Abstract
Aim:
A large number of drug-induced long QT syndromes are ascribed to blockage of hERG potassium channels. The aim of this study was to construct novel computational models to predict compounds blocking hERG channels.
Methods:
Doddareddy's hERG blockage data containing 2644 compounds were used, which divided into training (2389) and test (255) sets. Laplacian-corrected Bayesian classification models were constructed using Discovery Studio. The models were internally validated with the training set of compounds, and then applied to the test set for validation. Doddareddy's experimentally validated dataset with 60 compounds was used for external test set validation.
Results:
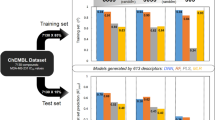
A Bayesian classification model considering the effects of four molecular properties (Mw, PPSA, ALogP and pKa_basic) as well as extended-connectivity fingerprints (ECFP_14) exhibited a global accuracy (91%), parameter sensitivity (90%) and specificity (92%) in the test set validation, and a global accuracy (58%), parameter sensitivity (61%) and specificity (57%) in the external test set validation.
Conclusion:
The novel model is better than those in the literatures for predicting compounds blocking hERG channels, and can be used for large-scale prediction.
Similar content being viewed by others
Introduction
Long QT syndrome (LQTs) occurs frequently and is associated with a potential risk of life-threatening torsades de pointes (TdP). Many drugs have been withdrawn from the market for LQTs toxicity1,2,3. The human ether-à-go-go-related gene (hERG) product is a tetrameric potassium channel that plays an important role in cardiac action potential. Currently, a large number of clinical cases of drug-induced or acquired LQTs have been ascribed to the blockage of the hERG channel4,5. Therefore, it is necessary to measure the hERG inhibition activity of candidate compounds during drug discovery. Many experimental assays have been used to detect hERG blockage, such as patch clamp assays, radioligand binding assays, and fluorescence-based assays6. However, these assays are time-consuming, labor-intensive and costly. It is of high interest to develop computational methods that can rapidly and accurately assess the hERG liability of a compound before it is synthesized or purchased from a commercial library.
In past years, many hERG blockage prediction approaches have been reported and can be roughly categorized as structure-based or ligand-based. Because the crystal structure of the hERG potassium channel has not been resolved as yet, the structure-based approaches rely mainly on homology modeling. These structure-based approaches are useful for explaining the binding effect of closely related analogues or a small collection of compounds. However, due to the lack of a suitable template three-dimensional (3D) structure with sufficient sequence identity, the modeled hERG structure shows limited ability to substantially improve the prediction capabilities of these approaches.
With the rapid accumulation of hERG blockage data for various compounds, more ligand-based approaches were reported. In 2002, Cavalli et al used the CoMFA method analysis for 31 drugs and built a four-point pharmacophore model that included a positively charged tertiary amine flanked by three aromatic or hydrophobic centers, which was validated using 6 compounds (r2=0.744)7. In 2003, Keseru et al developed traditional and hologram QSAR (HQSAR) models with five descriptors, including ClogP, molar refractivity (CMR), partial negative surface area (PNSA1), polarizability (W2), and hydrophobicity (D3), to predict hERG affinities, for a test set of 13 compounds (r=0.75)8. In 2006, Yoshida et al carried out 2D-quantitative structure activity relationship (2D-QSAR) studies on 104 hERG channel blockers with descriptors that included the octanol/water partition coefficient, topological polar surface area, molecular diameter, the summed surface area of the atoms and an indicator variable representing the experimental conditions for a test set containing 18 compounds (r2=0.688)9. In 2008, Li et al performed a classification model of hERG blockage for 495 compounds based on GRIND descriptors and the support vector machine (SVM) method, which achieved an accuracy of 92% for the training set but only an accuracy of 72% for the test set of 66 WOMBAT-PK compounds10. In 2011, Shen et al performed a model with 4D-fingerprints (4D-FPs) and traditional 2D and 3D VolSurf-like molecular descriptors, based on the PubChem hERG Bioassay data set containing 876 compounds — the accuracy for this model was 87% for an external test set of 356 compounds11. However, the PubChem hERG Bioassay data set was assembled from diverse sources and measured by various experimenters, which might cause the resulting model to be less reliable. In 2010, Doddareddy et al developed linear discriminant analysis (LDA) and SVM models based on a large dataset of 2644 compounds. Extended-connectivity fingerprints were used to describe chemical space. The best SVM-ECFP_6 model showed 88% accuracy for the external test set, which contained 255 compounds12. In 2013, Wang et al reviewed recent developments in computational prediction of hERG blockage, and they proposed that more reliable experimental data and a consensus modeling strategy are required to improve the performance of current computational models13.
hERG blockage data for chemicals are quickly accumulated, and a QSAR model based on a large dataset is a good approach to accurately predict the property of hERG blockage. Although Shen used PubChem containing a large amount of data and obtained a good prediction, the 4D-FP descriptors were generated based on estimations of the conformation energy profiles of molecules by molecular dynamics simulation, which is difficult to obtain11. So far, the largest dataset used for hERG blockage prediction was compiled by Doddareddy et al. They constructed SVM models that yielded good prediction accuracy but showed limited ability to interpret the mechanism of action. In this study, we used Doddareddy's data set and constructed models with a Laplacian-corrected Bayesian classification method, which considers the effects of some features in identifying toxic compounds. Simple molecular properties and extended-connectivity fingerprints were used as descriptors. These models performed well for the training and test sets and could be used for large-scale prediction.
Materials and methods
Datasets
The work reported here used the hERG blockage data from Doddareddy and co-workers12, which contained 2644 compounds, including 1479 literature compounds tested for effects on the hERG channel and 1165 FDA-approved drugs. The dataset was divided into training and test sets as described12. Activity data were expressed as an IC50 value, measured by either patch clamp or radioligand binding assays, which were conducted in human HEK-293 and CHO cell lines. Although there were some differences in the pIC50 values, these two assay results showed a consistent trend. Moreover, because we choose to build qualitative (classification) models, the quantitative variability in different assays can be considerably reduced by setting appropriate activity thresholds for separating blockers from nonblockers. Here, we employed Doddareddy's criteria, ie, the compounds with IC50 value <10 μmol/L were labeled as blockers and those with IC50 value >30 μmol/L and FDA-approved drugs as nonblockers.
Calculation of molecular properties
Many previous studies have revealed that molecular properties are relevant to hERG blockage, including the molecular lipid-water partition coefficient (ClogP), molecular weight (MW), and the molecular surface area information8,9. In this study, molecular weight (MW), molecular fractional polar surface area (FPSA), and lipophilicity (ALogP) were used. We also calculated the pKa of the most positive basic nitrogen (pKa basic) based on previous pharmacophore models7,14,15. These four molecular properties were used as simple molecular descriptors for hERG blockage modeling and were calculated with the “Calculate Molecular Properties” protocol within Discovery Studio (version 3.0; Accelrys, San Diego, CA, USA) and components in Pipeline Pilot (version 7.5; Accelrys, San Diego, CA, USA).
Extended-connectivity fingerprints (ECFPs) are a class of 2D fingerprints for capturing molecular features based on a variant of the Morgan algorithm16. An example of the generation process of ECFP for benzamide is shown in Figure 1. The ECFPs' capture atom information based on the Daylight atomic invariants rule17 and the identifiers are hashed into a single 32-bit integer value18. The number of iterations performed is determined by the maximum diameter of the neighborhoods requested. For example, “ECFP_4” generates features around each atom up to a diameter of 4, which requires two iterations. We also calculated the fingerprints with the “Calculate Molecular Properties” protocol within the Discovery Studio (version 3.0; Accelrys, San Diego, CA, USA).
Example to show the ECFP iterative generation procedure.
Modeling methods
Laplacian-corrected Bayesian classifier models were generated using Discovery Studio (version 3.0; Accelrys, San Diego, CA, USA)19,20,21. Bayesian categorization model is a statistical classification method, which use knowledge of probability and statistics based on Bayes' theorem (Eq 1):

Where A is the classification of model, B is the observed value, P(A) is the prior probability or marginal probability of classification A without considering any B factors, P(B) is the prior or marginal probability of the value, P(B|A) is the posterior probability (the probability of value B if classification A is true), and P(A|B) is the probability that classification A is true given the observed data B (also called the posterior probability)22.
We choose to build a Laplacian-corrected Bayesian classifier because it considers the complexity of the model as well as the likelihood and then picks the simplest model to explain observed data, which can avoid overfitting. The Bayesian classification technique was widely used in ADME/T predictions23,24,25. In our modeling process, the “good” samples (blockers) must be labeled first; then the model learns to distinguish the good samples from the bad samples (nonblockers). The learn-by-example process worked as follows: given a sample compound structure, the features of the sample were generated and then converted into Boolean forms. A bin was defined to count the frequency of the fingerprints and continuous values in a given range. Finally, the number of occurrences of each feature in the blocker subset, as well as in all samples, was collected. In addition, for each feature, a weight was calculated using the Laplacian-adjusted probability estimate. The Laplacian-adjusted process can be summarized as follows (Eq 2, 3, 4):



where a feature Fi is contained in B samples, and A of those B samples are active. K is a constant [virtual samples of P(Active) were added K times to stabilize the estimator to ensure more weight was assigned to the features that occurred more frequently and little weight was assigned to those that occurred less frequently]. When n features of a compound were generated, a cumulative score of feature contributions to the actives likeness (Pcombined) was computed. More details concerning the method have been described by Xia et al26.
Two different models were built: one used simple molecular properties as descriptors, and the other used simple molecular properties and ECFP fingerprints as descriptors. The Bayesian models with leave-one-out (LOO) validation were built with the training set and then evaluated with external test sets using the performance metrics described below.
Model evaluation
To evaluate the performance of the models, we used the parameter sensitivity (SE, also referred to as recall or the true-positive rate, the percentage of active compounds correctly predicted), specificity (SP, also known as the true-negative rate, the percentage of inactive compounds correctly predicted) and the global accuracy (AC, the total percentage correctly predicted). These parameters are defined as follows (Eq 5, 6, 7):



In Eqs 5,6,7, TP is the number of true positives (active compounds that are correctly predicted), TN is the number of true negatives (inactive compounds that are correctly predicted), FN is the number of false negatives (active compounds that are incorrectly predicted to be inactive), and FP is the number of false positives (inactive compounds that are incorrectly predicted to be active). The above parameters were calculated for the training and the test sets.
Results and discussion
Bayesian classifiers
The leave-one-out (LOO) statistical results for the training set for Bayesian classifiers are summarized in Table 1. The first Bayesian model based on the four simple descriptors (Model 1) showed a high sensitivity of 91% but with poor specificity of 70%. Then the ECFPs fingerprint descriptors were added to the simple descriptors; the best classifier was the model based on the simple descriptors+ECFP_14 (Model 2), which had a sensitivity of 96%, a specificity of 96%, and an overall accuracy of 96%, outperforming the best model of Doddareddy et al.
The two models were then validated with test set I, which comprised 255 molecules. As shown in Table 2, the sensitivity and specificity for Model 2 were 90% and 92%, respectively. Model 1 also gave a good sensitivity of 91% but a relatively poor specificity of 75%. Model 1 performed well on the blocking molecules but poorly on the non-blocking molecules. The performance was improved when ECFP_14 was added, especially for the predictive capability for nonblockers. This result suggested that ECFP_14 is useful for identifying the common structural features of compounds that may avoid interaction with the hERG channel.
Molecular features important for hERG
Many molecular descriptors have been proven to be helpful for hERG blockage prediction, including 2D descriptors, such as ClogP, compound molar refractivity (CMR), polarizability, partial positive/negative surface area, compound diameter, topological polar surface area, static polar surface area, fragment fingerprints and shape descriptors, 3D descriptors, such as GRIND descriptors, and 4D descriptors, such as 4D-FPs8,9,10,11,12,25,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49. Analysis of our simple, interpretable molecular properties for the compounds in the training set indicated that the molecular weight (MW), fractional polar surface area (FPSA), lipophilicity (ALogP) and pKa of the most positive basic nitrogen (pKa_basic) can preliminarily predict hERG blockage. The means and the standard deviations for the four descriptors in the training and test sets are summarized in Table 3. As shown, hERG blocking molecules have a larger molecular weight (mean value of approximately 430) compared with the nonblockers (mean value of approximately 330). In addition, compared with the nonblockers, the hERG blocking molecules have a lower FPSA value, a larger ALogP value and more basic nitrogen pKa. The mean values of the four molecular properties for the training set were listed in Figure 2. A test for significant differences is also shown. The four properties for the blockers are all significantly different from those for nonblockers (P≥0.01). The models also generated the features' statistics of the normalized probability contribution to the model (Figure 3). The features' normalized probability is the final contribution of a feature to the model's prediction, which means that the presence of a feature increases or decreases the likelihood for a compound to be a member of the good subset. The four molecular properties and their contributions to the activity identification were mapped. A positive normalized probability means that a molecule with the corresponding property value is likely to be hERG blockers, and a negative normalized probability means a molecule is likely to be nonblockers. From this figure, we can clearly find the favorable distribution ranges for properties of different molecules that cause a molecule to be more likely to be blockers or nonblockers. For example, molecules with an ALogP of −11.97∼−1, MW of 0∼227, FPSA of 0.43∼1.00, and pKa_basic of 0∼1.54 are likely to be nonblockers.
The mean value of (A) ALogP, (B) MW, (C) FPSA and (D) pKa_basic of the training set. cP<0.01 vs inactive.
The normalized probability of the four molecular properties (A) ALogP, (B) MW, (C) FPSA, and (D) pKa_basic of the training set.
Previous studies have revealed that most hERG inhibitors have two main characteristics: hydrophobic groups forming π–π stacking and protonated groups forming cation-π interaction with hERG channel residues50,51. It has also been suggested that, with increasing hydrophobicity and molecular diameter, the hERG inhibitory effect of a compound increases9,37. In our study, we found that molecules with larger ALogP and MW but smaller FPSA are more likely to be blockers, which could be ascribed to the large pore size in hERG as well as the lipophilic character of the pore cavity37. Generally, a nitrogen positive ionizable center has been considered a common feature for many hERG blockers7,50,51,52. We calculated the pKa value of the most basic nitrogen of compounds (pKa_basic) and found that the pKa_basic of non-blocking molecules is likely to be zero. The basic site with a large pKa is more likely to be protonated, and molecules with nitrogen positive ionizable centers are more likely to form a cation-π interaction with hERG channel residues.
During Laplacian-modified Bayesian modeling, the following statistics for each feature were calculated: the number of molecules in which the feature occurred, the number of those molecules that were blockers, and the measure of how different this is from the hit rate as a whole (the ratio that would be expected if the feature was occurring randomly across the blockers and nonblockers). The Bayesian score represents the final contribution of a feature to the model prediction, which takes into account the total number of occurrences of the feature, ensuring more weight is placed on features that occur more often and little weight on those for which there are very few occurrences. The top 20 fingerprint features that made the most positive contribution to the model and the top 20 that made the most negative contribution were generated as good and bad features. These fingerprint features are shown in Figure 4. The top 10 good features have a Bayesian score of 0.636, which matches to all 37 blockers (Figure 4A). These molecules are a series of 1,2,4-triazol-3-yl-thiopropyl-tetrapydrobenzazepines, which are potent and selective dopamine D3 receptor antagonists. It has been reported that D3 receptor antagonists tend to exhibit hERG toxicity53. The top 11–20 of the good features are from sertindole analogues, Wombat-PK database molecules and h5-HT2A receptor antagonists, which are mostly strong hERG blockers48,54,55. The good features can be used as structural alerts for hERG toxicity. The top 20 bad fingerprint features are listed in Figure 4B: all of these features only occurred in the nonblockers. From the prediction results we can see that the prediction for the nonblockers has been improved significantly when fingerprints were added into the descriptors: this may play important role in identifying nonblockers. For example, a series of sertindole analogues were included in our data set50, in which the most inactive analogue (compound 5) contains the fragment B13 (Figure 5B), which occurred 104 times in the whole data set but only 2 times in the blockers. As seen from the Figure 6, the two molecules (compound 4, 5) are different from each other only at one group, but the hERG channel IC50 values show a 100-fold difference. Thus, it can be hypothesized that the existence of the B13 fragment regulates the molecule to avoid the binding of the hERG channel. These fragments may affect the hERG binding affinity of compounds by changing their entire molecular shape and other physicochemical properties. Combined with the good features, the bad features may act as regulatory factors for the structural alerts listed in Figure 4A, which increases the overall accuracy of hERG toxicity prediction.
The good features (A) generated in the model. For the annotation of each fragment, G (good features) are bin IDs, the string of numbers show the bin value defining the fingerprints feature, the second line shows the frequencies this feature occurred in “good” samples and in the entire data set. The Bayesian score shows the final contribution of the feature to the model prediction.
The bad features (B) generated in the model. For the annotation of each fragment, B (bad features) are bin IDs, the string of numbers show the bin value defining the fingerprints feature, the second line shows the frequencies this feature occurred in “good” samples and in the entire data set. The Bayesian score shows the final contribution of the feature to the model prediction.
The structures of two Sertindole analogues compound 4 (A) and compound 5 (B). The B13 fragment presents in compound 5.
Prediction on an external dataset
We also applied our models to Doddareddy's experimentally validated dataset, (hereafter referred to as test set II), which contained 50 compounds predicted as blockers from the Chembridge database and 10 in-house predicted nonblockers. In Doddareddy's study, hERG blockage activity was measured by radioligand binding assays of displacement of [3H]astemizole binding to cell membranes expressing the hERG channel at 10 μmol/L concentration. Eighteen compounds of 50 predicted hERG blockers showed more than 50% displacement of [3H]astemizole in a single-point assay and 4 of those 18 showed a low micromolar and even lower nanomolar activity. We categorized the dataset into 18 blockers and 42 nonblockers based on Doddareddy's experimental results. The prediction results are listed in Table 4. Model 1 was able to identify 16 of 18 blockers, corresponding to a TP rate of 0.89, which showed a higher sensitivity than Doddareddy's. The 4 high activity compounds, VH01, VH06, VH19, and VH47, were also identified. In contrast, Model 2 correctly predicted 11 of 18 blockers and showed an increased SP of 0.57. The compounds in test set II were measured by a radioligand binding assay based on the displacement of [3H]astemizole binding, which is different from the assays used for the training set and test set I. From both Doddareddy's and our studies, we found these different assays show some variability in measuring hERG blockage, which may account for the lower prediction accuracies of these models on test set II. Doddareddy et al also developed models individually from patch clamp data and radioligand binding data, but obtained far fewer correct prediction results than the models generated using combined data from different assays. Nevertheless, compared to Doddareddy's results, our models showed more balanced results for identifying active and inactive hERG inhibitors, and they can avoid missing too many blockers during early screening.
Conclusion
In this study a Laplacian-modified Bayesian model was developed for a large data set using physicochemical properties and extended connectivity fingerprints as descriptors, and a good predictive performance was achieved, which was better than the results of the reference studies. The four simple descriptors (MW, FPSA, ALogP, pKa_basic) preliminarily predicted hERG blockage. Some structural alerts for hERG blockage and some regulating fragments contained in nonblockers were proposed. This work may provide guidance for medicinal chemists working in drug discovery and lead optimization.
Author contribution
Ming-yue ZHENG and Xiao-min LUO conceived and designed the experiments; Li-li LIU, Jing LU, and Yin LU performed the experiments; Li-li LIU, Ming-yue ZHENG and Xiao-min LUO analyzed the data; Li-li LIU, Ming-yue ZHENG and Xiao-min LUO wrote the paper: All authors discussed the results and commented on the manuscript. All authors have given approval to the final version of the manuscript.
References
Roden DM . Drug-induced prolongation of the QT interval. N Engl J Med 2004; 350: 1013–22.
Mitcheson JS, Chen J, Lin M, Culberson C, Sanguinetti MC . A structural basis for drug-induced long QT syndrome. Proc Natl Acad Sci U S A 2000; 97: 12329–33.
Curran ME, Splawski L, Timothy KW, Vincent GM, Green ED, Keating MT . A molecular basis for cardiac arrhythmia hERG mutations cause long QT syndrome. Cell 1995; 80: 795–803.
Keating MT, Sanguinetti MC . Molecular genetic insights into cardiovascular disease. Science 1996; 272: 681–5.
Babcock JJ, Li M . hERG channel function: beyond long QT. Acta Pharmacol Sin 2013; 34: 329–35.
Lin K, Liu YQ, Xu B, Gao JL, Fu YC, Chen Y, et al. Allocryptopine and benzyltetrahydropalmatine block hERG potassium channels expressed in HEK293 cells. Acta Pharmacol Sin 2013; 34: 847–58.
Cavalli A, Poluzzi E, Ponti FD, Recanatini M . Toward a phamacophore for drugs inducing the long QT syndrome: insights from a CoMFA study of HERG K+ channel blockers. J Med Chem 2002; 45: 3844–53.
Keser GM . Prediction of hERG potassium channel affinity by traditional and hologram qSAR methods. Bioorg Med Chem Lett 2003; 13: 2773–5.
Yoshida K, Niwa T . Quantitative structure-activity relationship studies on inhibition of hERG potassium channels. J Chem Inf Model 2006; 46: 1371–8.
Li QY, Jorgensen FS, Oprea T, Brunak S, Taboureau O . hERG classification model based on a combination of support vector machine method and GRIND descriptors. Mol Pharm 2008; 5: 117–27.
Shen MY, Su BH, Esposito EX, Hopfinger AJ, Tseng YJ . A comprehensive support vector machene binary hERG classification model based on extensive but biased end point hERG data set. Chem Rex Toxicol 2011; 24: 934–49.
Doddareddy MR, Klaasse EC, Shagufta I, Jzerman AP, Bender A . Prospective validation of a comprehensive in silico hERG model and its applications to commercial compound and drug databases. Chem Med Chem 2010; 5: 716–29.
Wang SC, Li YY, Xu L, Li D, Hou TJ . Recent developments in computational prediction of hERG blockage. Curr Topics Med Chem 2013; 13: 1317–26.
Ekins S, Crumb WJ, Sarazan RD, Wikel JH, Wrighton SA . Three-dimensional quantitative structure-activity relationship for inhibition of human ether-a-go-go-related gene potassium channel. J Pharmacol Exp Ther 2002; 301: 427–34.
Pearlstein R, Vaz R, Rampe D . Understanding the structure-activity relationship of the human ether-a-go-go-related gene cardiac K+ channel. A model for bad behavior. J Med Chem 2003; 46: 2017–22.
Morgan HL . The generation of a unique machine description for chemical structures-a techinque developed at chemical abstracts service. J Chem Doc 1965; 5: 107–12.
Weininger D, Weininger A, Weininger JL . SMILES. 2. Algorithm for generation of unique SMILES notation. J Chem Inf Comput Sci 1989; 29: 97–101.
David R, Mathew H . Extended-connectivity fingerprints. J Chem Inf Model 2010; 50: 742–54.
Prathipati P, Ma NL, Keller TH . Global bayesian models for the prioritization of antitubercular agents. J Chem Inf Model 2008; 48: 2362–70.
Klon AE, Lowrie JF, Diller DJ . Improved naïve bayesian modeling of numeriacl data for absorption, destribution, metabolism and excretion (ADME) property prediction. J Chem Inf Model 2006; 46: 1945–56.
Rogers D, Brown RD, Hahn M . Using extended-connectivity fingerprints with Laplacian-modified Bayesian analysis in high-throughput screening follow-up. J Biomol Screening 2005; 10: 682–6.
Zientek M, Stoner C, Ayscue R, McLeod-Klug J, Jiang Y, West M, et al. Integrated in silico-in vitro strategy for addressing cytochrome P450 3A4 time-dependent inhibition. Chem Res Toxicol 2010; 23: 664–76.
Chen L, Li YY, Zhao Q, Peng H, Hou TJ . ADME evaluation in drug discovery. 10. Predictions of p-glycoprotein inhibitors using recursive partitioning and naïve bayesian classification techniques. Mol Pharm 2011; 8: 889–900.
Tian S, Wang, JM, Li YY, Xu XJ, Hou TJ . Drug-likeness analysis of traditional Chinese medicines: prediction of drug-likeness using machine learning approaches. Mol Pharm 2012; 9: 2875–86.
Wang SC, Li YY, Wang JM, Chen L, Zhang LL, Yu HD, et al. ADMET evaluation in drug discovery. 12. development of binary classification models for pridiction of hERG potassium channel blockage. Mol Pharm 2012; 9: 996–1010.
Xia XY, Maliski EG, Gallant P, Rogers D . Classification of kinase inhibitors using a bayesian model. J Med Chem 2004; 47: 4463–70.
Roche O, Trube G, Zuegge J, Pflimlin P, Alanine A, Schneider G . A virtual screening method for prediction of the hERG potassium channel liability of compound libraries. Chem Biol Chem 2002; 3: 455–9.
Zolotoy1 AB, Plouvier BP, Beatch GB, Hayes ES, Wall RA, Walker MJ . Physicochemical determinants for drug induced blockade of HERG potassium channels effect of charge and charge shielding. Curr Med Chem Cardiovasc Hematol Agents 2003; 1: 225–41.
Aptula AO, Cronin MT . Prediction of hERG K+ blocking potency application of structural knowledge. SAR QSAR Environ Res 2004; 15: 399–411.
Song MH, Clark M . Development and evaluation of an in silico model for hERG binding. J Chem Inf Model 2006; 46: 392–400.
Bains W, Basman A, White C . HERG binding specificity and binding site structure: evidence from a fragment-based evolutionary computing SAR study. Prog Biophys Mol Biol 2004; 86: 205–233.
Ekins S, Balakin KV, Savchuk N, Ivanenkov Y . Insights for human ether-a-go-go-related gene potassium channel inhibition using recursive partitioning and kohonen and sammon mapping techniques. J Med Chem 2006; 49: 5059–71.
Aronov AM . Common pharmacophores for uncharged human ether-a-go-go-related gene (hERG) blockers. J Med Chem 2006; 49: 6917–21.
Chekmarev DS, Kholodovych V, Balakin KV, Ivanenkov Y, Ekins S, Welsh WJ . Shape signatures new descriptors for predicting cardiotoxicity in silico. Chem Res Toxicol 2008; 21: 1304–14.
Raschi E, Ceccarini L, Ponti FD, Recanatini M . hERG-related drug toxicity and models for predicting hERG liability and QT prolongation. Expert Opin Drug Metab Toxicol 2009; 5: 1005–21.
Clark M, Wiseman JS . Fragment-based prediction of the clinical occurrence of long QT syndrome and torsade de pointes. J Chem Inf Model 2009; 49: 2617–26.
Aronov A . Predictive in silico modeling for hERG channel blockers. Drug Discovery Today 2005; 10: 149–55.
Aronov AM, Goldman BB . A model for identifying HERG K+ channel blockers. Bioorg Med Chem 2004; 12: 2307–15.
Cianchetta G, Li Y, Kang J, Rampe D, Fravolini A, Cruciani G, et al. Predictive models for hERG potassium channel blockers. Bioorg Med Chem Lett 2005; 15: 3637–42.
Du-Cuny L, Chen L, Zhang S . A critical assessment of combined ligand- and structure-based approaches to HERG channel blocker modeling. J Chem Inf Model 2011; 51: 2948–60.
Gavaghan CL, Arnby CH, Blomberg N, Strandlund G, Boyer S . Development, interpretation and temporal evaluation of a global QSAR of hERG electrophysiology screening data. J Comput Aided Mol Des 2007; 21: 189–206.
Gepp MM, Hutter MC . Determination of hERG channel blockers using a decision tree. Bioorg Med Chem 2006; 14: 5325–32.
Kramer C, Beck B, Kriegl JM, Clark T . A composite model for HERG blockade. Chem Med Chem 2008; 3: 254–65.
Seierstad M, Agrafiotis DK . A QSAR model of HERG binding using alarge, diverse, and internally consistent training set. Chem Biol Drug Design 2006; 67: 284–96.
Tan Y, Chen Y, You Q, Sun H, Li M . Predicting the potency of hERG K(+) channel inhibition by combining 3D-QSAR pharmacophore and 2D-QSAR models. J Mol Model 2012; 18: 1023–36.
Thai KM, Ecker GF . A binary QSAR model for classification of hERG potassium channel blockers. Bioorg Med Chem 2008; 16: 4107–19.
Tobita M, Nishikawa T, Nagashima R . A discriminant model constructed by the support vector machine method for HERG potassium channel inhibitors. Bioorg Med Chem Lett 2005; 15: 2886–90.
Zachariae U, Giordanetto F, Leach AG . Side chain flexibilities in the human ether-a-go-go related gene potassium channel (hERG) together with matched-pair binding studies suggest a new binding mode for channel blockers. J Med Chem 2009; 52: 4266–76.
Su BH, Shen MY, Esposito EX, Hopfinger AJ, Tseng YJ . In silico binary classification QSAR models based on 4D-fingerprints and MOE descriptors for prediction of hERG blockage. J Chem Inf Model 2010; 50: 1304–18.
Pearlstein RA, Vaz RJ, Kang J, Chen XL, Preobrazhenskaya M, Shchekotikhin AE, et al. Characterization of HERG potassium channel inhibition using CoMSiA 3D QSAR and homology modeling approaches. Bioorg Med Chem Lett 2003; 13: 1829–35.
Pearlstein R, Vaz R, Rampe D . Understanding the structure-activity relationship of the human ether-a-go-go-related gene cardiac K+ channel. A model for bad behavior. J Med Chem 2003; 46: 2017–22.
Ekins S, Crumb WJ, Sarazan RD, Wikel JH, Wrighton SA . Three-dimensional quantitative structure-activity relationship for inhibition of human ether-a-go-go-related gene potassium channel. J Pharmacol Exp Ther 2002; 301: 427–34.
Micheli F, Bonanomi G, Blaney FE, Braggio S, Capelli AM, Checchia A, et al. 1,2,4-triazol-3-yl-thiopropyl-tetrahydrobenzazepines: A series of potent and selective dopamine D3 receptor antagonists. J Med Chem 2007; 50: 5076–89.
Taboureau O, Nielsen SK, Audouze K, Weinhold N, Edsgard D, Roque FS, et al. ChemProt: a disease chemical biology database. Nucleic Acids Res 2011; 39: 367–72.
Rowley M, Hallett DJ, Goodacre S, Moyes C, Crawforth J, Sparey TJ, et al. 3-(4-fluoropiperidin-3-yl)-2-phenylindoles as high affinity, selective, and orally bioavailable h5-HT2A receptor antagonists. J Med Chem 2001; 44: 1603–14.
Acknowledgements
This work was supported by Hi-TECH Research and Development Program of China (Grant 2012AA020308), National S&T Major Project (Grant 2012ZX09301, 2014ZX09507002), and National Natural Science Foundation of China (81220108025, 81001399, 2013ZX09507001).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Liu, Ll., Lu, J., Lu, Y. et al. Novel Bayesian classification models for predicting compounds blocking hERG potassium channels. Acta Pharmacol Sin 35, 1093–1102 (2014). https://doi.org/10.1038/aps.2014.35
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2014.35
Keywords
This article is cited by
-
Artificial intelligence and machine-learning approaches in structure and ligand-based discovery of drugs affecting central nervous system
Molecular Diversity (2023)
-
Facing small and biased data dilemma in drug discovery with enhanced federated learning approaches
Science China Life Sciences (2022)
-
A critical analysis of urea transporter B inhibitors: molecular fingerprints, pharmacophore features for the development of next-generation diuretics
Molecular Diversity (2022)
-
First structure–activity relationship analysis of SARS-CoV-2 virus main protease (Mpro) inhibitors: an endeavor on COVID-19 drug discovery
Molecular Diversity (2021)
-
Structural analysis of arylsulfonamide-based carboxylic acid derivatives: a QSAR study to identify the structural contributors toward their MMP-9 inhibition
Structural Chemistry (2021)