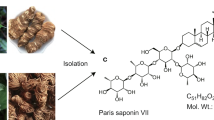
Abstract
Aim:
To investigate the effects of G226, a novel epipolythiodioxopiperazine derivative, on human breast cancer cells in vitro, and to explore its anticancer mechanisms.
Methods:
A panel of human breast cancer cell lines (MDA-MB-231, MDA-MB-468, MCF-7, ZR-75-30, BT474, BT549, SK-BR-3, T47D and HBL100) was examined. Cell proliferation was measured using sulforhodamine B assay, and cell apoptosis was detected with flow cytometry and caspase activity assay. Western blotting, immunofluorescence and targeted gene knockdowns were used to study autophagy in the cells.
Results:
G226 suppressed proliferation of the 9 breast cancer cell lines with a mean IC50 value of 48.5 nmol/L (the mean IC50 value of adriamycin, a reference compound, was 170.6 nmol/L). G226 induced dose-dependent apoptosis of MDA-MB-231 and MCF-7 cells, accompanied by markedly increased activities of caspase-8 and caspase-3/7, which were abolished by caspase inhibitors zVAD or zIETD. G226 also induced mitochondrial outer membrane permeabilization, resulted in the caspase-9 activation. Moreover, G226 dose-dependently enhanced the autophagy marker LC3-II and autophagy substrate p62 accumulation in the cells, which were co-localized with caspase-8. Silencing of p62 or LC3 partially diminished caspase-8 and subsequent caspase-3 activation. LC3 silencing partially reversed G226-induced apoptosis, but p62 silencing elicited a subtle effect on G226-induced apoptosis.
Conclusion:
The novel epipolythiodioxopiperazine derivative G226 exerts potent anticancer action against human breast cancer cells in vitro, via triggering autophagy and caspase-dependent apoptosis.
Similar content being viewed by others
Introduction
Natural products are rich sources of chemically diverse, potential therapeutic leads and have attracted tremendous interest due to their potential for considerable yields of new anti-tumor chemical entities. In this work, we aimed to identify potential anti-cancer drug candidates in natural products. Epipolythiodioxopiperazines (ETPs) are synthesized by fungi, and recent studies have highlighted the potential of ETPs as anticancer agents1. According to our previous studies, we found that 11,11′-dideoxyverticillin, an ETP, which is isolated from Shiraia bambusicola, exhibits potent cytotoxicity against a broad spectrum of cancer cell lines in vitro2. In this current study, we report that G226 (Figure 1A), a novel 11′-deoxyverticillin A derivative, exhibits potent cytotoxic activity against breast cancer. We also determine the mechanism by which G226 kills cancer cells and find that it significantly induces apoptosis and autophagy in breast cancer cells.
Cytotoxicity of G226 against human breast tumor cells. (A) Chemical structure of G226. (B) G226 inhibits human breast tumor cell proliferation in vitro.
Cell-killing intracellular events constitute a programmed cell death (PCD) phenomenon, which includes at least the following three different mechanisms: apoptosis, autophagy, and programmed necrosis3,4,5. These mechanisms are regulated and executed by different signaling pathways3.
Apoptosis (type I PCD) is characterized by chromatin condensation and fragmentation, cell shrinkage and the disintegration of cells into apoptotic bodies5. At the molecular level, apoptosis is a stringently controlled and ordered cellular suicide program that is critical for the development, immune regulation, and homeostasis of a multicellular organism. Moreover, apoptosis is thought to be an important and useful mechanism by which anti-tumor agents kill cancer cells. Although several different pathways can lead to apoptosis, a crucial event is the activation of caspases6,7.
Autophagy, or type II PCD, is a highly conserved “self-eating” process that requires lysosomes for the degradation and recycling of long-lived proteins and organelles. Autophagy is upregulated as a pro-survival response to extracellular or intracellular stresses and signals including starvation, growth factor deprivation, hypoxia, ER stress, and pathogen infection8,9,10,11. Defective autophagy is correlated with tumorigenesis8. Altered autophagy in cancer cells is paradoxical because autophagy functions as a tumor suppressive mechanism; however, it also represents a cytoprotective mechanism that cancer cells use to survive in hostile microenvironments. As certain tumors can exploit autophagy to maintain cell viability under nutrient-limiting conditions12 or to circumvent the effects of a wide spectrum of cytotoxic and targeted chemotherapeutic agents13, the inhibition of autophagy can kill tumor cells. Conversely, many studies have shown that the induction of autophagy can also contribute to caspase-dependent or caspase-independent apoptosis14,15,16,17. The inhibition of autophagy results in significant attenuation of cell death in many cellular circumstances, such as in cells with defective apoptotic machinery or in the presence of caspase inhibitors15,16.
During certain cellular circumstances and cell death scenarios, a mixed phenotype of more than one type of death has been observed within a single cell18. Because apoptosis invariably contributes to cell death and because autophagy can have either pro-survival or pro-death roles, these two forms of PCD jointly decide the fate of cells. Additionally, there must be crosstalk between apoptosis and autophagy that is complex and sometimes contradictory18,19. In this study, we sought to determine the role of autophagy activation in response to G226 and to explore whether autophagosomes are functionally involved in G226-induced cancer cell death. Here, we provide evidence that G226 induces the formation of the intracellular death-inducing signaling complex (iDISC) to activate Caspase-8, triggering LC3- and p62-dependent apoptosis in human breast tumor cells.
Materials and methods
Chemicals
G226 was kindly provided by Prof Yong-sheng CHE (Beijing Institute of Pharmacology & Toxicology, Beijing, China). The purity of the compound exceeded 98%. Z-VAD(OMe)-FMK and Z-IETD-FMK were obtained from Bachem AG (Bubendorf, Switzerland). 3-Methyladenine (3-MA) and chloroquine (CQ) were purchased from Sigma (St Louis, MO, USA). CQ was dissolved in sterile water at 100 mmol/L (used in vitro) as a stock solution and maintained at 4 °C, whereas the other compounds were dissolved in DMSO at 10 mmol/L (used in vitro) as stock solutions and maintained at −80 °C in the dark. All compounds were diluted to the desired concentrations in full medium immediately prior to each experiment. The final concentrations of DMSO did not exceed 0.1%.
Cell culture
The human breast cancer cell lines MDA-MB-231, MDA-MB-468, MCF-7, ZR-75-30, BT474, SK-BR-3, and T47D were purchased from the American Type Culture Collection (Manassas, VA, USA). The BT549 and HBL100 cell lines were obtained from the cell bank of the Shanghai Institute of Materia Medica, Chinese Academy of Sciences. The cells were cultured in RPMI-1640 medium (Gibco, Grand Island, NY, USA) (BT549, ZR-75-30, T47D), McCoy's 5A medium (Gibco) (SK-BR-3), LG-DMEM (Gibco) (MCF7, HBL100, BT474) or L-15 medium (Gibco) (MDA-MB-231, MDA-MB-468) supplemented with 10%-15% heat-inactivated fetal calf serum (FBS; Gibco).
Cell proliferation assay
The cytotoxicity of G226 was examined in a panel of human breast tumor cell lines; adriamycin (ADR) was used as a reference compound and was applied in parallel. Cells plated in 96-well plates were treated with gradient concentrations of compounds at 37 °C for 72 h. A cell proliferation assay was performed using sulforhodamine B (SRB; Sigma, St Louis, MO, USA).
Flow cytometry
Cells were stained with Annexin V-FITC and propidium iodide (PI) and then evaluated for apoptosis by flow cytometry according to the manufacturer's protocol (Invitrogen, Carlsbad, CA, USA). Briefly, after treatment, the cells were harvested by centrifugation. The cell pellets were suspended in 100 μL of binding buffer and incubated with 5 μL of Annexin V-FITC and 5 μL of a PI solution at room temperature for 15 min. Annexin V and PI staining was measured by flow cytometry on a FACSCalibur instrument (BD Biosciences, NJ, USA) followed by data analysis using FlowJo software.
For analysis of mitochondrial membrane potential, the JC-1 probe (Beyotime, Haimen, China) was used. Briefly, after drug treatment, the cells were stained with the JC-1 fluorescent probe at 37 °C for 20 min. Next, the cells were washed twice with binding buffer and resuspended in 1 mL of culture medium for fluorescence microscopy analysis. For the FACS assay, the cells stained with the JC-1 fluorescent probe were harvested by centrifugation, suspended in culture medium and then subjected to flow cytometry analysis on a FACSCalibur instrument (BD Biosciences) followed by data analysis using FlowJo software.
Transfection of small interfering RNAs (siRNAs)
p62, LC3 and scrambled control siRNAs were purchased from Santa Cruz Biotechnology. MDA-MB-231 and MCF-7 cells were transfected with Lipofectamine™ RNAiMAX reagent (Invitrogen) according to the manufacturer's protocols. The final concentration of all siRNAs was 34 nmol/L.
Western blot analysis
Treated cells were washed with cold PBS and lysed in RIPA buffer. Equal amounts of proteins were resolved by SDS-PAGE, transferred to nitrocellulose membranes (Thermo Scientific, Franklin, MA, USA), blocked with 5% (w/v) nonfat dry milk in TBST, and probed with antibodies against Caspase-8, Caspase-9, Caspase-3, PARP, Bid, XIAP, Mcl-1, FLIP, c-IAP1, c-IAP2, anti-LC3 I/II (Cell Signaling, Danvers, MA, USA), p62 (Santa Cruz Biotechnology, Santa Cruz, CA, USA), and GAPDH (Epitomics, CA, USA). The data shown are representative results obtained in three independent experiments.
Immunofluorescence assays
After treatments with the indicated agents, cells growing on glass coverslips were fixed with 4% paraformaldehyde and permeabilized with PBS containing 0.2% Triton X-100. After blocking with PBS containing 3% BSA for 30 min, the cells were incubated with LC3 (1:100; MBL, Woburn, MA, USA), p62 (1:200, Santa Cruz Biotechnology), and Caspase-8 (1:100, mouse monoclonal, Cell Signaling or 1:200, rabbit polyclonal, Santa Cruz) antibodies. Next, the cells were stained with Alexa Fluor 488-conjugated anti-rabbit secondary antibody or Alexa Fluor 633-conjugated anti-mouse secondary antibody (1:200; Invitrogen). Finally, after counterstaining with DAPI (Vector Laboratories, Burlingame, CA, USA), the cells were observed and imaged using an Olympus BX51 fluorescence microscope (Olympus, Japan).
Caspase activity assay
Caspase activity was determined using the Caspase-Glo 8 Assay Kit (Promega, Madison, WI, USA) and the Apo-ONE Homogeneous Caspase-3/7 Assay Kit (Promega) according to the manufacturer's protocols. Briefly, after treatment, floating and attached cells in 6-well plates were harvested and suspended in culture medium for detection. Cells were counted using a CYTORECON™ (GE healthcare, Piscataway, NJ, USA). Next, 50 μL of Caspase-Glo 8 or Apo-ONE Caspase-3/7 reagent was added to each well of a black 96-well plate containing samples or blank medium. After mixing the contents of the wells using a plate shaker and incubation at room temperature, the fluorescence was measured at 30 min intervals up to 6 h using a SpectraMax M5 luminescence reader (Molecular Devices, Sunnyvale, CA, USA).
Statistical analysis
The data are presented as the mean values±SD. Statistical significance was analyzed by two-tailed Student's t-test. P values <0.05 were considered statistically significant.
Results
G226 inhibits the proliferation of human breast cancer cells
The activity of G226 against human breast cancer was evaluated in a panel of breast cancer cell lines including estrogen receptor (ER)-positive, HER2-positive and triple-negative breast cancer cells. As shown in Figure 1B, G226 displayed potent cytotoxicity against these cell lines. The mean IC50 for all 9 tumor cell lines tested was 48.5 nmol/L, which was more potent than adriamycin (170.6 nmol/L).
G226 triggers caspase-dependent apoptosis
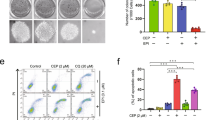
To investigate the detailed mechanism underlying G226-induced breast cancer cell death, we examined the ability of G226 to induce apoptosis. ER-negative MDA-MB-231 and ER-positive MCF-7 cells were used for these studies. G226 significantly increased the percentage of Annexin V+-labeled cells in a dose- and time-dependent manner, as indicated by flow cytometric analysis (Figure 2A and 2B). After treatment with 200 nmol/L G226 for 24 h, approximately 30 % cells underwent apoptosis. The apoptotic cleavage of poly (ADP-ribose) polymerase (PARP) and Caspase-8 was also increased in a dose-dependent manner by G226 (Figure 2C). Because Caspase-3 is deficient in MCF-7 cells, the increased cleaved Caspase-3 was only observed in MDA-MB-231 cells that were exposed to G226 (Figure 2C). Conversely, Caspase-Glo 8 and Caspase-3/7 activities were determined using commercially available kits. G226 significantly activated these caspases (Figure 2D). Taken together, these data indicate that caspases are involved in G226-induced apoptosis. Next, caspase inhibitors, such as zVAD and zIETD, were used to examine whether the inhibition of caspase cleavage would be sufficient to attenuate G226-induced apoptosis. As shown in Figure 2E–2G, both inhibitors abolished the activities of Caspase-8 and Caspase-3/7 and significantly attenuated G226-induced apoptosis. Collectively, these findings suggest that G226 induces caspase-dependent apoptosis.
G226 triggers apoptosis via a caspase-dependent pathway. MDA-MB-231 and MCF-7 cells were treated with increasing concentrations of G226 for 24 h (A) or with 200 nmol/L G226 for 0, 3, 6, 12, 24, or 48 h (B); apoptotic cells are indicated as Annexin V+-labeled cells. The cells were treated with increasing concentrations of G226 for 24 h and then collected for Western blotting with the indicated antibodies (C) and for determining the activities of the Caspase-8-like IETDase and the Caspase-3/7-like DEVDase (D). In the presence or absence of 20 μmol/L zVAD or 25 μmol/L zIETD, MDA-MB-231 cells were treated with 200 nmol/L G226 for 24 h (E-G). (E) The activities of the Caspase-8-like IETDase and the Caspase-3/7-like DEVDase were measured. The fold-changes were defined as the caspase activity of drug-treated cells/the caspase activity of respective DMSO-treated cells. (F) Apoptosis was detected by Annexin V+-PI staining and analyzed by flow cytometry. (G) The cell lysates were collected and examined by immunoblotting with the indicated antibodies. The data are shown as the mean±SD of independent experiments. The statistical significance was assessed by analysis of variance. bP<0.05, cP<0.01 compared with the DMSO group.
The mitochondrial pathway is involved in G226-induced apoptosis
Many agents induce apoptosis in cells via two major pathways: the death receptor-mediated pathway and the mitochondria-mediated pathway. Caspase-8 is an initiator caspase, which not only activates executioner caspases, such as procaspase-3 and procaspase-7, to complete the death receptor signaling pathway but also cleaves the Bcl-2 family member Bid to amplify apoptosis via the mitochondrial pathway20. Our results show that G226 increased the levels of JC-1 monomers at a concentration of 200 nmol/L, indicating that mitochondrial outer membrane permeabilization (MOMP) occurred after treatment with G226 (Figure 3A). However, this compound elicited no effect on the expression of Bcl-2, Bcl-XL, or Bax at concentrations within the 25–200 nmol/L range (Figure 3B). Only activated Caspase-9 and cleaved Bid (tBid) were increased in the cells (Figure 3B). These data suggest that in G226-treated cells, activated Caspase-8 cleaved Bid, resulting in the release of cytochrome c from the mitochondria and consequential activation of Caspase-9.
G226 induces mitochondrial outer membrane permeabilization. (A) MDA-MB-231 and MCF-7 cells were treated with 200 nmol/L G226 for 24 h, and the mitochondrial membrane potential of the cells was measured by flow cytometry after JC-1 staining. (B) MDA-MB-231 and MCF-7 cells were treated with increasing concentrations of G226 for 24 h, and the cell lysates were collected and examined by immunoblotting with the indicated antibodies.
G226 enhances autophagy in human breast cancer cell lines
Next, we determined whether G226-induced autophagy, another important cell-killing intracellular event, occurs. LC3-II is required for the elongation and closure of autophagosomal membranes and is considered as a marker of autophagy9,21. Compared with the control-treated cells, LC3-II accumulated in MDA-MB-231 and MCF-7 cells treated with G226 in a dose- and time-dependent manner (Figure 4A and 4B). Furthermore, treatment of cells with the lysosomal inhibitor chloroquine (CQ) resulted in the increased accumulation of LC3 puncta in response to G226 in MCF-7 cells (Figure 4C). These data suggest that G226 also induces autophagy in breast cancer cells. The degradation of p62 is considered to be another marker for autophagy because p62 is a substrate of autophagy. To our surprise, G226 resulted in the accumulation of p62 at lower doses and at early timepoints of treatment (Figure 4A and 4B). Interestingly, we also observed that the increased LC3-II expression (Figure 4B) resulting from G226 treatment occurred concomitantly with an increase in Annexin V+-labeled cells (Figure 2B) and in cleaved PARP (Figure 4B), indicating that G226-induced autophagy is associated with the apoptotic process.
G226 induces caspase-dependent apoptosis accompanied by induction of autophagy. MDA-MB-231 and MCF-7 cells were treated with increasing concentrations of G226 for 24 h (A) or with 200 nmol/L G226 for 0, 3, 6, 12, 24, or 48 h (B), and the cell lysates were collected and examined by immunoblotting with the indicated antibodies. (C) In the presence or absence of 25 μmol/L CQ, MCF-7 cells were treated with 100 nmol/L G226 for 24 h, and fluorescence deconvolution microscopy was performed using anti-LC3 (green) antibodies. DAPI (blue) was used to visualize nuclei. Representative images from three independent experiments are shown. In the presence or absence of 2 mmol/L 3-MA or 25 μmol/L CQ, the cells were treated with 200 nmol/L G226, and the cell lysates were examined by immunoblotting with the indicated antibodies (D). The apoptotic cells are indicated by Annexin V+-labeling (E). The data are expressed as the mean±SD of independent experiments. The statistical significance was assessed by analysis of variance. bP<0.05, cP<0.01 compared with the DMSO group.
We next used the autophagy inhibitors CQ and 3-methyladenine (3-MA) to explore the interplay between G226-induced autophagy and apoptosis. CQ and 3-MA inhibited cellular autophagy, as evidenced by the accumulation of LC3-II, resulting in a distinct suppression of G226-induced PARP cleavage and apoptosis (Figure 4D and 4E). These data demonstrate that the autophagy stimulated by G226 participates in G226-induced apoptosis.
LC3 and p62 are required for G226-induced activation of Caspase-8
Because p62 and LC3 promote the formation of the intracellular death-inducing signaling complex (iDISC), which mediates Caspase-8 activation22,23, we further explored whether the activation of Caspase-8 by G226 requires p62 or LC3. Immunofluorescence analysis revealed that endogenous Caspase-8, LC3, and p62 co-localized and interacted in G226-treated cells (Figure 5). As expected, the silencing of p62 or LC3 partially diminished Caspase-8 activation and subsequent Caspase-3 activation (Figure 6A). Moreover, the silencing of LC3 partially reversed G226-induced apoptosis, although p62 silencing only elicited a subtle effect (Figure 6B and 6C). Nrf2 is considered as a target gene of p62 and can upregulate the transcription of cytoprotective genes24,25. We found that Nrf2 was increased when p62 was overexpressed in response to G226. Moreover, knockdown of p62 also reversed the G226-induced Nrf2 upregulation (Figure 6D). This finding might explain why the silencing of p62 is insufficient to block G226-induced apoptosis.
G226 induces the formation of a complex containing Caspase-8, p62 and LC3. MDA-MB-231 and MCF-7 cells were treated with 100 nmol/L G226 or vehicle for 24 h. (A) Fluorescence deconvolution microscopy was performed; Caspase-8 is shown in green, whereas p62 is shown in red. (B) LC3 is shown in green and p62 is shown in red; (C) LC3 is green and Caspase-8 is red; Nuclei labeled with DAPI are shown in blue. Representative images of three independent experiments are shown.
Silencing of LC3 or p62 partially reverses the G226-induced activation of Caspase-8. MDA-MB-231 cells were transfected with a scrambled control siRNA or siRNAs against p62 or LC3 for 24 h followed by treatment with 200 nmol/L G226 for 24 h. (A) The activities of the Caspase-8-like IETDase and the Caspase-3/7-like DEVDase were measured; (B) Apoptotic cells were indicated by Annexin V+-labeling and determined by flow cytometry; (C) Cell lysates were collected and examined by immunoblotting with the indicated antibodies; and (D) Fluorescence deconvolution microscopy was performed using anti-Nrf2 (green) and anti-p62 (red) antibodies. DAPI was used to visualize the nuclei (blue). Representative images of three independent experiments are shown. The data are expressed as the mean±SD of independent experiments. The statistical significance was assessed by analysis of variance. bP<0.05, cP<0.01 compared with the DMSO group.
Decreased levels of FLIP, c-IAP1, XIAP, and MCL-1 are involved in G226-induced caspase-dependent apoptosis
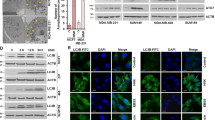
The above data demonstrate that apoptosis plays an essential role in G226-induced tumor cell death. Therefore, we further studied whether inhibitor of apoptosis proteins (IAPs), FLICE inhibitory proteins or Bcl-2 family proteins were also involved in G226-induced apoptosis. Indeed, the expression of FLIP, c-IAP1, XIAP, and MCL-1 was dose-dependently decreased after treatment with G226 (Figure 7A). The autophagy inhibitors CQ and 3-MA did not attenuate the downregulation of these proteins (Figure 7B). These data suggest that the downregulation of FLIP, c-IAP1, XIAP, and MCL-1 is involved in G226-induced caspase-dependent apoptosis but is not regulated by autophagy.
G226 decreases the levels of c-FLIPs, IAPs and MCL-1. (A) MDA-MB-231 and MCF-7 cells were treated with increasing concentrations of G226 for 24 h, and the cell lysates were examined by immunoblotting with the indicated antibodies. (B) In the presence or absence of 2 mmol/L 3-MA or 25 μmol/L CQ, MDA-MB-231 and MCF7 cells were treated with 200 nmol/L G226, and the cell lysates were collected and examined by immunoblotting with the indicated antibodies.
Discussion
Because gliotoxin elicits potent anti-tumor activity in vivo26, many ETPs have been explored. In this study, the novel ETP G226 was synthesized, which exhibits promising anti-proliferative activity against human breast cancer by triggering autophagy and caspase-dependent apoptosis.
Our previous study found that G226 inhibits topoisomerase II-mediated supercoiled pBR322 relaxation and kDNA decatenation and increases phospho-H2AX levels. The therapeutic efficacy of topoisomerase inhibitors is generally suggested to be attributable to activation of the apoptotic cascade. Exploration of the apoptotic pathway that becomes activated in response to G226 will enhance our understanding of the mechanisms underlying its anti-tumor activity. We demonstrated that G226 stimulates the activation of Caspase-8 and downstream caspases (Caspase-3/9) and induces mitochondrial outer membrane permeabilization but elicits no effect on the expression of Bcl-2 family proteins. These findings highlighted the critical roles of death receptor-mediated caspase activation in G226-mediated cellular effects.
Although apoptosis is considered as the principal mechanism by which chemotherapeutic agents kill cells, growing evidence has shown that anti-cancer agents also induce other forms of non-apoptotic cell death including necrosis, mitotic catastrophe, autophagy and senescence27,28,29. Upon the initiation of autophagy, LC3-I is converted to LC3-II and localizes to autophagosomal membranes to promote the subsequent dynamic rearrangement of intracellular membranes and the formation of autophagosomes21. Therefore, LC3-II represents an excellent autophagy marker for monitoring autophagy flux. In our study, compared with the control treatment, G226 treatment caused the accumulation of LC3-II and further increases in the levels of LC3 puncta. Autophagy can function as a tumor suppressor but also has a tumor-promoting effect in different models. Therefore, we have illustrated the role of autophagy in G226-induced cancer cell death in our studies. Immunofluorescence analysis revealed that autophagosomal membranes serve as a platform in breast cancer cells for recruiting p62, Caspase-8 and LC3 to form a complex, which mediates Caspase-8 activation and apoptosis. The suppression of autophagy by chemical inhibitors or by the silencing of LC3 or p62 partially prevented the activation of Caspase-8 and apoptosis. Taken together, our data demonstrate that LC3 and p62 are required for G226-induced caspase-dependent apoptosis.
The adaptor protein p62 plays crucial roles in a number of cellular functions, including bone remodeling, obesity, and cancer30. p62 contains an LC3-interacting motif as well as a ubiquitin binding domain. It can be incorporated into complete autophagosomes with proteins that are targeted for degradation by autophagy. Thus, p62 not only modulates but also serves as an indicator of autophagic degradation9,31. Although its degradation is generally considered as a marker for autophagy, enhanced p62 expression has also been reported to accompany autophagy. In our study, G226 treatment increased p62 protein expression and was accompanied by autophagy induction. Aside from its role as a marker for autophagy, p62 can also regulate the expression of other genes32. p62 has been demonstrated to serve as an endogenous inducer of Nrf2 protein expression, and Nrf2 is responsible for controlling the transcription of cytoprotective genes24,32,33. Increased Nrf2 was found after G226 treatment along with excess accumulation of p62, and p62 knockdown completely reversed G226-induced Nrf2 upregulation. We also found that the silencing of p62 did not significantly reverse G226-triggered apoptosis. This unexpected result might be at least partially attributed to the excessive decrease in Nrf2 levels34. The transcription factor Nrf2 controls cellular adaptation to oxidants and electrophiles by inducing cytoprotective genes, including antioxidant and detoxification genes that respond to redox stress35,36,37. Nrf2-knockout mice exhibit reduced expression of glutathione biosynthetic genes and GSTs, diminished detoxification capabilities, and enhanced susceptibility to oxidative stress-induced cell death. Conversely, Nrf2 overexpression in vitro and in vivo protects against oxidative stress. Furthermore, p62 has been shown to positively regulate the transcription of Nrf2 and result in the activation of Nrf2. Persistent activation of Nrf2 through the accumulation of phosphorylated p62 contributes to the growth of human hepatocellular carcinomas. In our study, we found that the silencing of p62 did not reverse G226-induced cell death but rather reversed G226-induced upregulation of Nrf2. Thus, we speculate that knockdown of p62 might suppress cellular survival pathways to enhance apoptosis.
In conclusion, this is the first report of G226, a novel ETP compound, which induces autophagy. Moreover, the autophagosomal membrane might serve as a platform for the formation of the iDISC, which activates Caspase-8 and the caspase cascade, leading to apoptosis. Due to its significant anti-breast cancer activities, G226 represents an attractive therapeutic agent for cancer therapy.
Author contribution
Jian DING, Yi CHEN, Qiao-jun HE, and Peng-xing HE designed the research; Yong-sheng CHE provided the G226 compound; Peng-xing HE performed the experiments, analyzed the results and wrote the manuscript; Yi CHEN and Jian DING revised the manuscript.
Abbreviations
ETPs, epipolythiodioxopiperazines; PCD, programmed cell death; iDISC, intracellular death-inducing signaling complex; CQ, chloroquine; 3-MA, 3-methyladenine; PARP, poly (ADP-ribose) polymerase; MOMP, mitochondrial outer membrane permeabilization; IAPs, inhibitor of apoptosis proteins; FLIP, FLICE inhibitory protein; ADR, adriamycin.
References
Gardiner DM, Waring P, Howlett BJ . The epipolythiodioxopiperazine (ETP) class of fungal toxins: distribution, mode of action, functions and biosynthesis Microbiology 2005; 151: 1021–32.
Son BW, Jensen PR, Kauffman CA, Fenical W . New cytotoxic epidithiodioxopiperazines related to verticillin A from a marine isolate of the Fungus Penicillium. Nat Product Lett 1999; 13: 213–22.
Bialik S, Zalckvar E, Ber Y, Rubinstein AD, Kimchi A . Systems biology analysis of programmed cell death. Trends Biochem Sci 2010; 35: 556–64.
Galluzzi L, Vanden Berghe T, Vanlangenakker N, Buettner S, Eisenberg T, Vandenabeele P, et al. Programmed necrosis from molecules to health and disease. Int Rev Cell Mol Biol 2011; 289: 1–35.
Ouyang L, Shi Z, Zhao S, Wang FT, Zhou TT, Liu B, et al. Programmed cell death pathways in cancer: a review of apoptosis, autophagy and programmed necrosis. Cell Prolif 2012; 45: 487–98.
Budihardjo I, Oliver H, Lutter M, Luo X, Wang X . Biochemical pathways of caspase activation during apoptosis. Annu Rev Cell Dev Biol 1999; 15: 269–90.
Hengartner MO . The biochemistry of apoptosis. Nature 2000; 407: 770–6.
He C, Klionsky DJ . Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 2009; 43: 67–93.
Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012; 8: 445–544.
Kimmelman AC . The dynamic nature of autophagy in cancer. Genes Dev 2011; 25: 1999–2010.
Li X, Xu HL, Liu YX, An N, Zhao S, Bao JK . Autophagy modulation as a target for anticancer drug discovery. Acta Pharmacol Sin 2013; 34: 612–24.
Eisenberg-Lerner A, Kimchi A . The paradox of autophagy and its implication in cancer etiology and therapy. Apoptosis 2009; 14: 376–91.
Marx J . Autophagy: Is it cancer's friend or foe? Science 2006; 312: 1160–1.
Laussmann MA, Passante E, Dussmann H, Rauen JA, Wurstle ML, Delgado ME, et al. Proteasome inhibition can induce an autophagy-dependent apical activation of caspase-8. Cell Death Differ 2011; 18: 1584–97.
Shimizu S, Kanaseki T, Mizushima N, Mizuta T, Arakawa-Kobayashi S, Thompson CB, et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat Cell Biol 2004; 6: 1221–8.
Yu L, Alva A, Su H, Dutt P, Freundt E, Welsh S, et al. Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 2004; 304: 1500–2.
Hou LL, Gao C, Chen L, Hu GQ, Xie SQ . Essential role of autophagy in fucoxanthin-induced cytotoxicity to human epithelial cervical cancer HeLa cells. Acta Pharmacol Sin 2013; 34: 1403–10.
Eisenberg-Lerner A, Bialik S, Simon HU, Kimchi A . Life and death partners: apoptosis, autophagy and the cross-talk between them. Cell Death Differ 2009; 16: 966–75.
Djavaheri-Mergny M, Maiuri MC, Kroemer G . Cross talk between apoptosis and autophagy by caspase-mediated cleavage of Beclin 1. Oncogene 2010; 29: 1717–9.
Degterev A, Boyce M, Yuan J . A decade of caspases. Oncogene 2003; 22: 8543–67.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–8.
Pan JA, Ullman E, Dou Z, Zong WX . Inhibition of protein degradation induces apoptosis through a microtubule-associated protein 1 light chain 3-mediated activation of caspase-8 at intracellular membranes. Mol Cell Biol 2011; 31: 3158–70.
Young MM, Takahashi Y, Khan O, Park S, Hori T, Yun J, et al. Autophagosomal membrane serves as platform for intracellular death-inducing signaling complex (iDISC)-mediated caspase-8 activation and apoptosis. J Biol Chem 2012; 287: 12455–68.
Liu Y, Kern JT, Walker JR, Johnson JA, Schultz PG, Luesch H . A genomic screen for activators of the antioxidant response element. Proc Natl Acad Sci U S A 2007; 104: 5205–10.
Fan W, Tang Z, Chen D, Moughon D, Ding X, Chen S, et al. Keap1 facilitates p62-mediated ubiquitin aggregate clearance via autophagy. Autophagy 2010; 6: 614–21.
Vigushin DM, Mirsaidi N, Brooke G, Sun C, Pace P, Inman L, et al. Gliotoxin is a dual inhibitor of farnesyltransferase and geranylgeranyltransferase I with antitumor activity against breast cancer in vivo. Med Oncol 2004; 21: 21–30.
Ricci MS, Zong WX . Chemotherapeutic approaches for targeting cell death pathways. Oncologist 2006; 11: 342–57.
Portugal J, Bataller M, Mansilla S . Cell death pathways in response to antitumor therapy. Tumori 2009; 95: 409–21.
Al-Ejeh F, Kumar R, Wiegmans A, Lakhani SR, Brown MP, Khanna KK . Harnessing the complexity of DNA-damage response pathways to improve cancer treatment outcomes. Oncogene 2010; 29: 6085–98.
Moscat J, Diaz-Meco MT . p62 at the crossroads of autophagy, apoptosis, and cancer. Cell 2009; 137: 1001–4.
Zhou L, Wang HF, Ren HG, Chen D, Gao F, Hu QS, et al. Bcl-2-dependent upregulation of autophagy by sequestosome 1/p62 in vitro. Acta Pharmacol Sin 2013; 34: 651–6.
Jain A, Lamark T, Sjottem E, Larsen KB, Awuh JA, Overvatn A, et al. p62/SQSTM1 is a target gene for transcription factor NRF2 and creates a positive feedback loop by inducing antioxidant response element-driven gene transcription. J Biol Chem 2010; 285: 22576–91.
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A, et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 2005; 171: 603–14.
Motohashi H, Yamamoto M . Nrf2-Keap1 defines a physiologically important stress response mechanism. Trends Mol Med 2004; 10: 549–57.
Hayes JD, McMahon M . NRF2 and KEAP1 mutations: permanent activation of an adaptive response in cancer. Trends Biochem Sci 2009; 34: 176–88.
Taguchi K, Motohashi H, Yamamoto M . Molecular mechanisms of the Keap1-Nrf2 pathway in stress response and cancer evolution. Genes Cells 2011; 16: 123–40.
Villeneuve NF, Lau A, Zhang DD . Regulation of the Nrf2-Keap1 antioxidant response by the ubiquitin proteasome system: an insight into cullin-ring ubiquitin ligases. Antioxid Redox Signal 2010; 13: 1699–712.
Acknowledgements
The project was supported by National Natural Science Foundation of China (81321092) and by a grant from the National Basic Research Program of China (2013CB932503).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
He, Px., Che, Ys., He, Qj. et al. G226, a novel epipolythiodioxopiperazine derivative, induces autophagy and caspase-dependent apoptosis in human breast cancer cells in vitro. Acta Pharmacol Sin 35, 1055–1064 (2014). https://doi.org/10.1038/aps.2014.47
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2014.47
Keywords
This article is cited by
-
Targeting NLRP3 Inflammasome in Inflammatory Bowel Disease: Putting out the Fire of Inflammation
Inflammation (2019)
-
Sequestosome 1 protects esophageal squamous carcinoma cells from apoptosis via stabilizing SKP2 under serum starvation condition
Oncogene (2018)
-
Carbazole alkaloids from Murraya koenigii trigger apoptosis and autophagic flux inhibition in human oral squamous cell carcinoma cells
Journal of Natural Medicines (2017)
-
Galangin Induces Autophagy via Deacetylation of LC3 by SIRT1 in HepG2 Cells
Scientific Reports (2016)
-
The roles of macrophage autophagy in atherosclerosis
Acta Pharmacologica Sinica (2016)