Abstract
Highly active antiretroviral therapy (HAART) can effectively suppress the replication of human immunodeficiency virus-1 (HIV-1) and block disease progression. However, chronic HIV-1 infection remains incurable due to the persistence of a viral reservoir, including the transcriptionally silent provirus in CD4+ memory T cells and the sanctuary sites that are inaccessible to drugs. Reactivation and the subsequent elimination of latent virus through virus-specific cytotoxic effects or host immune responses are critical strategies for combating the disease. Indeed, a number of latency reactivating reagents have been identified through mechanism-directed approaches and large-scale screening, including: (1) histone deacetylase inhibitors (HDACi); (2) cytokines and chemokines; (3) DNA methyltransferase inhibitors (DNMTI); (4) histone methyltransferase inhibitors (HMTI); (5) protein kinase C (PKC) activators; (6) P-TEFb activators; and (7) unclassified agents, such as disulfram. They have proved to be efficacious in latent cell line models and CD4+ T lymphocytes from HIV-1-infected patients. This review comprehensively summarizes the recent progress and relative challenges in this field.
Similar content being viewed by others
Introduction
During the last three decades, highly active antiretroviral therapy (HAART) has remarkably decreased the morbidity and mortality of HIV-infected patients. However, HAART fails to eradicate latent viral reservoirs in vivo. AIDS remains an incurable disease because the cessation of treatment inevitably leads to a rapid rebound of viremia, which is associated with the existence of a viral reservoir. A significant proportion of patients fail to maintain an undetectable plasma viral load (<50 copies/mL) because of a lack of adherence and/or the emergence of drug resistance1,2. Despite having a viral load below a detectable level, many patients also experience co-morbidities, such as cardiovascular diseases, bone disorders and cognitive impairments3. Moreover, immune functions fail to be fully restored after HAART treatment in patients chronically infected with HIV-14. The persistence of resting CD4+ T cells harboring a transcriptionally silent, yet replication-competent, provirus represents a major barrier to eradication. Reactivation of latent proviruses results in the rapid progression of AIDS. Recently, several new therapeutic approaches have been proposed that aim to control (functional cure) or eliminate (sterilizing cure) the viral reservoir, including novel drugs, gene therapy, bone marrow transplantation (BMT), immune-modulation and combination approaches5,6,7. The most widely discussed approach for eliminating the latent reservoir is the so-called “shock and kill” strategy. Specifically, this strategy seeks to activate HIV-1 from latent reservoirs using latency-reversing agents and then to target HIV-1-infected cells for eradication via the host immune response and/or cytotoxic drugs. Recently, a number of latency-reversing agents have been identified. The different anti-latency molecules have multiple different pharmacological targets. The molecular mechanisms responsible for latency are multiply determined; therefore, we will briefly introduce the transcriptional mechanisms governing latency, which are highly relevant for current anti-latency compounds.
Molecular mechanisms responsible for the development of HIV-1 latency
The major barrier to the eradication of HIV-1 is the establishment of a latent reservoir in different anatomical sites throughout host bodies. Resting CD4+ T cells in blood are considered to be the predominant viral reservoir. Moreover, other anatomical compartments, including the central nervous system (CNS), gut-associated lymphoid tissue, bone marrow and genital tract, are also believed to act as reservoirs for persistently infected cells. Latency happens at a very low frequency; only a small number of resting CD4+ T cells (approximately one million cells) harbor replication-competent latent provirus, which can be activated by mitogens8. Silencing of active infection in activated T cells and immediate silent integration in resting T cells both contribute to the establishment of latency, which has been demonstrated in vitro by several research groups9,10,11,12. The molecular mechanisms associated with latency are not completely understood13,14,15,16. In general, latency can be divided into pre-integration latency and post-integration latency; the latter accounts for the establishment of the latent viral reservoir. Latency can be maintained at transcriptional levels (eg, lack of transcriptional activators, condensed chromatin structure) or at post-transcriptional levels (eg, inhibition of nuclear RNA transport and inhibition of translation by microRNA), and it is generally assumed that maintenance at the transcriptional level is the case for the majority of latently infected cells. Molecular mechanisms targeting transcription consist of both trans-effects and cis-effects, including: (1) the chromatin environment around the integration site and transcriptional interference17,18; (2) the epigenetic control of the HIV-1 promoter or the presence of repressive nucleosomes (nuc-0 and nuc-1)19,20; (3) lack of crucial transcriptional factors, such as NF-κB or nuclear factor of activated T cells (NFAT)21, sub-optimal concentrations of transcriptional activators such as Tat22,23 or the presence of transcriptional suppressors such as CTIP2 (COUPTF Interacting Protein 2)24,25, DSIF (DRB-Sensitivity Inducing Factor)26 and NELF (Negative Elongation Factor)27; (4) the sequestration of positive transcriptional elongation factor b (P-TEFb) in its inactive form with HEXIM [hexamethylene bisacetamide (HMBA)-induced protein 1] and 7SK snRNA (small nuclear RNA)28,29,30.
Two nucleosomes (nuc-0 and nuc-1) that assemble around transcription sites harbor markers of silent heterochromatin, such as low acetylation of histones, lysine 9 trimethylated histone 3 (H3K9me3) and heterochromatin protein 1 (HP1)31. Disruption of nucleosomes by acetylation upon recruitment of histone acetyltransferase allows viral transcriptional activation to occur. Similarly, the reversible methylation of histone or CpG islands around the 5′ long terminal repeat (LTR) also regulates the activity of the HIV promoter epigenetically32.
It is worth noting that the CNS has several unique features that may result in differences in viral transcription and how latency is established. These include CNS-specific cell types, altered LTR activity, unique transcription factors and unique epigenetic factors. A large body of evidence has suggested that perivascular macrophages, microglia, and astrocytes, which are all susceptible to HIV infection, represent possible latent reservoirs in the CNS33. Phylogenetic and functional analyses showed reduced basal transcriptional activity of CNS-derived LTRs in astrocytes and T cells compared to that of non-CNS-derived LTRs, which most likely contributes to HIV-1 latency in the brain34. Unique transcription factors are also considered to be closely associated with HIV-1 latency in the brain. The most important transcription factors relevant to HIV transcription include Sp1-4, NF-κB, AP-1, and C/EBP. An abundance of Sp3 relative to Sp1 in astrocytes can, in some part, explain the restriction of HIV-1 transcription within these cells35. In addition, Desplats and his colleagues reported that levels of BCL11B, heterochromatin protein 1 (HP1)α, methyl CpG binding protein 2 (MeCP2), and histone deacetylase 1 (HDAC1) were significantly increased in CNS cells latently infected with HIV-1, favoring the idea that epigenetic factors are involved in silencing mechanisms36.
Therapeutics used in reactivating latent virus
In recent years, small molecular compounds targeting different mechanisms involved in maintaining latency have been identified. These can be categorized into the following groups, mainly based on pharmacological targets (Figure 1): (1) histone deacetylase inhibitors (HDACi); (2) cytokines and chemokines; (3) DNA methyltransferase inhibitors (DNMTI); (4) histone methyltransferase inhibitors (HMTI); (5) protein kinase C (PKC) activators; (6) P-TEFb activators; and (7) unclassified agents, such as disulfram9. These compounds have been suggested as agents to reactivate HIV-1 and eradicate viral reservoirs. In addition, compounds with known pharmacological targets have also been found in large-scale screenings. We summarize the clinical application of several typical reactivating agents below.
Multifactorial mechanisms governing HIV-1 latency and latency-reversing agents based on their pharmaceutical targets. Different compounds have been identified that reactivate HIV-1 transcription. In the cytoplasm, PKC agonists such as prostratin and brystatin 1 can activate PKC, which leads to the subsequent phosphorylation and degradation of NF-κB inhibitor IκB-α, followed by the freeing of p65/p50 and accumulation of p65/p50 in the nucleus. The binding of p65/p50 to the HIV LTR enhances initial transcription. Disulfram promotes the degradation of PTEN and upregulates HIV-1 transcription upon activation of the Akt pathway. Cytokines such as IL-7 and IL-15, which are involved in the JAK/STAT pathway, also have a positive effect on latent virus reactivation. In the nucleus, the recruitment of the P-TEFb complex, comprised of CDK9 and cyclin T1, by HIV Tat is the prerequisite for transcription initiation. JQ1, by binding to Brd4, can release P-TEFb and thereby promote transcription elongation. HDACi inhibits deacetylation of histones and non-histone transcription factors, allowing for acetylation by the HATs, which is required for a more favorable chromatin structure. In addition, DNMTIs, of which Aza-CdR is the prototype, and HMTIs, including DNZep and chaetocin, reactivate HIV-1 transcription by targeting DNMT and HMT, respectively.
Bromodomain and extra-terminal (BET) inhibitors
JQ1 is a cell-permeable compound with potent anti-cancer activity37. It interacts with bromodomains, exhibiting the highest specificity in Brd4. Recently, several lines of evidence indicated that JQ1 reactivates latent HIV in both the clonal and primary cell model of HIV latency, providing evidence for its multifunctional utility22,36. Specifically, it inhibits recognition of acetylated lysines relevant to actively transcribed chromatin by binding to the bromodomain-binding pocket. When transcription initiates, the P-TEFb complex, comprised of cyclin-dependent kinase 9 (CDK9) and its regulatory partner cyclin T1 (CycT1), is recruited onto the HIV TAR loop through interactions with HIV Tat. P-TEFb is maintained in a dynamic equilibrium between its inactive form when associated with the 7SK RNA complex and its active form when recruited to TAR by Tat. JQ1 is thought to increase the availability of P-TEFb by freeing it from its complex with 7SK RNA or Brd4. Notably, JQ1 and an HDACi or PKC agonist synergistically reactivated latent HIV in HAART treated patients, but JQ1 treatment alone did not22.
HDACi
Histones are reversibly acetylated by histone acetyltransferases (HATs) and deacetylated by histone deacetylases (HDACs), which can be divided into three main classes based on their homologs in yeast. Class I includes HDAC-1, -2, -3, and -8. Class II comprises HDAC-4, -5, -6, -7, -9, and -10. HDAC-11 has the properties of both Class I and Class II HDACs and therefore belongs to an individual group. Histone acetylation influences chromatin architecture at transcription sites and is crucial for active gene expression. Lysines 9 and 14 of histone 3 (H3K9, H3K14), and lysine 16 of histone 4 (H4K16) are generally involved in this process. Acetylation of the histone tail is believed to decrease the compaction of chromatin thereby increasing the accessibility of chromatin to transcription factors and co-activators. HDAC maintains HIV latency by deacetylating histones directly at provirus-integrated sites or indirectly by inducing deacetylation of non-histone proteins, such as NF-κB. Accordingly, reactivating latent HIV by inhibiting HDAC represents a practicable strategy for the eradication of the latent HIV reservoir. Furthermore, intensive investigations of the pharmaceutical and toxicological properties of HDACis as anti-cancer drugs make these molecules the most advanced anti-latency agents in clinical trials. HDACis are classified into four major structural families: short-chain aliphatic acids (including valproic acid and butyric acid), hydroxamic acids (such as trichostatin A and vorinostat), benzamides (including entinostat), and cyclic tetrapeptides and depsipeptides (such as trapoxin B and romidepsin). These four classes of HDACis share the same molecular structure, consisting of a cap for HDAC surface recognition, a hydrophobic aliphatic linker 5–6 carbons in length, and a head comprising a functional group to chelate the zinc cation in the catalytic center. This functional group can be a hydroxamic acid, a benzamide group, a carboxylic acid group or an epoxide group. Various HDACis were shown to reactivate the provirus in experimental systems or in primary cells isolated from patients under antiretroviral therapy (ART). Below, we introduce several typical HDACis that are either in ongoing clinical trials or approved by the FDA.
Short chain aliphatic acids
Butyric acid and valproic acid (VPA) belong to this category. VPA is a carboxylate HDAC inhibitor that is prescribed for seizures and psychiatric disorders. It was the first HDACi proposed to reverse HIV latency. However, VPA is weak and nonspecific in inducing HIV-1 out of latency in vitro. Clinical trials using a combination of HAART and VPA produced conflicting results. In the first study by Lehrman et al, the frequency of resting cell infection was significantly reduced after VPA was added into ART38. In the subsequent clinical studies, no detectable changes in the size of the latent reservoir were observed between patients receiving HAART alone and patients receiving HAART plus VPA39,40. Limited effects of VPA on latently infected resting CD4+ T cells were observed in a prospective study41, and these effects waned over time. These studies clearly demonstrated that VPA, with or without HAART, was of limited value in decreasing the reservoir of latently infected resting T cells. Nevertheless, other HDACis of high specificity and tolerability should be explored.
Hydroxamic acids: SAHA/vorinostat, givinostat (ITF2357) and panobinostat (LBH-589)
To date, suberoylanilide hydroxamic acid (SAHA/vorinostat, VOR) is the best studied HDAC inhibitor with respect to HIV latency. It has been named vorinostat and is currently approved by the FDA for the treatment of cutaneous T cell lymphoma. SAHA reactivates latent viruses in a variety of in vitro cell models of HIV latency, including ACH-2, U1, and J-LAT and in resting CD4+ T cells isolated from ART patients. This evidence constitutes a proof-of-concept that clinical exposure to SAHA might disrupt latent infection42,43. However, whether treatment with SAHA will lead to reduction of the latent viral reservoir remains unknown. Moreover, two other promising HDAC inhibitors are givinostat and panobinostat, which have been demonstrated to display potency in HIV infected cells44,45. Notably, givinostat decreases expression of the HIV co-receptors CXCR4 and CCR5 on the cell surface of infected CD4+ T cells, and panobinostat has displayed a potency superior to that of multiple other HDACis, including VPA, vorinostat, givinostat, and belinostat45.
Cyclic tetrapeptides and depsipeptides
Romidepsin (RMD; Istodax) is a natural cyclic peptide isolated from Chromobacterium violaceum that has been approved by the FDA to treat cutaneous T cell lymphoma. In a comparison with vorinostat, RMD was superior in the following aspects: 1) In an in vitro latency model, RMD displayed an EC50 value of 4.5 nmol/L and a CC50 value of 100 nmol/L, resulting in an approximately 20-fold selectivity window, whereas vorinostat was substantially less potent, with EC50 and CC50 values of 4 μmol/L and 25 μmol/L, respectively. RMD is also the most potent inducer of HIV with the lowest EC50 value to date; 2) The activation of HIV transcription is more durable with RMD than with vorinostat46. In both resting and memory CD4+ T cells isolated from combination antiretroviral therapy (cART)-treated patients, a 4-h exposure to 40 nmol/L RMD induced, on average, a 6-fold increase in intracellular HIV RNA levels, whereas a 24-h treatment with 1 mmol/L VOR resulted in 2- to 3-fold increases46. RMD is a more robust inducer of HIV expression in latently infected cells than all other HDACis in clinical tests46.
Benzamide HDAC inhibitors
Entinostat (MS275) and mocetinostat (MGCD0103) also induce HIV expression from in vitro HIV-latently infected cell lines and primary cell models. Entinostat mainly targets HDACis (HDAC 1, 2, 3), which are the HDACs most associated with HIV regulation. Mocetinostat also targets HADC 1 and is currently undergoing clinical trials for Hodgkin lymphoma47. Entinostat exhibits its highest potency against HDAC 1 (nanomolar range) and is less potent against HDACs 2 and 3 (micromolar range). Currently, entinostat is being evaluated in 23 Phase I or II trials for a range of malignancies and has been demonstrated to be well-tolerated and negative in the Ames test. In a mouse renal cancer model, entinostat has been shown to have suppressive activity for regulatory T cell function, which will be beneficial for eliminating HIV latency48. In latently infected CCL19-treated primary T cells and the ACH2 cell line, entinostat led to higher virus production than SAHA49. Taken together, benzamide HDACis, with entinostat as the prototype, are an attractive novel option for future clinical trials.
Collectively, HDACis are promising and efficacious agents to reactivate latent HIV, and several related regimens are being administered in clinical trials. Recent efforts have focused on developing novel HDACis with high specificity for a unique HDAC, as such inhibitors would be likely to have fewer “off-target” effects.
DNA methyltransferase inhibitors (DNMTI)
In an effort to identify novel mechanisms governing latency, methyl-CpG binding domain protein 2 (MBD2) was identified by a cDNA screen in the J-Lat cell model as a transcriptional repressor closely associated with latency. The HIV-1 promoter is hypermethylated at two CpG islands surrounding the HIV transcriptional start site. MBD2 binds specifically to methylated DNA and consequently recruits HDAC, which leads to chromatin remodeling and histone deacetylation. 5-Aza-2′-deoxycytidine (Aza-CdR), which is approved for use in humans to treat myelodysplastic syndrome50, was shown to reactivate latent virus on its own and to have an intensified synergistic effect when used in combination with NF-κB activators51. Aza-CdR plus tumor necrosis factor (TNF)α activated latent virus replication at least twice as well as TNFα alone in J-Lat cells. LTR dimethylation was observed after Aza-CdR treatment, which correlated with a synergistic increase in proviral activation52. However, Aza-CdR treatment led to complicated results depending upon the cell line used. For example, in J-Lat 10.6, ACH-2 and U1 cell lines, the combination of Aza-CdR and TNFα did not increase or maintain activation but instead decreased activation, most likely because of a detrimental effect of Aza-CdR when used in some latently infected cells52. Overall, the ability of Aza-CdR to activate HIV replication exhibited a strong cell line dependence, highlighting the necessity for fairly elaborate combinations of latency activators and co-activators, and careful optimization of anti-HIV strategies.
Histone methyltransferase inhibitors (HMTI)
Histone H3 lysine 9 (H3K9) methylation plays an important role in HIV transcriptional regulation. H3K9 trimethylation (H3K9me3), mediated by Suv39H153; H3K27 trimethylation (H3K27me3)54 and H3K9 dimethylation (H3K9me2), mediated by G9α19 lead to HIV transcriptional silencing in different cell models, including peripheral blood mononuclear cells (PBMCs) from infected individuals. Therefore, HMTIs could also be used to purge HIV-1 latent reservoirs. Three specific inhibitors of lysine methyltransferase have been developed: chaetocin, which specifically inhibits SUV39 H1; and 3-deaza-neplanocin A (DZNep) and BIX-01294, which act on the G9α HMT.
SUV39H1 has been implicated in maintaining HIV latency in microglial cells through its interaction with CTIP-2 and HP1 γ53. Similarly, G9α was also proposed to be involved in the establishment of HIV latency19. Chaetocin belongs to the 3–6-epi-dithio-diketopiperazines, which have been suggested to participate in immunosuppression55 and anti-inflammation56. As a specific SUV39H1 inhibitor, chaetocin was observed to cause a 25-fold increase in latent HIV expression without significant toxicity or global T cell activation in Jurkat T cells57. Subsequently, another group observed that chaetocin induced HIV-1 recovery in 50% of CD8-depleted PBMC cultures and in 86% of resting CD4+ T cell cultures isolated from HIV-1-infected, HAART-treated patients32. Furthermore, the combinations of chaetocin+SAHA and chaetocin+prostratin have higher potential than the compound alone to reactive HIV replication from latent CD4+ resting T cells from patients32. BIX-01294, as a diazepin-quinazolin-amine derivative, selectively suppressed the enzymatic activity of histone lysine methyltransferase (HKMT) G9α by binding to its SET catalytic domain. BIX-01294 reactivated HIV-1 expression in 80% of resting CD4+ T cell cultures isolated from similar patients32. However, in another experimental system, BIX-01294 was a comparatively poor inducer of latent proviruses and was only able to induce 21.1% of the latent proviruses in the E4 cell line after overnight exposure to the drug54. The broad-spectrum agent DZNep was reported to downregulate several HKMTs and to target EZH258. Exposure of cells to 10 μmol/L DZNep led to global reductions in H3K27me3 (44% reduction) and H3K9me2 (70% reduction), accompanied by a reactivation of latent virus. A synergistic effect was observed when DZNep and SAHA were combined54. The selective combination of DZNep and an HDACi provided the most effective means to reverse the epigenetic gene silencing; therefore, DZNep, which targets H3K27me3 formation, has great potential as a selective inducer of latent HIV-1 proviruses.
Protein kinase C activators
Prostratin
Prostratin, a non-tumor-promoting phorbol ester, was reported to antagonize HIV latency in J-Lat cells, the SICD-hu Thy/Liv system, and the blood of patients with HAART-suppressed HIV. Importantly, unlike other related phorbol esters (eg, phorbol-12-myristate-13-acetate and 12-deoxyphorbol-13-phenyl acetate (DPP)), prostratin is not a tumor promoter or an irritant. Furthermore, prostratin even protects against tumor-promoting effects of these other agents, making it a good candidate for HIV therapy59,60. In addition, prostratin elicits pleiotropic effects relevant to HIV treatment. It causes downregulation of CD4, CXCR4 and, in some cases, CCR5 and thereby protects CD4+ T cells from infection61,62. Further investigation revealed that prostratin induces PKC-mediated phosphorylation, and the degradation of IκB and enables the subsequent release and penetration of transcriptional factor NF-κB into the cell nucleus, where NF-κB binds to LTRs, leading to transcriptional initiation63. However, research on prostratin has relied exclusively on plant sources, which have low isolation yields. Beans et al designed and synthesized prostratin analogs that outperformed native prostratin. These analogs exhibited higher affinity for PKC, with Ki values of less than 2 nmol/L. More importantly, these analogs induced latent virus 20-fold more potently than the preclinical leading drug, prostratin, in latently infected U1 cell lines. Significantly, a potent lead analog also induced the expression of latent virus from PBMCs isolated from HAART-treated patients at the concentration at which prostratin lost its efficacy. These potent analogs will facilitate the research and preclinical advancement of HIV/AIDS eradication64.
Ingenol B (IngB)
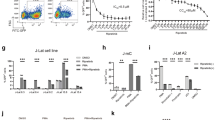
A recent study found that a newly modified ingenol ester compound originally isolated from Euphorbia tirucalli, Ingenol B (IngB), reactivated latent viruses in J-Lat A1 cell lines and purified CD4+ T cells from HIV-1 infected patients under long-term HAART65. IngB can effectively promote HIV transcription through activation of the protein kinase C (PKC) δ-Serine 664-NF-κB pathway or through a direct increase in NF-κB expression. Furthermore, IngB has minimal cell toxicity and is more potent than prostratin, SAHA and JQ1 in reactivating latent infection in vitro. A remarkable synergistic effect of IngB with JQ1 or HMBA in vitro would lead to a novel strategy in viral eradication.
In a recent report, a new semi-synthetic ingenol ester, 3-caproyl-ingenol (ING B), was shown to reactivate viral transcription in primary resting T cells by up to 12-fold upon activation of the PKC isoform. ING B outperforms a variety of current activators, such as SAHA, ingenol 3, 20-dibenzoate, PMA and HMBA and has a synergistic effect with SAHA66.
Cytokines and chemokines
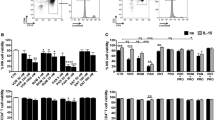
Cytokines have been proposed to act not only as an alternative immunomodulatory therapy to HAART but also to activate latent proviruses and increase the turnover rate of the latent reservoir. Therefore, several cytokines, eg, interleukin (IL)-2, IL-7, and IL-15, which share the same γ chain, have putative roles in latent reservoir clearance. The first attempt to purge a latent pool of HIV-1-infected CD4+ T cells was performed using IL-2 and other T cell activators, such as anti-CD3 antibody (OKT3). However, these agents induced global activation of T cells, which caused unacceptable toxicities and failed to reduce the latent reservoir67,68. In an investigation of the ability of IL-7 to purge latent virus, IL-7 was shown to have a greater capacity to activate latent reservoir from peripheral blood mononuclear cells (PBMCs) isolated from infected patients and CD4+ T lymphocytes than either IL-2 alone or IL-2 combined with phytohemagglutinin (PHA)69. Notably, IL-7 reactivated different provirus quasispecies than IL-2, indicating that different activators of proviral latency may perturb and deplete only a specific portion of proviruses in virally infected individuals.
Positive transcription elongation factor b activators
Hexamethylene bisacetamide (HMBA) is a hybrid bipolar compound that is a potent inducer of differentiation of various transformed cells and induces apoptosis70,71. Vlach et al observed that HMBA reactivated latent HIV-1 provirus without any detectable changes in the binding patterns of nuclear proteins to the enhancer or leader sequences of the HIV-1 LTR in cells of T lymphocytic and monocytic origin72. Subsequent mechanistic studies revealed that HMBA activates Akt via phosphatidylinositol-3-kinase (PI3K), leading to the phosphorylation of HEXIM-1 and the resultant release of active P-TEFb from its transcriptionally inactive complex with HEXIM-1 and 7SK snRNA72. Notably, HMBA induces HIV-1 expression in latent infected CD4+ resting memory T cells from aviremic patients under treatment with ART73. The nuclear content of CDK9 increased, allowing recruitment of CDK9 by Sp1 to the HIV-1 LTR. Concomitantly, the C-terminus of RNA polymerase II was hyperphosphorylated at the Ser2 and Ser5 residues, which is consistent with its roles in transcriptional initiation and elongation73. This evidence indicated that HMBA may be a candidate to reactivate latent virus.
Unclassified agents
Disulfram (bis(diethylthiocarbamoyl) disulfide) is an FDA-approved drug used to treat alcoholism. It can inhibit aldehyde dehydrogenase and increase acetaldehyde levels in patients. Xing et al found that disulfram reactivated latent HIV-1 in a Bcl2-transduced CD4+ T cell model without inducing global T cell activation74. However, the molecular mechanism of in vitro disulfram-induced HIV latency reactivation is ambiguous. A recent report found that disulfram induced-degradation of tensin homolog (PTEN) protein might contribute to upregulation of HIV-1 transcription through the Akt signaling pathway in U1 cell lines75. Importantly, only the parent compound and its first metabolite, diethyldithiocarbamic acid (DDTC), can reactivate latent HIV-1, whereas subsequent metabolites, such as diethyldithiocarbamate methyl ester (DDTC-Me), lose their efficacy. In a pilot clinical trial in which disulfram was administered for 14 d to HIV-1 infected individuals on ART, no significant changes in either the residual viremia or in the size of the latent reservoir were observed76. Strikingly, the pharmacokinetics and pharmacodynamics of disulfram were highly variable among subjects. The intersubject variability suggested that higher doses of disulfram might be more effective, and combining disulfram with other latency-reversing agents might have a more pronounced effect on the eradication of the reservoir.
In addition to disulfram, a number of compounds have been identified by large-scale screening of chemical and siRNA libraries in in vitro cell models of HIV latency; however, their mechanisms of action remain elusive. Two classes of derivatives of quinoline-8-ol; Mannich adducts of 5-chloroquinolin-8-ol and quinolin-8-yl carbamates were discovered to reactivate HIV-1 expression without causing T cell activation using a high-throughput system (HTS)77. Furthermore, Micheva-Viteva et al discovered antiviral 6 (AV6) using cell-based HTS assays. In subsequent studies, they demonstrated that AV6 required NFAT to reactivate latent infection, and demonstrated the broad spectrum of its latency-reversing activity in a variety of cells, including lymphocyte-based clonal cell lines and latently infected patient cells78.
Summary
The extensive current studies of molecular mechanisms associated with HIV latency have expanded our knowledge of new drug targets, allowing for the development of novel anti-latency therapeutics. Several groups of latency-reversing compounds have been identified and tested in laboratories and preclinical trials. However, the tested eradication cures so far have yielded complicated or unsatisfactory results, and several setbacks have occurred. A major obstacle in evaluating new latency-reversing agents is the lack of a suitable model representing latency in vivo. Currently available cell lines and several primary cell models fail to fully mimic the process in AIDS patients. Therefore, to provide a standard representation of persistent HIV-1 infection to evaluate new agents, non-human primate animal models can be used because these models faithfully represent the in vivo latently infected cells in the ART setting. Another unresolved issue in this strategy is the poor capacity to reduce the latent reservoir in clinical trials when a single reactivating agent is used. In addition, single agents likely purge only a highly specific quasispecies of latent virus or purge virus from only specific cell types. Therefore, a combination of several latency reactivating agents is required to obtain a satisfactory result. Another overarching question is whether the increase in viral protein expression in reactivated cells induced by these compounds will lead to viral cytopathogenicity and/or elimination by cytotoxic T cells. To this end, Siliciano et al recently reported that unstimulated HIV-specific cytotoxic T cell lymphocytes (CTLs) failed to kill the latently infected cells reactivated by SAHA. In contrast, antigen-specific stimulation of patients' CTLs led to rapid killing of these SAHA-reactivated cells79. Moreover, monocytes, macrophages and myeloid dendritic cells are more resistant to viral cytotoxic effects. Therefore, significant boosting of the CTL response before viral reactivation represents a necessary supplemental strategy for the latency-reversing approach.
Despite the hurdles in HIV-1 eradication in vivo, remarkable progress has been made in this field. In particular, novel strategies utilizing new anti-latency agents have been proposed and are in both preclinical and clinical trials, which will enable the field to move closer to effective clearance of HIV-1 in the future.
References
Sluis-Cremer N . The emerging profile of cross-resistance among the nonnucleoside HIV-1 reverse transcriptase inhibitors. Viruses 2014; 6: 2960–73.
Liu H, Ma Y, Su Y, Smith MK, Liu Y, Jin Y, et al. Emerging trends of HIV drug resistance in Chinese HIV-infected patients receiving first-line highly active antiretroviral therapy: a systematic review and meta-analysis. Clin Infect Dis 2014; 59: 1495–502.
Waters L, Patterson B, Scourfield A, Hughes A, de Silva S, Gazzard B, et al. A dedicated clinic for HIV-positive individuals over 50 years of age: a multidisciplinary experience. Int J STD AIDS 2012; 23: 546–52.
Zhang Z, Fu J, Zhao Q, He Y, Jin L, Zhang H, et al. Differential restoration of myeloid and plasmacytoid dendritic cells in HIV-1-infected children after treatment with highly active antiretroviral therapy. J Immunol 2006; 176: 5644–51.
Zhang JY, Zhang Z, Wang X, Fu JL, Yao J, Jiao Y, et al. PD-1 up-regulation is correlated with HIV-specific memory CD8+ T-cell exhaustion in typical progressors but not in long-term nonprogressors. Blood 2007; 109: 4671–8.
Herrera-Carrillo E, Berkhout B . Potential mechanisms for cell-based gene therapy to treat HIV/AIDS. Expert Opin Ther Targets 2015; 19: 245–63.
Kent SJ, Reece JC, Petravic J, Martyushev A, Kramski M, De Rose R, et al. The search for an HIV cure: tackling latent infection. Lancet Infect Dis 2013; 13: 614–21.
Finzi D, Hermankova M, Pierson T, Carruth LM, Buck C, Chaisson RE, et al. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science 1997; 278: 1295–300.
Tyagi M, Pearson RJ, Karn J . Establishment of HIV latency in primary CD4+ cells is due to epigenetic transcriptional silencing and P-TEFb restriction. J Virol 2010; 84: 6425–37.
Duverger A, Jones J, May J, Bibollet-Ruche F, Wagner FA, Cron RQ, et al. Determinants of the establishment of human immunodeficiency virus type 1 latency. J Virol 2009; 83: 3078–93.
Jeeninga RE, Westerhout EM, van Gerven ML, Berkhout B . HIV-1 latency in actively dividing human T cell lines. Retrovirology 2008; 5: 37.
Lassen KG, Hebbeler AM, Bhattacharyya D, Lobritz MA, Greene WC . A flexible model of HIV-1 latency permitting evaluation of many primary CD4 T-cell reservoirs. PLoS One 2012; 7: e30176.
Colin L, Van Lint C . Molecular control of HIV-1 postintegration latency: implications for the development of new therapeutic strategies. Retrovirology 2009; 6: 111.
Marsden MD, Zack JA . Eradication of HIV: current challenges and new directions. J Antimicrob Chemother 2009; 63: 7–10.
Donahue DA, Wainberg MA . Cellular and molecular mechanisms involved in the establishment of HIV-1 latency. Retrovirology 2013; 10: 11.
Mbonye U, Karn J . Control of HIV latency by epigenetic and non-epigenetic mechanisms. Curr HIV Res 2011; 9: 554–67.
Shan L, Yang HC, Rabi SA, Bravo HC, Shroff NS, Irizarry RA, et al. Influence of host gene transcription level and orientation on HIV-1 latency in a primary-cell model. J Virol 2011; 85: 5384–93.
Gallastegui E, Millan-Zambrano G, Terme JM, Chavez S, Jordan A . Chromatin reassembly factors are involved in transcriptional interference promoting HIV latency. J Virol 2011; 85: 3187–202.
Imai K, Togami H, Okamoto T . Involvement of histone H3 lysine 9 (H3K9) methyltransferase G9a in the maintenance of HIV-1 latency and its reactivation by BIX01294. J Biol Chem 2010; 285: 16538–45.
Van Lint C, Emiliani S, Ott M, Verdin E . Transcriptional activation and chromatin remodeling of the HIV-1 promoter in response to histone acetylation. EMBO J 1996; 15: 1112–20.
Chan JK, Greene WC . NF-kappaB/Rel: agonist and antagonist roles in HIV-1 latency. Curr Opin HIV AIDS 2011; 6: 12–8.
Boehm D, Calvanese V, Dar RD, Xing S, Schroeder S, Martins L, et al. BET bromodomain-targeting compounds reactivate HIV from latency via a Tat-independent mechanism. Cell Cycle 2013; 12: 452–62.
Karn J . The molecular biology of HIV latency: breaking and restoring the Tat-dependent transcriptional circuit. Curr Opin HIV AIDS 2011; 6: 4–11.
Marban C, Suzanne S, Dequiedt F, de Walque S, Redel L, Van Lint C, et al. Recruitment of chromatin-modifying enzymes by CTIP2 promotes HIV-1 transcriptional silencing. EMBO J 2007; 26: 412–23.
Marban C, Redel L, Suzanne S, Van Lint C, Lecestre D, Chasserot-Golaz S, et al. COUP-TF interacting protein 2 represses the initial phase of HIV-1 gene transcription in human microglial cells. Nucleic Acids Res 2005; 33: 2318–31.
Ping YH, Rana TM . DSIF and NELF interact with RNA polymerase II elongation complex and HIV-1 Tat stimulates P-TEFb-mediated phosphorylation of RNA polymerase II and DSIF during transcription elongation. J Biol Chem 2001; 276: 12951–8.
Natarajan M, Schiralli Lester GM, Lee C, Missra A, Wasserman GA, Steffen M, et al. Negative elongation factor (NELF) coordinates RNA polymerase II pausing, premature termination, and chromatin remodeling to regulate HIV transcription. J Biol Chem 2013; 288: 25995–6003.
Ott M, Geyer M, Zhou Q . The control of HIV transcription: keeping RNA polymerase II on track. Cell Host Microbe 2011; 10: 426–35.
Hoque M, Shamanna RA, Guan D, Pe'ery T, Mathews MB . HIV-1 replication and latency are regulated by translational control of cyclin T1. J Mol Biol 2011; 410: 917–32.
Kim YK, Mbonye U, Hokello J, Karn J . T-cell receptor signaling enhances transcriptional elongation from latent HIV proviruses by activating P-TEFb through an ERK-dependent pathway. J Mol Biol 2011; 410: 896–916.
Sadowski I, Lourenco P, Malcolm T . Factors controlling chromatin organization and nucleosome positioning for establishment and maintenance of HIV latency. Curr HIV Res 2008; 6: 286–95.
Bouchat S, Gatot JS, Kabeya K, Cardona C, Colin L, Herbein G, et al. Histone methyltransferase inhibitors induce HIV-1 recovery in resting CD4(+) T cells from HIV-1-infected HAART-treated patients. AIDS 2012; 26: 1473–82.
Brew BJ, Gray L, Lewin S, Churchill M . Is specific HIV eradication from the brain possible or needed? Expert Opin Biol Ther 2013; 13: 403–9.
Gray LR, Cowley D, Crespan E, Welsh C, Mackenzie C, Wesselingh SL, et al. Reduced basal transcriptional activity of central nervous system-derived HIV type 1 long terminal repeats. AIDS Res Hum Retroviruses 2013; 29: 365–70.
Churchill MJ, Cowley DJ, Wesselingh SL, Gorry PR, Gray LR . HIV-1 transcriptional regulation in the central nervous system and implications for HIV cure research. J Neurovirol 2015; 21: 290–300.
Desplats P, Dumaop W, Smith D, Adame A, Everall I, Letendre S, et al. Molecular and pathologic insights from latent HIV-1 infection in the human brain. Neurology 2013; 80: 1415–23.
Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, et al. Selective inhibition of BET bromodomains. Nature 2010; 468: 1067–73.
Lehrman G, Hogue IB, Palmer S, Jennings C, Spina CA, Wiegand A, et al. Depletion of latent HIV-1 infection in vivo: a proof-of-concept study. Lancet 2005; 366: 549–55.
Siliciano JD, Lai J, Callender M, Pitt E, Zhang H, Margolick JB, et al. Stability of the latent reservoir for HIV-1 in patients receiving valproic acid. J Infect Dis 2007; 195: 833–6.
Sagot-Lerolle N, Lamine A, Chaix ML, Boufassa F, Aboulker JP, Costagliola D, et al. Prolonged valproic acid treatment does not reduce the size of latent HIV reservoir. AIDS 2008; 22: 1125–9.
Archin NM1, Eron JJ, Palmer S, Hartmann-Duff A, Martinson JA, Wiegand A, et al. Valproic acid without intensified antiviral therapy has limited impact on persistent HIV infection of resting CD4+ T cells. AIDS 2008; 22: 1131–5.
Wightman F, Ellenberg P, Churchill M, Lewin SR . HDAC inhibitors in HIV. Immunol Cell Biol 2012; 90: 47–54.
Archin NM, Liberty AL, Kashuba AD, Choudhary SK, Kuruc JD, Crooks AM, et al. Administration of vorinostat disrupts HIV-1 latency in patients on antiretroviral therapy. Nature 2012; 487: 482–5.
Matalon S, Rasmussen TA, Dinarello CA . Histone deacetylase inhibitors for purging HIV-1 from the latent reservoir. Mol Med 2011; 17: 466–72.
Rasmussen TA, Schmeltz Sogaard O, Brinkmann C, Wightman F, Lewin SR, Melchjorsen J, et al. Comparison of HDAC inhibitors in clinical development: effect on HIV production in latently infected cells and T-cell activation. Hum Vaccin Immunother 2013; 9: 993–1001.
Wei DG, Chiang V, Fyne E, Balakrishnan M, Barnes T, Graupe M, et al. Histone deacetylase inhibitor romidepsin induces HIV expression in CD4 T cells from patients on suppressive antiretroviral therapy at concentrations achieved by clinical dosing. PLoS Pathog 2014; 10: e1004071.
Boumber Y, Younes A, Garcia-Manero G . Mocetinostat (MGCD0103): a review of an isotype-specific histone deacetylase inhibitor. Expert Opin Investig Drugs 2011; 20: 823–9.
Deeks SG . HIV: Shock and kill. Nature 2012; 487: 439–40.
Wightman F, Lu HK, Solomon AE, Saleh S, Harman AN, Cunningham AL, et al. Entinostat is a histone deacetylase inhibitor selective for class 1 histone deacetylases and activates HIV production from latently infected primary T cells. AIDS 2013; 27: 2853–62.
Fenaux P . Inhibitors of DNA methylation: beyond myelodysplastic syndromes. Nat Clin Pract Oncol 2005; 2: S36–44.
Kauder SE, Bosque A, Lindqvist A, Planelles V, Verdin E . Epigenetic regulation of HIV-1 latency by cytosine methylation. PLoS Pathog 2009; 5: e1000495.
Fernandez G, Zeichner SL . Cell line-dependent variability in HIV activation employing DNMT inhibitors. Virol J 2010; 7: 266.
du Chene I, Basyuk E, Lin YL, Triboulet R, Knezevich A, Chable-Bessia C, et al. Suv39H1 and HP1gamma are responsible for chromatin-mediated HIV-1 transcriptional silencing and post-integration latency. EMBO J 2007; 26: 424–35.
Friedman J, Cho WK, Chu CK, Keedy KS, Archin NM, Margolis DM, et al. Epigenetic silencing of HIV-1 by the histone H3 lysine 27 methyltransferase enhancer of Zeste 2. J Virol 2011; 85: 9078–89.
Yamada A, Kataoka T, Nagai K . The fungal metabolite gliotoxin: immunosuppressive activity on CTL-mediated cytotoxicity. Immunol Lett 2000; 71: 27–32.
Kawahara N, Nozawa K, Yamazaki M, Nakajima S, Kawai K . Structures of novel epipolythiodioxopiperazines, emethallicins B, C, and D, potent inhibitors of histamine release, from Emericella heterothallica. Chem Pharm Bull (Tokyo) 1990; 38: 73–8.
Bernhard W, Barreto K, Saunders A, Dahabieh MS, Johnson P, Sadowski I . The Suv39H1 methyltransferase inhibitor chaetocin causes induction of integrated HIV-1 without producing a T cell response. FEBS Lett 2011; 585: 3549–54.
Miranda TB, Cortez CC, Yoo CB, Liang G, Abe M, Kelly TK, et al. DZNep is a global histone methylation inhibitor that reactivates developmental genes not silenced by DNA methylation. Mol Cancer Ther 2009; 8: 1579–88.
Zayed S, Sorg B, Hecker E . Structure activity relations of polyfunctional diterpenes of the tigliane type, VI. Irritant and tumor promoting activities of semisynthetic mono and diesters of 12-deoxyphorbol. Planta Med 1984; 50: 65–9.
Szallasi Z, Krsmanovic L, Blumberg PM . Nonpromoting 12-deoxyphorbol 13-esters inhibit phorbol 12-myristate 13-acetate induced tumor promotion in CD-1 mouse skin. Cancer Res 1993; 53: 2507–12.
Hezareh M, Moukil MA, Szanto I, Pondarzewski M, Mouche S, Cherix N, et al. Mechanisms of HIV receptor and co-receptor down-regulation by prostratin: role of conventional and novel PKC isoforms. Antivir Chem Chemother 2004; 15: 207–22.
Kulkosky J, Culnan DM, Roman J, Dornadula G, Schnell M, Boyd MR, et al. Prostratin: activation of latent HIV-1 expression suggests a potential inductive adjuvant therapy for HAART. Blood 2001; 98: 3006–15.
Trushin SA, Bren GD, Asin S, Pennington KN, Paya CV, Badley AD . Human immunodeficiency virus reactivation by phorbol esters or T-cell receptor ligation requires both PKCalpha and PKCtheta. J Virol 2005; 79: 9821–30.
Beans EJ, Fournogerakis D, Gauntlett C, Heumann LV, Kramer R, Marsden MD, et al. Highly potent, synthetically accessible prostratin analogs induce latent HIV expression in vitro and ex vivo. Proc Natl Acad Sci U S A 2013; 110: 11698–703.
Jiang G, Mendes EA, Kaiser P, Sankaran-Walters S, Tang Y, Weber MG, et al. Reactivation of HIV latency by a newly modified Ingenol derivative via protein kinase Cdelta-NF-kappaB signaling. AIDS 2014; 28: 1555–66.
Pandelo Jose D, Bartholomeeusen K, da Cunha RD, Abreu CM, Glinski J, da Costa TB, et al. Reactivation of latent HIV-1 by new semi-synthetic ingenol esters. Virology 2014; 462–3: 328–39.
Dybul M, Hidalgo B, Chun TW, Belson M, Migueles SA, Justement JS, et al. Pilot study of the effects of intermittent interleukin-2 on human immunodeficiency virus (HIV)-specific immune responses in patients treated during recently acquired HIV infection. J Infect Dis 2002; 185: 61–8.
Bowman MC, Archin NM, Margolis DM . Pharmaceutical approaches to eradication of persistent HIV infection. Expert Rev Mol Med 2009; 11: e6.
Wang FX, Xu Y, Sullivan J, Souder E, Argyris EG, Acheampong EA, et al. IL-7 is a potent and proviral strain-specific inducer of latent HIV-1 cellular reservoirs of infected individuals on virally suppressive HAART. J Clin Invest 2005; 115: 128–37.
Siegel DS, Zhang X, Feinman R, Teitz T, Zelenetz A, Richon VM, et al. Hexamethylene bisacetamide induces programmed cell death (apoptosis) and down-regulates BCL-2 expression in human myeloma cells. Proc Natl Acad Sci U S A 1998; 95: 162–6.
Richon VM, Webb Y, Merger R, Sheppard T, Jursic B, Ngo L, et al. Second generation hybrid polar compounds are potent inducers of transformed cell differentiation. Proc Natl Acad Sci U S A 1996; 93: 5705–8.
Vlach J, Pitha PM . Hexamethylene bisacetamide activates the human immunodeficiency virus type 1 provirus by an NF-kappa B-independent mechanism. J Gen Virol 1993; 74: 2401–8.
Choudhary SK, Archin NM, Margolis DM . Hexamethylbisacetamide and disruption of human immunodeficiency virus type 1 latency in CD4+ T cells. J Infect Dis 2008; 197: 1162–70.
Xing S, Bullen CK, Shroff NS, Shan L, Yang HC, Manucci JL, et al. Disulfiram reactivates latent HIV-1 in a Bcl-2-transduced primary CD4+ T cell model without inducing global T cell activation. J Virol 2011; 85: 6060–4.
Doyon G, Zerbato J, Mellors JW, Sluis-Cremer N . Disulfiram reactivates latent HIV-1 expression through depletion of the phosphatase and tensin homolog. AIDS 2013; 27: F7–F11.
Spivak AM, Andrade A, Eisele E, Hoh R, Bacchetti P, Bumpus NN, et al. A pilot study assessing the safety and latency-reversing activity of disulfiram in HIV-1-infected adults on antiretroviral therapy. Clin Infect Dis 2014; 58: 883–90.
Xing S, Bhat S, Shroff NS, Zhang H, Lopez JA, Margolick JB, et al. Novel structurally related compounds reactivate latent HIV-1 in a Bcl-2-transduced primary CD4+ T cell model without inducing global T cell activation. J Antimicrob Chemother 2012; 67: 398–403.
Micheva-Viteva S, Kobayashi Y, Edelstein LC, Pacchia AL, Lee HL, Graci JD, et al. High-throughput screening uncovers a compound that activates latent HIV-1 and acts cooperatively with a histone deacetylase (HDAC) inhibitor. J Biol Chem 2011; 286: 21083–91.
Shan L, Deng K, Shroff NS, Durand CM, Rabi SA, Yang HC, et al. Stimulation of HIV-1-specific cytolytic T lymphocytes facilitates elimination of latent viral reservoir after virus reactivation. Immunity 2012; 36: 491–501.
Author information
Authors and Affiliations
Corresponding author
PowerPoint slides
Rights and permissions
About this article
Cite this article
Shang, Ht., Ding, Jw., Yu, Sy. et al. Progress and challenges in the use of latent HIV-1 reactivating agents. Acta Pharmacol Sin 36, 908–916 (2015). https://doi.org/10.1038/aps.2015.22
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2015.22
Keywords
This article is cited by
-
The CREB Regulated Transcription Coactivator 2 Suppresses HIV-1 Transcription by Preventing RNA Pol II from Binding to HIV-1 LTR
Virologica Sinica (2021)
-
Mesenchymal stem cells are attracted to latent HIV-1-infected cells and enable virus reactivation via a non-canonical PI3K-NFκB signaling pathway
Scientific Reports (2018)
-
BET inhibitors RVX-208 and PFI-1 reactivate HIV-1 from latency
Scientific Reports (2017)
-
Reactivation of HIV-1 from Latency by an Ingenol Derivative from Euphorbia Kansui
Scientific Reports (2017)
-
Underlying mechanisms of HIV-1 latency
Virus Genes (2017)