Abstract
In Ph-positive (Ph+) leukemia, the quiescent cell state is one of the reasons for resistance to the BCR-ABL-kinase inhibitor, imatinib. In order to examine the mechanisms of resistance due to quiescence and the effect of the mammalian target of rapamycin inhibitor, everolimus, for such a resistant population, we used Ph+ acute lymphoblastic leukemia patient cells serially xenotransplanted into NOD/SCID/IL2rγnull (NOG) mice. Spleen cells from leukemic mice showed a higher percentage of slow-cycling G0 cells in the CD34+CD38− population compared with the CD34+CD38+ and CD34− populations. After ex vivo imatinib treatment, more residual cells were observed in the CD34+CD38− population than in the other populations. Although slow-cycling G0 cells were insensitive to imatinib in spite of BCR-ABL and CrkL dephosphorylation, combination treatment with everolimus induced substantial cell death, including that of the CD34+CD38− population, with p70-S6 K dephosphorylation and decrease of MCL-1 expression. The leukemic non-obese diabetic/severe combined immunodeficiency (NOD/SCID) mouse system with the in vivo combination treatment with imatinib and everolimus showed a decrease of tumor burden including CD34+ cells. These results imply that treatment with everolimus can overcome resistance to imatinib in Ph+ leukemia due to quiescence.
Similar content being viewed by others
Introduction
Marked clinical improvement was reported when a BCR-ABL-kinase inhibitor, imatinib, was combined with chemotherapy for the treatment of Ph-positive (Ph+) acute lymphoblastic leukemia (ALL).1, 2 However, further improvement is needed to decrease relapses owing to residual or resistant leukemic cells. Leukemia stem cells (LSCs) are reported to be responsible for leukemia relapse.3 Mathematical models of the chronic phase of chronic myeloid leukemia suggested that imatinib does not eradicate LSCs,4, 5 and the survival of malignant cells is reportedly attributable to the quiescence of LSCs.6
In order to overcome drug resistance, rational combinations of molecular targeting drugs of different signal pathways have been explored.7 The mammalian target of rapamycin (mTOR) has attracted attention as a therapeutic target of LSCs.8, 9 mTOR regulates cell growth and apoptosis through the phosphatidilinositide-3-kinase (PI3K)/AKT/mTOR pathway, which was reported to be constitutively activated in most AMLs10 or in T-ALL cell lines.11 The inhibitory effect of an mTOR inhibitor, rapamycin, on the Ph+ leukemia cell lines with T315I was reported,7 and more recently, the effect of everolimus (RAD001) on human childhood B-cell progenitor ALL was reported in a non-obese diabetic/severe combined immunodeficiency (NOD/SCID) model.12 The mTOR inhibitor everolimus is an orally available mTOR inhibitor which was approved by the Food and Drug Administration for advanced renal cell carcinoma. A phase I/II study has been performed in patients with acute leukemia,13 and clinical trials alone or in combination with other drugs are also currently ongoing for lymphoma and myeloma.14 Effects of everolimus on human Ph+ ALL have not been well examined.
In this study, we examined the efficacy of everolimus in combination with imatinib utilizing Ph+ ALL cell lines and an NOD/SCID/IL2rγnull (NOG) mouse model of human BCR-ABL+ leukemia,15 in which the hierarchy of leukemia cells was maintained.
Materials and methods
Leukemic cells
Xenografts were established in NOD/SCID/IL2rγnull (NOG) mice as previously described.15 Briefly, Ph+ ALL patient cells were serially xenotransplanted into immunodeficient NOG mice, and engrafted spleen cells were obtained 8–10 weeks after injection. Erythrocytes were removed by erythrocyte lysis buffer (EL-buffer; Qiagen, Hilden, Germany), and the remaining leukemic cells were preserved in liquid nitrogen until use. Leukemic repopulated cells were thawed and washed, resuspended in RPMI containing 10% fetal bovine serum, 5 mM MgCl2 and 0.2 mg/ml DNase I (Roche Diagnostics, Mannheim, Germany) and incubated at 37 °C for 10 min. Cells were washed and resuspended at 1 million cells per ml in RPMI containing 20% fetal bovine serum with cytokines (human stem cell factor, 50 ng/ml, human interleukin-3 20 ng/ml and human granulocyte/macrophage-colony-stimulating factor, 20 ng/ml), and incubated with imatinib for 48 h at 37 °C in a CO2 incubator. In an in vitro long-term culture, spleen (SP) cells derived from leukemic NOG mice were co-cultured with S17 stromal cells and treated with imatinib and everolimus.16 S-17 cells and leukemic cells were passaged twice weekly.
Reagents
Everolimus and imatinib were supplied by Novartis Institutes for Biomedical Research (Basel, Switzerland). Imatinib was dissolved in dH2O and used for in vitro and in vivo experiments. Everolimus was stored as 10−2 M stock solution in dimethylsulfoxide for an in vitro experiment. For in vivo experiments, everolimus was formulated at 2% (wt/wt) in a microemulsion vehicle. Aliquots of everolimus and control vehicle were stored at −20 °C.
Immunoblotting
Antibodies against the phospho(p)-Abl (Tyr245), p-CrkL (Tyr207), p-mTOR (Ser2448), p-p70 S6 kinase (Thr389), p-4EBP1 (Thr37/46), MCL-1, p-AKT (Ser473), AKT and p-FOXO1(Thr24)/FoxO3a(Thr32) were from Cell Signaling (Boston, MA, USA). Immunoblotting was performed with the standard protocols as previously described.17
Flow cytometric analysis and cell sorting
After the treatment period, cells were washed at 4 °C and then stained with anti-CD34-allophycocyanin (APC), anti-CD38-PE-Cy7 (Becton Dickinson, San Jose, CA, USA), and anti-CD45-APC-Alexa Fluor 750 (Invitrogen, Carlsbad, CA, USA) antibodies (1:100) for 30 min on ice. Cells were subsequently labeled with annexin-V-fluorescein isothiocyanate and propidium iodide (PI) according to the manufacturer's protocol (Annexin-V-FLUOS Staining Kit; Roche Diagnostics). The cells were acquired by FACS Aria (Becton Dickinson) and analyzed by Flow Jo software. DNA contents analysis was assessed using the standard procedure as previously described.18 For CD34+ selection, leukemic cells were subjected to immunomagnetic separation using a MACS CD34 MicroBead Kit (Miltenyi Biotech, Auburn, CA, USA) following the manufacturer's recommendations. The collected cells were applied to a second column and the purification step was repeated. Staining of cells with Hoechst 33342 (Sigma, St Louis, MO, USA) with PyroninY (Polysciences, Warrington, PA, USA) was performed as previously described.19 Briefly, thawed leukemic spleen cells were separated with the MACS CD34 MicroBead Kit (Miltenyi Biotech) into CD34+ cells and flow-through cells containing CD34− cells. MACS-separated cells or drug-treated cells on S17 were washed and stained with Hoechst 33342 and PyroninY, and washed at 4 °C. MACS-separated CD34+ cells were then stained with anti-CD38-APC, and flow-through cells containing CD34− cells were stained with anti-CD34-APC. Flow cytometric analysis was performed using FACS Aria.
For cell sorting, leukemic spleen cells were stained with anti-CD34-APC, anti-CD38-PE-Cy7 and anti-CD45-APC-Cy7 antibodies and labeled with PI. PI− CD45+ cells were sorted for CD34 and CD38 expression using FACS Aria, incubated with treatment drugs for 6 h at 37 °C in a CO2 incubator as described above.
Mouse models
Humanized leukemic mouse model
NOG and NOD/SCID mice were obtained from the Central Institute for Experimental Animals (Kawasaki, Japan) and Clea Japan (Tokyo, Japan), respectively. Seven-week-old male NOD/SCID mice received 2 Gy of total body irradiation from an X-ray source (MBR-1520R-3; Hitachi Medico Technology Corporation, Tokyo, Japan), 24 h before administration of leukemic 2 × 107 cells. Everolimus (5 mg/kg), imatinib (100 mg/kg) or vehicle were diluted with dH2O, and given daily at 10 ml/kg for 10 days by gavage. Bone marrow and spleen cells were stained with anti-human CD19-PE (Becton Dickinson) and anti-mouse CD45-PerCP, and acquired with FACS Aria to analyze chimerism. Cells were also stained with anti-CD34-APC, anti-CD38-PE-Cy7 and anti-CD45-APC-Alexa Fluor 750, and were subsequently labeled with annexin-V–fluorescein isothiocyanate and PI as explained above. Protocols were approved by the Nagoya University Animal Ethics Committee.
Histopathology
Livers, spleens and femurs were fixed in 15% buffered formalin. Hematoxylin and eosin, and CD34 staining were performed as previously described.20, 21, 22 Slides were examined at room temperature and images were captured by a fluorescence microscope with a charge-coupled device camera (BZ-8000; Keyence, Osaka, Japan) fitted with × 20 and × 60 objective. Images were acquired by BZ-analyzer software (Keyence).
Statistical analysis
Differences among more than two groups were analyzed with the Bonferroni test followed by one-way analysis of variance. Statistical analyses were performed with STATA 9.2 software (StataCorp, College Station, TX, USA).
Results
Ex vivo treatment with imatinib for more residual quiescent CD34+CD38− population in Ph+ ALL cells
We analyzed the cell cycle status of untreated spleen cells derived from the humanized Ph+ ALL leukemia murine model reported previously.15 In the CD34+CD38− population, a higher percentage of Hoechstlow/PyroninYlow slow-cycling quiescent cells was observed than in the CD34+CD38+ population (P<0.05) and CD34− population (P<0.01). A lower percentage of the S+G2/M population was also observed among CD34+CD38− cells (P<0.05; Figure 1a, Supplementary Figure S1a).
Ex vivo analysis of humanized mouse positive (Ph+) acute lymphoblastic leukemia cells. (a) Leukemic spleen cells were CD34 positively selected with MACS column. CD34+ cells were stained with Hoechst, PyroninY and CD38-allophycocyanin (APC). Cells including CD34− population that had flowed through the column were stained with Hoechst, PyroninY and CD34-APC. (b) Leukemic spleen cells were ex vivo cultured with cytokines and treated with or without imatinib (IM) for 48 h. Human CD45+ propidium iodide (PI)− Annexin-V− viable population was analyzed for CD34 and CD38 distribution. Panels show a representative experiment. (c) After treated with IM for 48 h, CD34+ cells were positively selected with MACS column, and stained with Hoechst, PyroninY and CD38-APC. Cells including CD34− population that had flowed through the column were stained with Hoechst, PyroninY and CD34-APC. Graphs show the number of forward scatter/side scatter gated G0 cells in each CD34/CD38 sub-population, each relative to the untreated control. Bars indicate mean±s.d. values of three independent experiments (*P=0.03 between control and IM 3 μM for CD34+ CD38−, and **P=0.02 between control and IM 3 μM for CD34−, by one-way analysis of variance followed by Bonferroni). (d) CD34/CD38 sorted populations were treated with or without IM 3 μM for 6 h. Expression of BCR-ABL and phosphorylation of BCR-ABL and CrkL in each population was examined by western blotting analysis.
We next treated these cells with imatinib for 48 h and analyzed the distribution of CD34/CD38 in residual viable cells. After treatment with imatinib at 0.3, 1 and 3 μM, more residual CD34+CD38− cells were observed than non-treated cells (12.8 vs 27.2%; Figure 1b, P<0.05; Supplementary Figure S1b). Significantly more slow-cycling quiescent (G0) cells were observed within CD34+38− population after treated with 3 μM of imatinib (P=0.03), and less G0 cells in CD34− population (P=0.02, Figure 1c).
Treatment of CD34/CD38 sorted cells for 6 h with 3 μM imatinib caused equivalent inhibition of the phosphorylation of BCR-ABL and direct-substrate CrkL in each CD34/CD38 sub-population (Figure 1d). Inhibition of phosphorylation with imatinib (1 and 3.3 μM) was also observed with intracellular staining of phospho-CrkL (Supplementary Figure S1c). Expressions of BCR-ABL and ABL were equivalent in each CD34/CD38 sub-population (Figure 1d). These results suggested that the slow-cycling population derived from Ph+ leukemia NOG mice was insensitive to imatinib in spite of BCR-ABL dephosphorylation.
Ex vivo effects of everolimus on leukemic spleen cells, alone and in combination with imatinib
Furthermore, we have introduced S-17 murine stromal cell lines to support the leukemic spleen cells in order to assess longer-term effect of treatment drugs,16 as the CD34+ population of leukemic cells from the NOG mice eventually differentiated into CD34− cells and could not be maintained only with cytokines for a longer period. If cultured with S17 cells, leukemic spleen cells were viable for more than 30 days (Figure 2a).
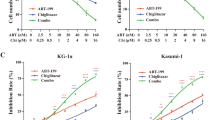
Ex vivo effects of everolimus on leukemic spleen cells in combination with imatinib (IM). (a) Leukemic spleen cells were co-cultured with S-17 stromal cells for up to 35 days. Cells were counted with Trypan blue, and viable cells were maintained. (b) Cells were treated with or without everolimus (Eve, 100 nM) and imatinib (1 μM) alone and in combination for 5 days on S-17 cells. DNA contents were assessed (upper panels) and Hoechst/PyroninY cell cycle analysis (lower panels) was performed. (c) Percentages of G0 population in total acquired cells were compared with dimethylsulfoxide (DMSO) control after 5-day treatment with imatinib (1 μM), everolimus (100 nM) or in combination. Graph shows the means±s.d. values of three independent experiments (*P<0.05 by one-way analysis of variance followed by Bonferroni). (d) In the stromal-culturing system, leukemic spleen cells were treated with DMSO (upper column) or in combination of imatinib and everolimus (lower column) for 5 days. Cells were stained with CD34, CD38, human CD45, propidium iodide, and Annexin-V. Cells were gated for human CD45+ and CD34/CD38, and cell viabilities in CD34+38− population are shown. Panels show a representative analysis.
To examine the potential of everolimus to overcome resistance due to quiescence in Ph+ leukemia cells, everolimus treatment was investigated ex vivo alone and in combination with imatinib on S17 stromal cells. Everolimus treatment at 100 nM for 5 days increased the sub-G1 population (14 vs 7.9%; control), and combination of everolimus (100 nM) and imatinib (1 μM) further increased the sub-G1 population (68%, Figure 2b, upper panel). Cell cycle status was also investigated after treatment with S-17 for 5 days. Although the untreated control and imatinib-treated cells showed 35% of Hoechstlow/PyroninYlow cells in total acquired cells, combination of imatinib and everolimus decreased these quiescent cells to 13% of total acquired cells (Figure 2b, lower panels). Significant difference was found between imatinib alone and combination of imatinib plus everolimus (Figure 2c). Treatment with everolimus and imatinib for 5 days induced substantial cell death in CD34+38− population relative to dimethylsulfoxide control (Figure 2d, Supplementary Figure S1d). These results indicated that ex vivo combination treatment with imatinib and everolimus was also effective for the quiescent CD34+38− cells.
Evaluation of molecular biomarkers during cell death induced by treatment with imatinib and everolimus
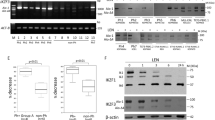
We next investigated the effects of imatinib and everolimus on BCR-ABL and mTOR signaling. Separated CD34+ cells were treated with and without imatinib (1 μM) or everolimus (100 nM) for 4 h. After imatinib treatment, phosphorylation of BCR-ABL was clearly inhibited in each population, but it was not affected after everolimus treatment (Figure 3a). After everolimus treatment, the phosphorylation of S6 K, which is a direct substrate of mTOR, was clearly inhibited; however, the phosphorylation of mTOR and 4EBP1 was not changed (Figure 3b). These results imply that everolimus inhibited mTOR signaling of CD34+ cells and induced cell death independently of the BCR-ABL signaling pathway. Both imatinib alone and in combined treatment inhibited phosphorylation of BCR-ABL. Conversely, everolimus alone and in combination both inhibited phosphorylation of S6 K in both CD34+38− and CD34+38+ sub-populations (Figure 3c). Everolimus alone or in combination with imatinib decreased the expression of the antiapoptotic BCL-2 family protein, MCL-1, after 4 h, and the combination of everolimus and imatinib also decreased the expression of MCL-1, not BCL-2, after 12 h (Figure 3c). These results implied that combination treatment with imatinib and everolimus induced cell death in quiescent Ph+ leukemia cells.
Western blot analysis of ex vivo-treated leukemic cells. (a) CD34+ cells were separated with MACS column and cells were treated with and without imatinib (IM, 3 μM) or everolimus (Eve, 300 nM) for 4 h. Each sample was lysed and western blotting analysis was performed with each antibody. (b) MACS-separated cells were treated with or without everolimus (100 nM) for 4 h. Each sample was lysed and western blotting was performed with p-mTOR, p-S6 K, p-4EBP1 antibodies. Immunoblotting by antitubulin was performed for the control. (c) CD34+38− and CD34+38+ sorted cells were treated with treatment drugs for 4 or 12 h. Each sample was lysed and western blotting analysis was performed with each antibody.
In vivo investigation of effects of everolimus, alone and in combination with imatinib
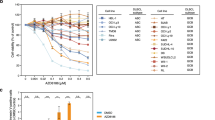
To elucidate the in vivo efficacy of everolimus treatment, its effects were investigated alone (5 mg/kg) and in combination with imatinib (100 mg/kg+everolimus 5 mg/kg) using NOD/SCID mice intravenously injected with leukemic spleen cells from humanized NOG mouse (Figure 4a). Percentage of CD19+ leukemic cells in peripheral blood was lowest in the imatinib-plus-everolimus-treated group, compared with the vehicle or imatinib alone (Figure 4b). Overall tumor burden, as assessed by spleen weight (Figure 4d) and the total number of splenic human CD19+ leukemic cells (Figure 4e), was observed to be lowest in the imatinib-plus-everolimus-treated group. Immunohistochemistry showed that the combination of imatinib plus everolimus decreased the infiltrated CD34+ human leukemic cells in spleen, liver and bone marrow (Figure 4f, Supplementary Figure S2). Everolimus alone also decreased the percentage of G0 cells in the CD34+ leukemic cells of the treated bone marrow (Supplementary Figure S3). These results indicated the in vivo efficacy of everolimus treatment in a Ph+ leukemia murine model.
In vivo effects of everolimus on leukemic spleen cells in combination with imatinib. (a) Non-obese diabetic/severe combined immunodeficiency ( mice were irradiated (IR), and leukemic spleen (SP) cells from NOG (2 × 107) were injected. Experiments were performed twice with three in each treatment group. Control vehicle (n=6), imatinib 100 mg/kg (n=6), everolimus 5 mg/kg (n=6) or a combination of both (n=6) were administered for 10 days, and mice were dissected 24 h after the last administration on day 28 following the tumor injection. (b, c) Percentages of CD19+ cells in peripheral blood (b) and bone marrow (c) are shown, respectively. (d) Spleen weight was relatively compared with the average of control mice in each experiment. Bars indicate average of spleen weight in each study group. (e) CD19+ leukemic spleen cell numbers were relatively compared with the average of control (b–e: *P<0.001, **P<0.005 and ***P<0.05 by one-way analysis of variance followed by Bonferroni). (f) Hematoxylin and eosin (HE) staining (left panels) and immunohistochemical analysis with CD34 (right panels) of the spleen from control mice and imatinib and/or everolimus-treated mice were performed. A charge-coupled device camera provided images at approximately × 100 of the original magnification.
Discussion
In this study, we showed the effects of everolimus in combination with imatinib against Ph+ ALL quiescent cells. Ex vivo imatinib treatment of Ph+ leukemia cells from a humanized mouse model showed more residual cells in the CD34+CD38− population, which contains significantly more quiescent cells. Our data showed an ex vivo effect against these residual cells, and the combination of imatinib and everolimus showed an in vivo effect. These data have shown the potential of everolimus to overcome imatinib resistance in quiescent cells. LSCs are reported to be responsible for the resistance to chemotherapy and molecular targeting agents.23, 24 In chronic myeloid leukemia, non-proliferating quiescent CD34+ cells have been found to be more resistant than proliferating leukemic cells after treatment with several chemotherapeutic agents.25 Other studies have shown that inhibitors of mTOR with conventional therapies induced apoptosis and reduced LSCs.8 The definition of LSCs or cancer stem cells is sometimes controversial in certain diseases. In human AML, LSCs have been phenotypically identified within a CD34+CD38− fraction.23, 26 In contrast, it is controversial whether ALL LSCs exist within the CD34+ fraction and how CD34, CD38, CD19 and CD133 relate to ALL LSCs.15, 27, 28, 29 In our current study, the potential of everolimus to overcome imatinib-resistant quiescent cells was demonstrated by using a humanized leukemic mouse model that maintains the differentiation hierarchy of Ph+ leukemia.15 However, it cannot be determined at this point if the real LSCs of Ph+ ALL can be diminished until the LSCs in this disease category are clarified.
MCL-1, an antiapoptotic member of the BCL-2 protein family, reportedly regulates the self-renewal of human hematopoietic stem cells as well as LSCs.30 Mills et al.31 also reported that MCL-1 was translationally regulated by mTORC1. Together with these reports, our results showing decreased expression of MCL-1 by combination treatment of imatinib and everolimus suggested that the combination treatment induced cell death of quiescent Ph+ leukemia cells by interfering with the mitochondrial-mediated cell death pathway. Rapamycin and its analogs are also known to induce autophagic cell death,32 and Bellodi et al.33 reported that target autophagy potentiates tyrosine kinase-induced cell death in Ph+ leukemia cells. We are also investigating the relation of autophagy in cell death in our experimental systems.
In this study, everolimus treatment of Ph+ leukemia cells from a humanized mouse model decreased the phosphorylation of S6 K, but it increased the phosphorylation of AKT (Ser473) and FOXO1/3a (Supplementary Figure S4a). Rapamycin and its analogs, such as everolimus and temsirolimus, are allosteric mTOR inhibitors that function at a distance from the adenosine triphosphate-catalytic binding site. Of the two cellular protein complexes of mTOR molecule, mTORC1 and mTORC2, mTORC1 is sensitive to these allosteric mTOR inhibitors and mTORC2 is resistant.34 mTORC2 directly activates AKT, and this AKT activation in a feedback loop has been reported to correlate with rapamycin failure.35 This feedback loop might also be related to our data on upregulated AKT.
Recently, a new generation of mTOR inhibitors has been developed. Dual PI3K/mTOR inhibitors, such as BEZ235, EX147 and PI-103, inhibit PI3 K and both small molecules of mTORC1/2.36 Adenosine triphosphate-competitive mTOR inhibitors that selectively inhibit TORC1/2 molecules also have been reported to be effective against Ph+ transformed leukemia cells and to be less immunosuppressive than PI3K/mTOR inhibitors.37 The effectiveness of a new generation of mTOR inhibitors should also be investigated in our future studies, in particular, the efficacy of these inhibitors against quiescent or leukemic stem cells using a humanized leukemic mouse model. However, it was suggested that dual PI3K/mTOR inhibitors may cause a greater degree of immune suppression by affecting normal cell functions.14 Although we have examined the colony formation of CD34+ human umbilical cord blood and it was suggested that everolimus did not severely interfere with hematopoietic colony formation (data not shown), the effects of everolimus and the new-generation mTOR inhibitors on normal cells and immune functions must be investigated in future studies.
Acquired mutation in the BCR-ABL gene also causes primary and secondary treatment failure in Ph+ leukemia. Our data suggest that imatinib-resistant cell lines with T315I mutation (Supplementary Figures S4b and c) can be inhibited with everolimus with downregulation of the mTOR pathway (Supplementary Figure S4d). The in vivo effect of everolimus on T315I-mutated Ph+ leukemic cells is also indicated (Supplementary Figure S4e). Further study is needed to determine the effect of everolimus on T315I-mutated leukemia, especially in combination with a T315I inhibitor such as AP24534 (ponatinib).38
In conclusion, we have investigated the imatinib and everolimus combination effect against human Ph+ quiescent leukemic cells utilizing a mouse model. Everolimus can improve the treatment of resistant Ph+ leukemia. These mice also provide the opportunity to evaluate the effects of new therapeutic modalities on leukemic cells in different stages of cell cycle.
References
Wassmann B, Pfeifer H, Goekbuget N, Beelen DW, Beck J, Stelljes M et al. Alternating versus concurrent schedules of imatinib and chemotherapy as front-line therapy for Philadelphia-positive acute lymphoblastic leukemia (Ph+ ALL). Blood 2006; 108: 1469–1477.
Quintas-Cardama A, Kantarjian H, Cortes J . Imatinib and beyond—exploring the full potential of targeted therapy for CML. Nat Rev Clin Oncol 2009; 6: 535–543.
Jordan CT . The leukemic stem cell. Best Pract Res Clin Haematol 2007; 20: 13–18.
Michor F, Hughes TP, Iwasa Y, Branford S, Shah NP, Sawyers CL et al. Dynamics of chronic myeloid leukaemia. Nature 2005; 435: 1267–1270.
Roeder I, Horn M, Glauche I, Hochhaus A, Mueller MC, Loeffler M . Dynamic modeling of imatinib-treated chronic myeloid leukemia: functional insights and clinical implications. Nat Med 2006; 12: 1181–1184.
O'Hare T, Corbin AS, Druker BJ . Targeted CML therapy: controlling drug resistance, seeking cure. Curr Opin Genet Dev 2006; 16: 92–99.
Mohi MG, Boulton C, Gu TL, Sternberg DW, Neuberg D, Griffin JD et al. Combination of rapamycin and protein tyrosine kinase (PTK) inhibitors for the treatment of leukemias caused by oncogenic PTKs. Proc Natl Acad Sci USA 2004; 101: 3130–3135.
Krause DS, Van Etten RA . Right on target: eradicating leukemic stem cells. Trends Mol Med 2007; 13: 470–481.
Yilmaz OH, Valdez R, Theisen BK, Guo W, Ferguson DO, Wu H et al. Pten dependence distinguishes haematopoietic stem cells from leukaemia-initiating cells. Nature 2006; 441: 475–482.
Xu Q, Thompson JE, Carroll M . mTOR regulates cell survival after etoposide treatment in primary AML cells. Blood 2005; 106: 4261–4268.
Cullion K, Draheim KM, Hermance N, Tammam J, Sharma VM, Ware C et al. Targeting the Notch1 and mTOR pathways in a mouse T-ALL model. Blood 2009; 113: 6172–6181.
Crazzolara R, Cisterne A, Thien M, Hewson J, Baraz R, Bradstock KF et al. Potentiating effects of RAD001 (Everolimus) on vincristine therapy in childhood acute lymphoblastic leukemia. Blood 2009; 113: 3297–3306.
Yee KW, Zeng Z, Konopleva M, Verstovsek S, Ravandi F, Ferrajoli A et al. Phase I/II study of the mammalian target of rapamycin inhibitor everolimus (RAD001) in patients with relapsed or refractory hematologic malignancies. Clin Cancer Res 2006; 12: 5165–5173.
Chapuis N, Tamburini J, Green AS, Willems L, Bardet V, Park S et al. Perspectives on inhibiting mTOR as a future treatment strategy for hematological malignancies. Leukemia 2010; 24: 1686–1699.
Tanizaki R, Nomura Y, Miyata Y, Minami Y, Abe A, Hanamura A et al. Irrespective of CD34 expression, lineage-committed cell fraction reconstitutes and re-establishes transformed Philadelphia chromosome-positive leukemia in NOD/SCID/IL-2Rgammac-/- mice. Cancer Sci 2010; 101: 631–638.
Minami Y, Stuart SA, Ikawa T, Jiang Y, Banno A, Hunton IC et al. BCR-ABL-transformed GMP as myeloid leukemic stem cells. Proc Natl Acad Sci USA 2008; 105: 17967–17972.
Minami Y, Yamamoto K, Kiyoi H, Ueda R, Saito H, Naoe T . Different antiapoptotic pathways between wild-type and mutated FLT3: insights into therapeutic targets in leukemia. Blood 2003; 102: 2969–2975.
Minami Y, Kiyoi H, Yamamoto Y, Yamamoto K, Ueda R, Saito H et al. Selective apoptosis of tandemly duplicated FLT3-transformed leukemia cells by Hsp90 inhibitors. Leukemia 2002; 16: 1535–1540.
Holyoake T, Jiang X, Eaves C, Eaves A . Isolation of a highly quiescent subpopulation of primitive leukemic cells in chronic myeloid leukemia. Blood 1999; 94: 2056–2064.
Ninomiya M, Abe A, Katsumi A, Xu J, Ito M, Arai F et al. Homing, proliferation and survival sites of human leukemia cells in vivo in immunodeficient mice. Leukemia 2007; 21: 136–142.
Zhou Y, Pan Y, Zhang S, Shi X, Ning T, Ke Y . Increased phosphorylation of p70 S6 kinase is associated with HPV16 infection in cervical cancer and esophageal cancer. Br J Cancer 2007; 97: 218–222.
Abe A, Kiyoi H, Ninomiya M, Yamazaki T, Murase T, Ozeki K et al. Establishment of a stroma-dependent human acute myelomonocytic leukemia cell line, NAMO-2, with FLT3 tandem duplication. Int J Hematol 2006; 84: 328–336.
Ishikawa F, Yoshida S, Saito Y, Hijikata A, Kitamura H, Tanaka S et al. Chemotherapy-resistant human AML stem cells home to and engraft within the bone-marrow endosteal region. Nat Biotechnol 2007; 25: 1315–1321.
Misaghian N, Ligresti G, Steelman LS, Bertrand FE, Basecke J, Libra M et al. Targeting the leukemic stem cell: the Holy Grail of leukemia therapy. Leukemia 2009; 23: 25–42.
Holtz MS, Forman SJ, Bhatia R . Nonproliferating CML CD34+ progenitors are resistant to apoptosis induced by a wide range of proapoptotic stimuli. Leukemia 2005; 19: 1034–1041.
Bonnet D, Dick JE . Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med 1997; 3: 730–737.
Cox CV, Diamanti P, Evely RS, Kearns PR, Blair A . Expression of CD133 on leukemia-initiating cells in childhood ALL. Blood 2009; 113: 3287–3296.
Hong D, Gupta R, Ancliff P, Atzberger A, Brown J, Soneji S et al. Initiating and cancer-propagating cells in TEL-AML1-associated childhood leukemia. Science 2008; 319: 336–339.
Kong Y, Yoshida S, Saito Y, Doi T, Nagatoshi Y, Fukata M et al. CD34+CD38+CD19+ as well as CD34+CD38-CD19+ cells are leukemia-initiating cells with self-renewal capacity in human B-precursor ALL. Leukemia 2008; 22: 1207–1213.
Campbell CJ, Lee JB, Levadoux-Martin M, Wynder T, Xenocostas A, Leber B et al. The human stem cell hierarchy is defined by a functional dependence on Mcl-1 for self-renewal capacity. Blood 2010; 116: 1433–1442.
Mills JR, Hippo Y, Robert F, Chen SM, Malina A, Lin CJ et al. mTORC1 promotes survival through translational control of Mcl-1. Proc Natl Acad Sci USA 2008; 105: 10853–10858.
Codogno P, Meijer AJ . Autophagy and signaling: their role in cell survival and cell death. Cell Death Differ 2005; 12 (Suppl 2): 1509–1518.
Bellodi C, Lidonnici MR, Hamilton A, Helgason GV, Soliera AR, Ronchetti M et al. Targeting autophagy potentiates tyrosine kinase inhibitor-induced cell death in Philadelphia chromosome-positive cells, including primary CML stem cells. J Clin Invest 2009; 119: 1109–1123.
Wullschleger S, Loewith R, Hall MN . TOR signaling in growth and metabolism. Cell 2006; 124: 471–484.
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM . Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 2005; 307: 1098–1101.
Meric-Bernstam F, Gonzalez-Angulo AM . Targeting the mTOR signaling network for cancer therapy. J Clin Oncol 2009; 27: 2278–2287.
Janes MR, Limon JJ, So L, Chen J, Lim RJ, Chavez MA et al. Effective and selective targeting of leukemia cells using a TORC1/2 kinase inhibitor. Nat Med 2010; 16: 205–213.
O'Hare T, Shakespeare WC, Zhu X, Eide CA, Rivera VM, Wang F et al. AP24534, a pan-BCR-ABL inhibitor for chronic myeloid leukemia, potently inhibits the T315I mutant and overcomes mutation-based resistance. Cancer Cell 2009; 16: 401–412.
Acknowledgements
We thank the staff of the Center for Research of Laboratory Animals and the Division for Medical Research Engineering, Nagoya University Graduate School of Medicine for their technical support. We are grateful to the Novartis Institutes for Biomedical Research for providing imatinib and everolimus for these experiments. We also are indebted to Y Nomura, R Tanizaki, T Kawake and C Wakamatsu for their technical assistance. This work was supported by Grants-in-Aids from the National Institute of Biomedical Innovation and from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Hitoshi Kiyoi received research grants from Novartis Pharma, Kyowa-Hakko Kirin Co. Ltd and Chugai Pharmaceutical Co. Ltd. Tomoki Naoe received research grants from Janssen Pharma, Novartis Pharma, Kyowa-Hakko Kirin Co. Ltd and Chugai Pharmaceutical Co. Ltd. The other authors have no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Blood Cancer Journal website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Kuwatsuka, Y., Minami, M., Minami, Y. et al. The mTOR inhibitor, everolimus (RAD001), overcomes resistance to imatinib in quiescent Ph-positive acute lymphoblastic leukemia cells. Blood Cancer Journal 1, e17 (2011). https://doi.org/10.1038/bcj.2011.16
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bcj.2011.16
Keywords
This article is cited by
-
Obesity and Leukemia: Biological Mechanisms, Perspectives, and Challenges
Current Obesity Reports (2023)
-
Non ABL-directed inhibitors as alternative treatment strategies for chronic myeloid leukemia
Molecular Cancer (2018)