Abstract
In drug design and discovery, binding affinity and selectivity are two basic properties of a drug candidate. Opioid receptors (ORs) are the main targets of strong analgesics. Like some other class A members of G-protein-coupled receptors (GPCRs), ORs exhibit complex selectivity on their ligands. The diversity of binding activity and selectivity among opioids has deeply attracted researchers for a long time. To investigate the subtype selectivity of μ, δ and κ ORs in detail, using the κ-selective antagonist JDTic as a probe, we performed a series of computational simulations, including molecular dynamics and metadynamics, on JDTic-μ/δ/κ-OR complexes. From the simulations, we found that the decisive factor of JDTic selectivity on the μ-subtype was the 2.63 position, which affected the efficacy of JDTic through changing the dynamics of the Q2.60 residue. In addition to the 2.63-position residue, the 7.35 position was the other crucial aspect of JDTic selectivity for the δ-subtype. Based on the results, we suggest a new concept, the “message-address-efficacy” hypothesis, to explain the relationships among the affinity, selectivity and function between ORs and opioids. Thus, all the detailed dynamics of JDTic-bound ORs might be helpful to deeply understand the subtype selectivity and binding mechanisms of other GPCRs.
Similar content being viewed by others
Introduction
G protein-coupled receptors (GPCRs) are the largest super-family of drug targets1. For their ligands, the selectivity may be divided into two types, subtype2 and functional selectivities3. The two types of selectivity are very common in GPCR ligands. For instance, 5-serotonin receptors, dopamine receptors and adrenergic receptors contain multiple subtypes, and most ligands on each subtype show different affinity or activity. In signal transduction, different ligands exhibit various subtype selectivity on G-protein-biased pathway or β-arrestin pathway, and even the same ligand shows different functional selectivity in different species or in different organs of the same species4. In general, an antagonist only displays subtype selectivity, while an agonist exhibits both subtype and functional selectivities.
Members of class A subfamily of GPCRs5 share a similar ligand binding pocket, which is located in the upper half of the receptor helix area near the extracellular loops6. Comparing these crystal structures, it is easy to find that the ligand binding sites of different receptors vary, leading to binding affinity and selectivity issues. As two basic elements of drug discovery, both affinity and selectivity should be considered for a druggable candidate, further indicating that a more reasonable mode of action is necessary for new ligands7.
The opioid receptors (ORs), members of the class A subfamily of GPCRs6, are the targets of analgesics. Researchers began to pay attention to the affinity and selectivity of opiate problems in 1970s and put forward the “message-address” hypothesis8,9,10 to explain the interaction mechanism of opioids. In the hypothesis, the “message” and “address” are two subsites for the receptor to recognize peptide ligand. The message component is involved in receptor recognition and related signal transduction, while the address could provide additional binding affinity and receptor subtype selectivity. The endogenous opioid peptides are hence recognized according to the message-address concept, in which a constant tetrapeptide sequence (Tyr-Gly-Gly-Phe) is contained11. The residue Tyr constitutes the message component, and the sequence starting with Phe comprises the address, whereas Gly-Gly functions as a spacer. This hypothesis is identical with the well-known structure-activity relationships of nonpeptide opioid ligands that comprise one aromatic ring to presumably mimic the Tyr residue. The “message-address” hypothesis has been used to design many OR ligands, such as the first non-peptide δ-OR antagonist Naltrindole (NTI)12 and its corresponding derivatives11,13.
According to the critical groups of opioids, the functions of some crucial residues14,15,16 were inferred, that were also validated by site-directed mutagenesis experiments17. Residues D3.32 (Ballesteros/Weinstein numbering18) and H6.52, as well as their surrounding areas, are basic residues of opioid recognition, and play "message" roles, while "address" residues are responsible for subtype selectivity on the upper part of the binding pocket and second extracellular loop (ECL2) area.
However, the selectivity mechanism of address residues is intriguing, such as that for NTI19,20. Observed from the crystal structure, the crux of ligand efficacy is W6.58 at the upper part of the binding pocket. However, the molecular superposition19 and mutation results21 showed that the 7.35-position is the critical factor for NTI subtype selectivity. Moreover, the "message-address" hypothesis was not suitable for all opioids20,22. For instance, the κ-subtype selective ligand Salvinorin A is a special type of ligand, and its mode of action is different from that of morphinan compounds.
Residue mutation experiments identified the selective site of ligands 5′-GNTI and 6′-GNTI at the 6.58 position23,24,25 and NTI at the 7.35 position21,25; the chimera method26 demonstrated that the ECL2 region is the crucial aspect for the subtype selectivity of ligand U50488. These experimental methods are efficient in the study of ligand selectivity, but consume many reagents and raw materials, while the simulated mutation study could save these resources.
Among the selective ligands of ORs, JDTic27 and its analogs28,29 are a class of highly selective κ-OR antagonists (Figure 1). It was difficult to explain their mechanisms of high selectivity in the absence of OR crystal structures30. After the crystal structures were determined, Wu et al20 and Granier et al19 tried explaining related selectivity mechanisms of opioid ligands through molecular superposition and homology modeling methods. They focused on the key roles of the 7.35- and 2.63-position residues on the selectivity of JDTic. However, this was still static cognition and limits our understanding of the selectivity mechanism.
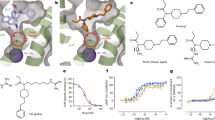
Two-dimensional structures of the antagonists JDTic and JPP6. The key parts were labeled with a dotted circle.
There are many ligands that exhibit high selectivity for each subtype of ORs. Elucidation of the related selective mechanism would deepen our understanding of their interactions and help to design new selective ligands. Moreover, the selectivity mechanism of many reported compounds is not clear, suggesting that much work should be performed on ligand selectivity.
Stevens et al resolved the crystal structure of the human κ-OR in complex with its selective antagonist JDTic20, and the crystal structures of μ/δ-OR were also reported recently19,31,32. These structures provide atomic structural information on ORs14,15 and a new opportunity to study JDTic selectivity on ORs from the dynamics level.
Here, we selected JDTic as a probe and performed unbiased molecular dynamics (MD) and metadynamics (MT) simulations to elucidate the ligand selectivity on μ/δ/κ-OR subtypes (Table 1). Combined with the simulated mutation method, we found some key residues to subtype selectivity of JDTic. Furthermore, to interpret the relationships among the affinity, selectivity and function between opioid receptors and ligands, we proposed the "message-address-efficacy" hypothesis mainly based on the receptor interaction mechanisms. Thus, our study provided a useful method and a new idea for the subtype selectivity and binding mechanism of ORs.
Materials and methods
Protein preparation
The crystal structures of inactive μ/δ/κ-OR were obtained from the Protein Data Bank (PDB, entry codes: 4DKL31, 4N6H32 and 4DJH20), which were engineered proteins with T4 lysozyme (T4L)20 or nano antibody. To perform MD simulations on the wild-type receptor, T4L and other unnecessary parts were removed from the crystal structures, and the loop was reconstructed by adding the missing residues with the loop refinement protocol in Discovery Studio 3.5, such as S262 and T302–S303–H304–S305–T306 in κ-OR. The generated 10 loop models were clustered as a dendrogram based on the pairwise main-chain root-mean-square deviation (RMSD) of the loop region33, and the one with the lowest RMSD value was chosen for loop reconstruction. The Na+ ion located at the allosteric pocket position32 was exactly positioned according to the crystal structure of NTI-complexed δ-OR (PDB entry code: 4N6H).
The Protein Preparation Tool (ProPrep) module in the Schrödinger 2012 suite software package was then used to prepare the above integrated three-dimensional structures of μ/κ/δ-OR. Residues Asn, Gln, and His were checked for the protonated state automatically in ProPrep. The PROPKA tool in Maestro was utilized to add the hydrogen atoms into the three structures at physiological pH with an optimized hydrogen bond network.
Preparation of ligand JDTic and related residues
The ligand JDTic in the crystal structure of κ-OR was subjected to ProPrep together with κ-OR. Hydrogen atoms were added through the PROPKA tool. The protonated state of nitrogen atoms on JDTic was determined through the Epik module. The pose of JDTic in the final complex was extracted and employed as the starting point for subsequent simulations.
The Paramchem webserver34,35,36 (https://cgenff.paramchem.org/), a program coupled with the CHARMM force field, was used to generate the force field parameters of JDTic. We imported JDTic with a correct configuration into the webserver and obtained the parameters of the compound through a script.
In μ/δ-OR, the same JDTic structure was adopted as that in κ-OR. Due to the position collision with JDTic, the poses of residues Q2.60 and Y7.43 located at the binding pocket were adjusted based on those in the crystal structure of κ-OR (Figure 2).
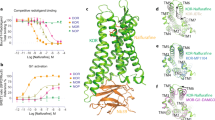
(A) Key interactive residues within 4 Å of JDTic in the initial JDTic-OR complexes among MD simulations. (B) The initial angle and measured atoms of JDTic in the crystal structure. The backbones of residues and JDTic were colored in green (μ-OR), cyan (δ-OR), orange (κ-OR) and slate-blue (JDTic).
According to the residue Val at the 2.63 position in the crystal structure of κ-OR, related residues Asn and Lys at the 2.63 position in μ/δ-OR were mutated to Val using the Residue Mutation module in DS3.5.
MD preparations
Five systems, JDTic-wild type μ/δ/κ-OR and JDTic-mutated μ/δ-OR complexes, were built for the simulations. A POPC (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine) bilayer with the surface area of 75×75 Å2 on the X-Y plane was generated under the Charmm36 force field using the VMD program (Version 1.9.1) (Table 1). For each system, our in-house program was first used to embed the receptor into the POPC bilayer pre-aligned in the OPM (Orientations of Proteins in Membranes) database37,38,39,40. Thereafter, pre-equilibrated POPC lipids coupled with TIP3P water molecules in a box ∼ 75×75×100 Å3 were used to solvate the protein (Table 1). Water molecules in the bilayer and lipid molecules within 0.85 Å of the heavy atoms on the protein structure were removed. 51 Na+ and 66/68/58 Cl− ions were added into the JDTic-bound μ/δ/κ-OR systems to neutralize the systems with 0.15 mol/L NaCl in the water phase. The protein was described using the CHARMM36 force field with cmap correction.
MD simulations
All the simulations were carried on the high-performance cluster of the State Key Laboratory of Bioreactor Engineering using Gromacs V.4.6.541 and the CHARMM36 parameters for all compositions. In the first simulation step, a 10000-step energy minimization was implemented with 1000.0 kJ/mol/nm as the force threshold for the systems. Next, a 100-ps initial equilibration was produced to gradually heat the systems from 0 K to 310 K at a constant volume and temperature at 310 K (NVT). The systems were subjected to an additional 1-ns equilibration at constant pressure and temperature (NPT ensemble; 310 K, 1 atm) with three thermostats (stabilizing temperature independently for the protein-ligand system, lipid POPCs and water-ion system) at 2-ps time steps. The cut-off value for vdW and short-range electrostatic interactions was set at 12 Å. The Particle Mesh Ewald (PME) summation scheme was used to compute long-range electrostatic interactions. Finally, the production MD simulations of the five systems were performed for 100 ns under NPT conditions using the Nose-Hoover thermostat for temperature coupling and Parrinello-Rahman pressure coupler methods. The integrator leap-frog algorithm was employed, and the time step for MD simulation was 2 fs.
Metadynamics simulations
Theory: In an MT simulation, an additional history-dependent biased potential VG(S,t) was introduced into the system,

where S represents the collective variables, t represents time, ω is the energy rate and σi controls the width of the Gaussian for the ith collective variable. With the evolution of the system, the wells in the FES (free energy surface) of the collective variables are filled up with the biased potential VG. The underlying free energy -F(s) is assumed to be estimated from the biased potential once all the wells have been filled after a sufficiently long time,

The correctness of the relationship as shown in formula 2 was proven to be empirical by extensive tests under the assumption that the stochastic dynamics in the collective variable space is memoryless in the absence of the bias. Under the assumption, the error in FES construction was proven to be:

where D is the intrinsic system diffusion coefficient in the collective variable space, κB is the Boltzmann constant, and T is the temperature of the system42.
In fact, a “well-tempered” and “smoothly converging” algorithm was introduced by Barducci et al to improve the biased potential43. In the well-tempered MT, the deposition rate for the biased potential is decreased by rescaling the Gaussian height (W) over the simulation time:

where VG(S,t) is the biased potential at the current position and current time, ΔT is a temperature-like parameter, and τG is the deposit ion stride. The underlying free energy is a scaled approximation to the VG(S,t), with

With respect to the standard MT, the biased potential decreases as 1/t when the simulation proceeds, allowing smooth converge to an approximation of F(s).
Simulation details: We carried out approximately 40 ns of well-tempered MT simulations for the systems of JDTic-wild μ/δ/κ-OR complexes. The crystal structure of inactive JDTic-bound κ-OR was directly used as a system for MT simulation, while the conformation of JDTic-wild μ/δ-OR complexes used for MT analysis was the frame at the final 100 ns of the unbiased MD simulation. For each system, the MT simulations were carried out one time using plumed 2.0244,45 implemented in Gromacs 4.6.5. The RMSD evolutions of the heavy atoms in each diffusing ligand were used as the first collective variable. The second collective variables were selected based on the binding experiments and our unbiased MD simulation. The residue D3.32 was proven to be indispensable for JDTic binding20, and then the Z-component of the vector connecting the protonated nitrogen of JDTic and two carboxyl oxygen atoms of D3.32 was thus selected as the second collective variable for JDTic egress. For the MT simulations, the biasing potential was added every 250 steps, with the width and height of the Gaussian hills set to 0.05 and 0.3 kJ/mol, respectively, and ΔT=2700.
Analysis of MD simulations
RMSD and RMSF (root-mean-square fluctuation) calculations, hydrogen bond and angle analyses, and distance evolutions were produced by the program Gromacs. The interval time of trajectory calculations, including RMSD, RMSF, angle, distance and principal component analysis, was 100 ps in these 100-ns simulations. RMSD values were calculated by comparison with the initial simulation conformations. All the smooth curves in Figures were fitting groups, which were identical to the corresponding Angle/RMSD calculations in color.
Results
Initial binding pockets of three OR subtypes with the ligand JDTic
In μ/δ/κ subtypes, the initial structure of ligand JDTic, extracted from the crystal structure of JDTic-κ-OR complex20, was stabilized in a V-shaped conformation, and its stretching angle was 88.5° (Figure 2). Although the three subtypes were all in the antagonistic state, residues within 4 Å of JDTic showed some structural differences among them (Figure 2A). For instance, residues at positions 2.63 and 7.35 were different from each other. In the μ/δ/κ subtypes, residues at position 2.63 were Asn, Lys and Val, while those at position 7.35 were Trp, Leu and Tyr, respectively. The orientation of some common residues, such as D3.32 and Y3.33, were similar in the μ/δ subtypes but different in the κ subtype. The differences might have resulted from the diverse ligand chemotypes because the original ligands in the crystal structures of the μ/δ subtypes belong to the family morphinans, which contain multiple rigid rings. With JDTic binding to the three OR subtypes, we built JDTic-wild type μ/δ/κ-OR systems to be used for the unbiased MD simulations.
Unbiased MD simulations on the three wild-type OR systems
In our MD simulations, the three complex systems reached equilibrium at approximately 30 ns as indicated by the protein RMSD values (Supplementary Figure S1A). Compared with the δ/κ subtypes after stabilization, the μ-OR subtype exhibited higher RMSD values, varying between 2.7 and 3.5 Å with respect to its initial structure. The δ-OR subtype showed the lowest RMSD changes, ranging from 2.0 to 2.7 Å. All the RMSDs could generally reflect the structural diversities among these subtypes, and the protein flexibility of μ-OR might be larger than the other two subtypes.
The ligand in the κ-OR system exhibited excellent stability, and ligand RMSD essentially fluctuated only by 0.4 Å, occasionally up to approximately 0.8 Å (Supplementary Figure S1B). In the μ/δ subtypes, the performance of the ligand was much unstable, with RMSD values even up to 2 Å. From the sampling of the final 100-ns complexes (Figure 3), we could see that the V-shaped conformation of the ligand was largely distorted in both the μ and δ subtypes. Comparing JDTic reverse trends and key residues within 4 Å of the ligand in these complexes, we could find that residue Q2.60 in the μ/δ subtypes adopted different orientations from that in the κ subtype. Because the Q2.60 residue is common among the three subtypes, we speculated that this conformational variety was more likely to be caused by residues Asn and Lys at the 2.63 positions, which were particularly close to Q2.60. To verify the abovementioned conjecture, we mutated 2.63-position residues to Val in the μ/δ subtypes according to the κ subtype, and then constructed JDTic-bound-mutated μ/δ-OR systems for additional MD simulations.
Key interactions within 4 Å of JDTic in JDTic-OR complexes at final 100 ns. The backbones of residues and JDTic were colored in the same way as mentioned in Figure 2.
Interaction modes of JDTic with the 2.63-position-mutated μ/δ-subtype
The average RMSD values of the receptors and ligands were lower in the mutant systems than in the corresponding wild-type ones (Figure 4A and 4B). The average RMSD value of the μ-mutant showed a great decrease, varying from an average of approximately 3.3 Å to 2.8 Å, while the RMSD change of the δ-mutant was not notable. Compared with the receptor change, the RMSD values of JDTic in the mutant systems decreased significantly. The average RMSD of JDTic in the μ-mutant was reduced to 0.45 Å from 1.6 Å, while that in the δ-subtype decreased from 1 Å to 0.5 Å, both of which were close to JDTic in the κ-subtype, especially that in the μ-mutant.
RMSD calculations of receptors (A) and JDTic (B) in unbiased JDTic-mutated OR systems. (C) The key interactions of JDTic-mutated OR complexes at the final 100 ns. The backbones of residues and JDTic were colored in forest green (mute-μ-OR), deep teal-cyan (mute-δ-OR) and slate-blue (JDTic). Hydrogen bond interactions are shown in the yellow dotted lines.
The number of hydrogen bonds (HBs) between JDTic and D3.32 in the mutant increased from 2 in the wild type to 3, accounting for 59.3% of the simulations, close to 63.1% in the κ-subtype (Table 2). However, the performance of the δ-mutant was relatively poor, and the hydrogen bonds changed from 3 to 2. From the samples at 100 ns (Figure 3, 4C and 4D) and the initial structure of κ-subtype (Figure 2), we could see that the 7.35-position residues of the μ-mutant and κ-subtype formed HBs with the carbonyl group on JDTic through water molecules, and the carbonyl group maintained a certain orientation. However, we did not find similar residues in this position of the δ-mutant or around to keep the carbonyl group under the same orientation (Figure 4D), and then the isoquinoline ring in the δ-mutant occurred in some tilt and away from D3.32. Therefore, in the case of JDTic remaining at a similar bending angle, it was very difficult for the ligand to form 3 hydrogen bonds with D3.32.
The bending angle of JDTic also attracted our attention. The stability of the ligand in the two mutants was better than that in the wild type, with an average angle of approximately 92°, which was close to that of the κ-subtype, approximately 88.5° (Figure 5A). However, the average angle of the μ-mutant, accounting for approximately 43%, close to that of the κ-subtype, 50%, was significantly higher than that of the δ-mutant, 25%. It was found that the Q2.60 residue was not stable in the δ-mutant (Figure 5B and Supplementary Table S1), especially at 10–65 ns. From the sampling of the two mutant systems at 100 ns and conformation of the Q2.60 residue in the κ-subtype, the residue was located at the position close to TM7. However, there was no suitable residue to form a stable water environment and then make the Q2.60 residue less steady in the δ-mutant, which should be the main factor leading to the large change in the ligand bending angle.
(A) Ligand angle calculations and distributions in all the five systems. The measured method is the same as that in Figure 2B. (B) The torsion angle values and distributions of Q2.60 in all unbiased JDTic-wild type/mutated OR systems.
Unbinding pathways of JDTic in the three wild-type OR subtypes
To further understand the interaction diversities between JDTic and the three subtypes, we performed well-tempered metadynamics analysis on ligand dissociation channels of the κ-subtype crystal structure and μ/δ-wild type at 100 ns among unbiased MD simulations.
In our MT simulations, JDTic left the antagonist binding pocket in the three OR subtypes and explored the binding pathways to exit the receptors through a series of conformational changes.
Representative conformations (Figure 6) obtained from the dissociation of each system showed that JDTic adopted different unbinding pathways in the three subtypes. In the μ-subtype, JDTic left the receptor from the TM2 region near the TM1 position. In the δ-subtype, JDTic realized the dissociation from the area between TM5 and TM6. In the κ-subtype, JDTic left the receptor from the TM7 region near the TM1 direction. In each unbinding pathway of the μ/δ/κ-subtype, the gating residue varied, namely, Y2.64, W6.58 and Y7.35, respectively. The 6.58 and 7.35 positions were “address” sites of ORs. Although Y2.64 is common for the three subtypes, the adjacent residue at position 2.63 was different and had a significant impact on the selectivity of JDTic as discussed above. Therefore, the differences in the gating residues could also reflect the ligand selectivity varieties of ORs.
Alignment of representative poses and gating residues in JDTic-μ-OR (A)/δ-OR (B)/κ-OR (C) complexes for the trajectory analysis of JDTic unbinding.
Discussion
In this study, we performed a series of MD simulations around the selectivity of JDTic on the three OR subtypes. The simulations showed that the 2.63 position was the key factor of the selective ligand JDTic on the μ-subtype. For the δ-subtype, the 7.35 position was the other crucial aspect of the JDTic subtype selectivity in addition to the 2.63-position residue. Binding affinity assays46 displayed that the binding affinity ratio of JDTic was 339 for the subtype κ/μ and 4935 for the subtype κ/δ. From these affinities, we could see the influence factors for the δ-subtype activity of JDTic were more complex than that in the μ-subtype. In addition, those JDTic analogs28,30 generally had high selectivity on the μ/δ-subtype, in particular δ-OR, and our results could also reveal this phenomenon well.
From the existing structural modification on JDTic analogs, we could find that the amino group of the isoquinoline ring was the key aspect for the activity of these analogs. For instance, the binding affinity of compound JPP-6 (Figure 1B) was decreased to 1/10 of JDTic due to the absence of the amino group30, which could be explained by the crystal structure, because the nitrogen atom was involved in hydrogen bonding with the D3.32 residue. At the same time, the loss of the nitrogen atom would increase the selectivity of the JPP-6 compound on the μ/δ-subtype, which should be closely related to the residues at the 2.63 and 7.35 positions of ORs and the stability of JPP-6. The deletion of the nitrogen atoms damaged the rigid structure of the isoquinoline ring, thus increasing the flexibility of JPP-6. The affinity varieties of other JDTic analogs might be related to the mode of action and other residues at different positions20.
Site-directed mutagenesis experiments20 demonstrated that D3.32 was the critical residue for the binding affinity of JDTic on the κ-subtype, while the 2.63 position basically did not form direct interaction with the κ-subtype. Mapped to the μ/δ-subtype, the 2.63-position residue became the vital factor influencing the JDTic binding affinity for the two subtypes. This mode of action was similar to that of the δ-OR-selective ligand NTI. All the results also reflected the diversity between the activity and selectivity mechanism to a certain extent.
Observed from the unbinding pathway of JDTic, the 2.63- and 7.35-position residues were located in the gating area, and participated in ligand recognition and dissociation activities. Therefore, it was supposed that “address” residues were not only closely related to the static interaction of ligands but also directly involved in the binding and dissociation process.
The importance of the 7.35- and 2.63-position residues for the selectivity of JDTic was the focus of work by Wu et al20 and Granier et al19, but they were consented only with the static perspective through molecular superposition and homology modeling. We started using the molecular dynamics method and carefully investigated the selectivity mechanism of JDTic for the three OR subtypes. In addition to the significance of the 2.63-position residues on JDTic selectivity, we also observed whether the efficacy of JDTic was affected by the dynamics of Q2.60, which was driven by the 2.63-position residue. These observations not only deepened our understanding of the selective mechanism of JDTic regarding three OR subtypes but also had some reference values for the mechanism elucidation of other selective opioids. At the same time, this virtual mutation method could save on experimental reagents and materials.
Here, we incidentally summarized the critical sites for OR subtype selectivity, including the 2.63 position (JDTic), ECL2 (U5048826), the 6.58 position (GNTIs23,24,25) and the 7.35 position (NTI21,25 and JDTic). These sites were indeed located at the upper part of the binding pocket; therefore, these positions were closely related to ligand identification and dissociation. This suggests that the interaction between a ligand and a target might be determined by the time of their encounter. Furthermore, the G-protein-biased agonist PZM2147, discovered in 2016, showed good selectivity on the three OR subtypes. The selectivity site was supposed to be at the 2.63 position from the docking mode, while the PZM21 ligand exhibited an excellent activation efficacy on the μ-subtype.
In opioid receptors, the mutation of some key residues could lead to a significant decline in ligand activity; these residues include D3.32 (JDTic20), E6.58 and I6.55 (5′-GNTI and Nor-BNI20), W6.58 (NTI21,25), and so on; in the other side, some mutated residues contributed to ligand functional changes. For instance, mutations of D2.50, N7.45 and N7.49 residues could reverse NTI from an antagonist to a β-arrestin protein-biased agonist32. The role of these two typical residues cannot be explained by the “message-address” concept32. Therefore, we suggest a new concept, namely, the “message-address-efficacy” hypothesis, to more appropriately interpret the relationships among the affinity, selectivity and function between ORs and opioids. In the new hypothesis, the “efficacy” site in a receptor is closely related to opioid activity and function similarly to the abovementioned residues, and these sites are both in consistency and diversity with those functioning with “message” and “address”. Taking the binding process of JDTic20 as an example, the D3.32 residue plays the “message” role and acts as an “efficacy” site; additionally, in the 5′-GNTI20 binding activity, the E6.58 residue is the critical “address” site, but the “efficacy” function is completed through the cooperation of E6.58 and I6.55 residues. In addition, the abovementioned residues32, closely correlated with the functional selectivity of ORs, should also be regarded as “efficacy” residues.
The concept of “message-address-efficacy” can not only promote our comprehensive understanding of residue functions on ORs but also update our recognition of the “message-address” concept proposed based on traditional opiates8,9,10, such as the indolyl group of NTI functioning both as “address” and “efficacy” sites21,25,32. Taking JDTic20 as an example again, three nitrogen atoms and areas around are the “efficacy” sites, while the hydroxyphenol position acts an “address” role. Considering the crucial interaction information of the opioid receptor, the thought of “message-address-efficacy” would be more suitable to be applied to obtain highly active and selective ligands. The idea can be used to design new corresponding ligands and modify existing opioids.
Currently, it was difficult to quantitatively identify the specific effect intensity of the 2.63 and 7.35 positions on three OR subtypes. We thought the related difference values would be further computed through combining free energy calculation methods. Certainly, it is necessary to build more systems to study the dissociation of ligands.
Conclusions
In this work, we studied the JDTic-induced subtype selectivity of μ/δ/κ-ORs through a series of computational simulations, including molecular dynamics and metadynamics methods. To clarify the related subtype selectivity mechanism in detail, we combined the simulated mutation method with our simulations.
Subtype selectivity mechanism of JDTic. We found the 2.63 position was the decisive factor of JDTic selectivity for the μ-subtype. However, for the δ-subtype, the 7.35 position was the other crucial aspect of JDTic subtype selectivity in addition to the 2.63-position. The two positions affected the efficacy of JDTic both through changing the dynamics of the Q2.60 residue. The 2.63- and 7.35-position residues are located in the gating area and participate in ligand recognition and dissociation activities among the unbinding process of JDTic.
“Message-address-efficacy” hypothesis. Based on the existing critical interaction mechanism, we proposed the “message-address-efficacy” hypothesis to interpret the relationships among the affinity, selectivity and function between ORs and opioids. Considering the receptor interaction mechanism, the above hypothesis is thought to be more suitable than the “message-address” concept in understanding the functions of opioids and their receptors.
Thus, our studies provide the detailed dynamics mechanism of OR-subtype selectivity, and the “message-address-efficacy” hypothesis might be helpful for deep understanding of the action of ORs and the new ligand design.
Author contribution
Jian-xin CHENG carried out the molecular dynamics simulations; Jian-xin CHENG and Yun TANG designed the study and analyzed the data; Yun TANG was responsible for the project; Jian-xin CHENG, Tao CHENG, Wei-hua LI, Gui-xia LIU, Wei-liang ZHU and Yun TANG contributed to writing and commenting on the manuscript.
References
Lagerstrom MC, Schioth HB . Structural diversity of G protein-coupled receptors and significance for drug discovery. Nat Rev Drug Discov 2008; 7: 339–57.
Lee SM, Booe JM, Pioszak AA . Structural insights into ligand recognition and selectivity for classes A, B, and C GPCRs. Eur J Pharmacol 2015; 763: 196–205.
Mallipeddi S, Janero DR, Zvonok N, Makriyannis A . Functional selectivity at G-protein coupled receptors: Advancing cannabinoid receptors as drug targets. Biochem Pharmacol 2017; 128: 1–11.
Zhou L, Bohn LM . Functional selectivity of GPCR signaling in animals. Curr Opin Cell Biol 2014; 27: 102–8.
Kolakowski LF Jr . GCRDb: a G-protein-coupled receptor database. Receptors Channels 1994; 2: 1–7.
Katritch V, Cherezov V, Stevens RC . Structure-function of the G protein-coupled receptor superfamily. Annu Rev Pharmacol Toxicol 2013; 53: 531–56.
Beaulieu JM . In vivo veritas, the next frontier for functionally selective GPCR ligands. Methods 2016; 92: 64–71.
Eberle A, Leukart O, Schiller P, Fauchere JL, Schwyzer R . Hormone--receptor interactions: [4-carboranylalanine, 5-leucine]-enkephalin as a structural probe for the opiate receptor. FEBS Lett 1977; 82: 325–8.
Chavkin C, Goldstein A . Specific receptor for the opioid peptide dynorphin: structure--activity relationships. Proc Natl Acad Sci U S A 1981; 78: 6543–7.
Schwyzer R . ACTH: a short introductory review. Ann N Y Acad Sci 1977; 297: 3–26.
Portoghese P, Sultana M, Takemori A . Design of peptidomimetic delta opioid receptor antagonists using the message-address concept. J Med Chem 1990; 33: 1714–20.
Portoghese PS, Sultana M, Takemori A . Naltrindole, a highly selective and potent non-peptide δ opioid receptor antagonist. Eur J Pharmacol 1988; 146: 185–6.
Portoghese PS . Bivalent ligands and the message-address concept in the design of selective opioid receptor antagonists. Trends Pharmacol Sci 1989; 10: 230–5.
Filizola M, Devi LA . Grand opening of structure-guided design for novel opioids. Trends Pharmacol Sci 2013; 34: 6–12.
Cox BM . Recent developments in the study of opioid receptors. Mol Pharmacol 2013; 83: 723–8.
Lipkowski AW, Tam SW, Portoghese PS . Peptides as receptor selectivity modulators of opiate pharmacophores. J Med Chem 1986; 29: 1222–5.
Mansour A, Taylor LP, Fine JL, Thompson RC, Hoversten MT, Mosberg HI, et al. Key residues defining the mu-opioid receptor binding pocket: a site-directed mutagenesis study. J Neurochem 1997; 68: 344–53.
Ballesteros JA, Weinstein H . Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci 1995; 25: 366–428.
Granier S, Manglik A, Kruse AC, Kobilka TS, Thian FS, Weis WI, et al. Structure of the delta-opioid receptor bound to naltrindole. Nature 2012; 485: 400–4.
Wu H, Wacker D, Mileni M, Katritch V, Han GW, Vardy E, et al. Structure of the human kappa-opioid receptor in complex with JDTic. Nature 2012; 485: 327–32.
Bonner G, Meng F, Akil H . Selectivity of mu-opioid receptor determined by interfacial residues near third extracellular loop. Eur J Pharmacol 2000; 403: 37–44.
Filizola M, Devi LA . Structural biology: how opioid drugs bind to receptors. Nature 2012; 485: 314–7.
Sharma SK, Jones RM, Metzger TG, Ferguson DM, Portoghese PS . Transformation of a kappa-opioid receptor antagonist to a kappa-agonist by transfer of a guanidinium group from the 5′- to 6′-position of naltrindole. J Med Chem 2001; 44: 2073–9.
Jones RM, Hjorth SA, Schwartz TW, Portoghese PS . Mutational evidence for a common kappa antagonist binding pocket in the wild-type kappa and mutant mu[K303E] opioid receptors. J Med Chem 1998; 41: 4911–4.
Metzger TG, Paterlini MG, Ferguson DM, Portoghese PS . Investigation of the selectivity of oxymorphone- and naltrexone-derived ligands via site-directed mutagenesis of opioid receptors: exploring the "address" recognition locus. J Med Chem 2001; 44: 857–62.
Metzger TG, Ferguson DM . On the role of extracellular loops of opioid receptors in conferring ligand selectivity. FEBS Lett 1995; 375: 1–4.
Thomas JB, Atkinson RN, Rothman RB, Fix SE, Mascarella SW, Vinson NA, et al. Identification of the first trans-(3R,4R)-dimethyl-4-(3-hydroxyphenyl)piperidine derivative to possess highly potent and selective opioid κ receptor antagonist activity. J Med Chem 2001; 44: 2687–90.
Mitch CH, Quimby SJ, Diaz N, Pedregal C, de la Torre MG, Jimenez A, et al. Discovery of aminobenzyloxyarylamides as kappa opioid receptor selective antagonists: application to preclinical development of a kappa opioid receptor antagonist receptor occupancy tracer. J Med Chem 2011; 54: 8000–12.
Carroll FI . Carlezon WA Jr. Development of kappa opioid receptor antagonists. J Med Chem 2013; 56: 2178–95.
Thomas JB, Atkinson RN, Vinson NA, Catanzaro JL, Perretta CL, Fix SE, et al. Identification of (3R)-7-hydroxy-N-((1S)-1-[[(3R,4R)-4-(3-hydroxyphenyl)- 3,4-dimethyl-1-piperidinyl]methyl]-2-methylpropyl)-1,2,3,4-tetrahydro- 3-isoquinolinecarboxamide as a novel potent and selective opioid kappa receptor antagonist. J Med Chem 2003; 46: 3127–37.
Manglik A, Kruse AC, Kobilka TS, Thian FS, Mathiesen JM, Sunahara RK, et al. Crystal structure of the micro-opioid receptor bound to a morphinan antagonist. Nature 2012; 485: 321–6.
Fenalti G, Giguere PM, Katritch V, Huang XP, Thompson AA, Cherezov V, et al. Molecular control of delta-opioid receptor signalling. Nature 2014; 506: 191–6.
Spassov VZ, Flook PK, Yan L . LOOPER: a molecular mechanics-based algorithm for protein loop prediction. Protein Eng Des Sel 2008; 21: 91–100.
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, et al. CHARMM general force field: A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 2010; 31: 671–90.
Vanommeslaeghe K . MacKerell AD Jr. Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J Chem Inf Model 2012; 52: 3144–54.
Vanommeslaeghe K, Raman EP . MacKerell AD Jr. Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J Chem Inf Model 2012; 52: 3155–68.
Lomize AL, Pogozheva ID, Mosberg HI . Anisotropic solvent model of the lipid bilayer. 2. Energetics of insertion of small molecules, peptides, and proteins in membranes. J Chem Inf Model 2011; 51: 930–46.
Lomize MA, Pogozheva ID, Joo H, Mosberg HI, Lomize AL . OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res 2012; 40: D370–6.
Klauda JB, Venable RM, Freites JA, O'Connor JW, Tobias DJ, Mondragon-Ramirez C, et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J Phys Chem B 2010; 114: 7830–43.
Humphrey W, Dalke A, Schulten K . VMD: visual molecular dynamics. J Mol Graph 1996; 14: 33–8.
Hess B, Kutzner C, van der Spoel D, Lindahl E . GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 2008; 4: 435–47.
Barducci A, Bonomi M, Parrinello M . Metadynamics. Wiley Interdiscip. Rev Comp Mol Sci 2011; 1: 826–43.
Barducci A, Bussi G, Parrinello M . Well-tempered metadynamics: a smoothly converging and tunable free-energy method. Phys Rev Lett 2008; 100: 020603.
Laio A, Parrinello M . Escaping free-energy minima. Proc Natl Acad Sci U S A 2002; 99: 12562–6.
Tribello GA, Bonomi M, Branduardi D, Camilloni C, Bussi G . PLUMED 2: New feathers for an old bird. Comput Phys Commun 2014; 185: 604–13.
Rorick-Kehn LM, Witkin JM, Statnick MA, Eberle EL, McKinzie JH, Kahl SD, et al. LY2456302 is a novel, potent, orally-bioavailable small molecule kappa-selective antagonist with activity in animal models predictive of efficacy in mood and addictive disorders. Neuropharmacology 2014; 77: 131–44.
Manglik A, Lin H, Aryal DK, McCorvy JD, Dengler D, Corder G, et al. Structure-based discovery of opioid analgesics with reduced side effects. Nature 2016; 537: 185–90.
Acknowledgements
All the simulations were performed using the high-performance cluster kindly provided by Prof Bang-ce YE of the State Key Laboratory of Bioreactor Engineering, East China University of Science and Technology. This work was supported by the National Key Research and Development Program of China (Grant 2016YFA0502304), the National Natural Science Foundation of China (Grants 81673356 and U1603122) and the 111 Project (Grant B07023).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary files are available at the website of Acta Pharmacologica Sinica.
Supplementary information
Supplementary Information
Supplementray Figures and table (DOC 2321 kb)
Rights and permissions
About this article
Cite this article
Cheng, Jx., Cheng, T., Li, Wh. et al. Computational insights into the subtype selectivity and “message-address-efficacy” mechanisms of opioid receptors through JDTic binding and unbinding. Acta Pharmacol Sin 39, 482–491 (2018). https://doi.org/10.1038/aps.2017.132
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2017.132