance of relative mutant level for evaluating impact on outcome of KIT, FLT3 and CBL mutations in core-binding factor acute myeloid leukemia. Leukemia 2013; 27: 1891–1901.
Boissel N, Leroy H, Brethon B, Philippe N, de Botton S, Auvrignon A et al. Incidence and prognostic impact of c-Kit, FLT3, and Ras gene mutations in core binding factor acute myeloid leukemia (CBF-AML). Leukemia 2006; 20: 965–970.
Park SH, Chi HS, Min SK, Park BG, Jang S, Park CJ . Prognostic impact of c-KIT mutations in core binding factor acute myeloid leukemia. Leuk Res 2011; 35: 1376–1383.
Wakita S, Yamaguchi H, Miyake K, Mitamura Y, Kosaka F, Dan K et al. Importance of c-kit mutation detection method sensitivity in prognostic analyses of t(8;21)(q22;q22) acute myeloid leukemia. Leukemia 2011; 25: 1423–1432.
Cairoli R, Beghini A, Grillo G, Nadali G, Elice F, Ripamonti CB et al. Prognostic impact of c-KIT mutations in core binding factor leukemias: an Italian retrospective study. Blood 2006; 107: 3463–3468.
Riera L, Marmont F, Toppino D, Frairia C, Sismondi F, Audisio E et al. Core binding factor acute myeloid leukaemia and c-KIT mutations. Oncol Rep 2013; 29: 1867–1872.
Kim HJ, Ahn HK, Jung CW, Moon JH, Park CH, Lee KO et al. KIT D816 mutation associates with adverse outcomes in core binding factor acute myeloid leukemia, especially in the subgroup with RUNX1/RUNX1T1 rearrangement. Ann Hematol 2013; 92: 163–171.
Gilliland DG . Hematologic malignancies. Curr Opin Hematol 2001; 8: 189–191.
Qin YZ, Zhu HH, Jiang Q, Jiang H, Zhang LP, Xu LP et al. Prevalence and prognostic significance of c-KIT mutations in core binding factor acute myeloid leukemia: a comprehensive large-scale study from a single Chinese center. Leuk Res 2014; 38: 1435–1440.
Becker H, Pfeifer D, Afonso JD, Nimer SD, Veelken H, Schwabe M et al. Two cell lines of t(8;21) acute myeloid leukemia with activating KIT exon 17 mutation: models for the 'second hit' hypothesis. Leukemia 2008; 22: 1792–1794.
Newton CR, Graham A, Heptinstall LE, Powell SJ, Summers C, Kalsheker N et al. Analysis of any point mutation in DNA. The amplification refractory mutation system (ARMS). Nucleic Acids Res 1989; 17: 2503–2516.
Bench AJ, White HE, Foroni L, Godfrey AL, Gerrard G, Akiki S et al. Molecular diagnosis of the myeloproliferative neoplasms: UK guidelines for the detection of JAK2 V617F and other relevant mutations. Br J Haematol 2013; 160: 25–34.
Lu Q, Huang X, Chen H, Zhao X . A novel melting curve-based method for detecting mutations in acute myeloid leukemia. Oncol Lett 2014; 8: 99–104.
Lee ST, Kim JY, Kown MJ, Kim SW, Chung JH, Ahn MJ et al. Mutant enrichment with 3′-modified oligonucleotides a practical PCR method for detecting trace mutant DNAs. J Mol Diagn 2011; 13: 657–668.
Pollard JA, Alonzo TA, Gerbing RB, Ho PA, Zeng R, Ravindranath Y et al. Prevalence and prognostic significance of KIT mutations in pediatric patients with core binding factor AML enrolled on serial pediatric cooperative trials for de novo AML. Blood 2010; 115: 2372–2379.
Meshinchi S, Woods WG, Stirewalt DL, Sweetser DA, Buckley JD, Tjoa TK et al. Prevalence and prognostic significance of Flt3 internal tandem duplication in pediatric acute myeloid leukemia. Blood 2001; 97: 89–94.
Cheson BD, Cassileth PA, Head DR, Schiffer CA, Bennett JM, Bloomfield CD et al. Report of the National Cancer Institute-sponsored workshop on definitions of diagnosis and response in acute myeloid leukemia. J Clin Oncol 1990; 8: 813–819.
Kanda Y . Investigation of the freely available easy-to-use software 'EZR' for medical statistics. Bone Marrow Transplant 2013; 48: 452–458.
Shih LY, Liang DC, Huang CF, Chang YT, Lai CL, Lin TH et al. Cooperating mutations of receptor tyrosine kinases and Ras genes in childhood core-binding factor acute myeloid leukemia and a comparative analysis on paired diagnosis and relapse samples. Leukemia 2008; 22: 303–307.
Shih LY, Huang CF, Wu JH, Lin TL, Dunn P, Wang PN et al. Internal tandem duplication of FLT3 in relapsed acute myeloid leukemia: a comparative analysis of bone marrow samples from 108 adult patients at diagnosis and relapse. Blood 2002; 100: 2387–2392.
Wada H, Asada M, Nakazawa S, Itoh H, Kobayashi Y, Inoue T et al. Clonal expansion of p53 mutant cells in leukemia progression in vitro. Leukemia 1994; 8: 53–59.
Radich JP, Kopecky KJ, Willman CL, Weick J, Head D, Appelbaum F et al. N-ras mutations in adult de novo acute myelogenous leukemia: prevalence and clinical significance. Blood 1990; 76: 801–807.
Wang YY, Zhou GB, Yin T, Chen B, Shi JY, Liang WX et al. AML1-ETO and C-KIT mutation/overexpression in t(8;21) leukemia: implication in stepwise leukemogenesis and response to Gleevec. Proc Natl Acad Sci USA 2005; 102: 1104–1109.
Choi S, Henderson MJ, Kwan E, Beesley AH, Sutton R, Bahar AY et al. Relapse in children with acute lymphoblastic leukemia involving selection of a preexisting drug-resistant subclone. Blood 2007; 110: 632–639.
Wang Y, Wu DP, Liu QF, Qin YZ, Wang JB, Xu LP et al. In adults with t(8;21)AML, posttransplant RUNX1/RUNX1T1-based MRD monitoring, rather than c-KIT mutations, allows further risk stratification. Blood 2014; 124: 1880–1886.
Yoon JH, Kim HJ, Kim JW, Jeon YW, Shin SH, Lee SE et al. Identification of molecular and cytogenetic risk factors for unfavorable core-binding factor-positive adult AML with post-remission treatment outcome analysis including transplantation. Bone Marrow Transplant 2014; 49: 1466–1474.
Acknowledgements
This study was supported by a grant of the Korean Health Technology R&D Project, Ministry for Health and Welfare, Korea (HI14C3417).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Blood Cancer Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Jang, W., Yoon, JH., Park, J. et al. Significance of KIT exon 17 mutation depends on mutant level rather than positivity in core-binding factor acute myeloid leukemia. Blood Cancer Journal 6, e387 (2016). https://doi.org/10.1038/bcj.2015.116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bcj.2015.116
This article is cited by
-
Non-age-related neoplastic loss of sex chromosome correlated with prolonged survival in real-world CBF-AML patients
International Journal of Hematology (2022)
-
Prognostic values of D816V KIT mutation and peri-transplant CBFB-MYH11 MRD monitoring on acute myeloid leukemia with CBFB-MYH11
Bone Marrow Transplantation (2021)
-
Heterogeneous prognosis among KIT mutation types in adult acute myeloid leukemia patients with t(8;21)
Blood Cancer Journal (2018)